For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VTTPTRWAGVPLTDRRAERRALLVDAAFRLFGDGGETALSVRSVCRECGLNTRYFYESFADTDDLIGAVY DEVSALLAADVEAAMEGAGNSLRARTRAGIAAVLGFSSADPRRGRVLFTDARANPVLAQRRAVTQDLLRE AVLSEGGRLNPGSDPVAAQVGAAMYTGAMAELAQQWLAGNLGDDLDAVVDHALRLVLGR
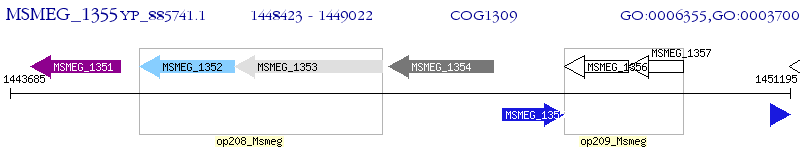
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_1355 | - | - | 100% (199) | transcriptional regulator, TetR family protein |
| M. smegmatis MC2 155 | MSMEG_5768 | - | 2e-22 | 34.85% (198) | hypothetical protein MSMEG_5768 |
| M. smegmatis MC2 155 | MSMEG_0612 | - | 9e-19 | 33.67% (199) | transcriptional regulator, TetR family protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0281c | - | 3e-24 | 36.22% (196) | TetR family transcriptional regulator |
| M. gilvum PYR-GCK | Mflv_5109 | - | 8e-87 | 83.42% (193) | TetR family transcriptional regulator |
| M. tuberculosis H37Rv | Rv0275c | - | 2e-23 | 35.71% (196) | TetR family transcriptional regulator |
| M. leprae Br4923 | MLBr_01733 | - | 3e-05 | 27.49% (171) | putative TetR-family transcriptional regulator |
| M. abscessus ATCC 19977 | MAB_3300c | - | 2e-24 | 36.87% (198) | TetR family transcriptional regulator |
| M. marinum M | MMAR_4855 | - | 2e-24 | 36.27% (193) | hypothetical protein MMAR_4855 |
| M. avium 104 | MAV_3986 | - | 1e-85 | 79.70% (197) | transcriptional regulator, TetR family protein |
| M. thermoresistible (build 8) | TH_4000 | - | 1e-91 | 82.61% (207) | PUTATIVE transcriptional regulator, TetR family |
| M. ulcerans Agy99 | MUL_0446 | - | 2e-24 | 36.27% (193) | hypothetical protein MUL_0446 |
| M. vanbaalenii PYR-1 | Mvan_1244 | - | 9e-90 | 86.67% (195) | TetR family transcriptional regulator |
CLUSTAL 2.0.9 multiple sequence alignment
Mb0281c|M.bovis_AF2122/97 MTRS-----DRP--YRGVE-------------------------------
Rv0275c|M.tuberculosis_H37Rv MTRS-----DRP--YRGVE-------------------------------
MMAR_4855|M.marinum_M MQAG-----QRRGRWTGVP-------------------------------
MUL_0446|M.ulcerans_Agy99 MQAG-----QRRGRWTGVP-------------------------------
MSMEG_1355|M.smegmatis_MC2_155 MT--------TPTRWAGVP-------------------------------
TH_4000|M.thermoresistible__bu MT--------SPTRWAGVP-------------------------------
Mflv_5109|M.gilvum_PYR-GCK MSGAP-----HATRWAGVP-------------------------------
Mvan_1244|M.vanbaalenii_PYR-1 MSSAASRPAATPTRWAGVP-------------------------------
MAV_3986|M.avium_104 MS--------SPSRWAGVP-------------------------------
MAB_3300c|M.abscessus_ATCC_199 ---------MPSRTYGGAT-------------------------------
MLBr_01733|M.leprae_Br4923 MTRRS-IHTRKPLKWSGCAGQHVRSGKVFDVGRISLSYPSMKPGVKIDAR
: *
Mb0281c|M.bovis_AF2122/97 ---AAERLATRRRQLLSAGLDLLGSDQHDIAELTIRTICRRAGLSVRYFY
Rv0275c|M.tuberculosis_H37Rv ---AAERLATRRRQSLSAGLDLLGSDQHDIAELTIRTICRRAGLSVRYFY
MMAR_4855|M.marinum_M ---LEDRHAVRRDTLIAAGVQLLGEEAG--PALTVRAVCRQAALTERYFY
MUL_0446|M.ulcerans_Agy99 ---LEDRHAVRRDTLIAAGVQLLGEEAG--PALTVRAVCRQAALTERYFY
MSMEG_1355|M.smegmatis_MC2_155 ---LTDRRAERRALLVDAAFRLFGDGGE--TALSVRSVCRECGLNTRYFY
TH_4000|M.thermoresistible__bu ---LTDRRAERRALLIAAAYRLFGEGGE--AALSVRSVCRESELNSRYFY
Mflv_5109|M.gilvum_PYR-GCK ---LTDRRAERRALLVDAAFRLFGDGGE--SAVSVRSVSRECGLNTRYFY
Mvan_1244|M.vanbaalenii_PYR-1 ---LTDRRAERRALLVDAAFRLFGDGGE--QALSVRSVSREAGLNTRYFY
MAV_3986|M.avium_104 ---LKDRRAERRALLVEAAYRLFGSEGE--AAVSVRSVCRECGLNTRYFY
MAB_3300c|M.abscessus_ATCC_199 ---SGERRARRRAALLDAALDLVAGSGV--KGLTVRGVCAEARLNDRYFY
MLBr_01733|M.leprae_Br4923 SERWREHRKKVRGEIVDAAFRALDRLGP---EVSVREIAEEAGTAKPKIY
:: * : *. . :::* :. .. :*
Mb0281c|M.bovis_AF2122/97 ESFTDKDEFVGRVFDWVVAELVATTQAAV----------TAVPAREQTRA
Rv0275c|M.tuberculosis_H37Rv ESFTDKDEFVGRVFDWVVAELVATTQAAV----------TAVPAREQTRA
MMAR_4855|M.marinum_M ESFSDRDHFVRAVYDDVCTRAMATLMSAN----------TPRDAVER---
MUL_0446|M.ulcerans_Agy99 ESFSDRDHFVRAVYDDVCTRAMATLMSAN----------TPRDAVER---
MSMEG_1355|M.smegmatis_MC2_155 ESFADTDDLIGAVYDEVSALLAADVEAAM--------EGAGNSLRARTRA
TH_4000|M.thermoresistible__bu ESFADVDELLGAVYDEVSAQLAAEVETAMAAAVEGAGEGAGDSLRARTRA
Mflv_5109|M.gilvum_PYR-GCK ESFGGTDDLLGAVYDRVSAELSVAVADAM--------DHAEDSLRARTRA
Mvan_1244|M.vanbaalenii_PYR-1 ESFTDTDDLLGAVYDRVSAELAEVVEAAM--------SGAGASLPARTRA
MAV_3986|M.avium_104 ESFRDTDDLLGAVYDRVSEQLEEVIAAAI--------EQAGDSLRARTRA
MAB_3300c|M.abscessus_ATCC_199 ESFRSTDELVLAMLDEQMLAGSATIIEAIAKS------STSKDPVVRTRA
MLBr_01733|M.leprae_Br4923 RHFTDKSDLFEAIGIRLRDMLWAAIFPAID--------LATDSAREIIRR
. * . ..:. : * :
Mb0281c|M.bovis_AF2122/97 GMANIVRTITADARVGRLLFST--QLANAVITRKRAESSALFAMLSGQ-H
Rv0275c|M.tuberculosis_H37Rv GMANIVRTITADARVGRLLFST--QLANAVITRKRAESSALFAMLSGQ-H
MMAR_4855|M.marinum_M ----FVALMVDDPVRGRVLLLA--PAVEPVLTRSGADWMPNFIDLLQR-K
MUL_0446|M.ulcerans_Agy99 ----FVALMVDDPVRGRVLLLA--PAVEPVLTRSGADWMPNFIDLLQR-K
MSMEG_1355|M.smegmatis_MC2_155 GIAAVLGFSSADPRRGRVLFTD--ARANPVLAQRRAVTQDLLREAVLS-E
TH_4000|M.thermoresistible__bu GIVAVLGFSSADPRRGRILFTD--ARANPVLAARRAATQDQLREAVLT-E
Mflv_5109|M.gilvum_PYR-GCK GMAAVLHFSSADPRRGRVLFTD--ARSNPVLTARRAATQDLLREAVLT-E
Mvan_1244|M.vanbaalenii_PYR-1 GMTAVLHFSSADPRRGRVLFTD--ARANPVLTQRRAATQDLLREAVLT-E
MAV_3986|M.avium_104 GIAAVLGFSSADPRRGRVLFTD--ARTNPVLAARRRATQDMLRQGVLS-E
MAB_3300c|M.abscessus_ATCC_199 AVGSGLRFLTADPRRGRLLVES--QATEGLAARRRDLVKTLAQVMADQGR
MLBr_01733|M.leprae_Br4923 SVDEYVSLVDKHPNVLRVFIQGRFSAQSEATVRTLDEGREITLTMAEMFN
: .. *::. . . .
Mb0281c|M.bovis_AF2122/97 AVDTLHAPANDHVKAVAHFAVGGVGQTISAWLAGDVR----LDPDQLVDQ
Rv0275c|M.tuberculosis_H37Rv AVDTLHAPANDHVKAVAHFAVGGVGQTISAWLAGDVR----LDPDQLVDQ
MMAR_4855|M.marinum_M LSQIADP---VLQKLVATSLIGALTGLFTAYLNGQLG----ATRQQFIDY
MUL_0446|M.ulcerans_Agy99 LSQIADP---VLQKLVATSLIGALTGLFTAYLNGQLG----ATRQQFIDY
MSMEG_1355|M.smegmatis_MC2_155 GGRLNPGSDPVAAQVGAAMYTGAMAELAQQWLAGNLG----DDLDAVVDH
TH_4000|M.thermoresistible__bu GWRLHPKSDPVAAQVGAAMYTGAMAELAQQWLAGNLG----DDLDRVVDY
Mflv_5109|M.gilvum_PYR-GCK GWRLHPDTDPVAAATGAAMYTGAMAELAQQWLSGNLG----DDLDVVVDH
Mvan_1244|M.vanbaalenii_PYR-1 GWRLHPDSDPTAAMVGAAMYTGAMAELAQQWLAGHLG----DDLDAVVGY
MAV_3986|M.avium_104 GWRLNPRSDPVAAEVAAAMYTGAMAELAQQWLAGHLG----GDLDAVVDY
MAB_3300c|M.abscessus_ATCC_199 VLLPADDALDPDLDLATFTLVSGGLDLVTTWFRGELD----ITRDRLEDF
MLBr_01733|M.leprae_Br4923 NELREMELNRAASELAAFAAFGSVASATEWWLGPEPDSPRRMPREQFVAH
: .. :: . : .
Mb0281c|M.bovis_AF2122/97 LAALLDELTDPNLSRPRVAATAAKSGANDPQPPEVAGQPPSSARPARRS
Rv0275c|M.tuberculosis_H37Rv LAALLDELTDPNLSRPRVAATAAKSGANDPQPPEVAGQPPSSARPARRS
MMAR_4855|M.marinum_M CVDLLLG---------TAAAYAPHRDRGEAERPVAARQHD---------
MUL_0446|M.ulcerans_Agy99 CVDLLLG---------TAAAYAPHRDRGEAEHPVAARQHD---------
MSMEG_1355|M.smegmatis_MC2_155 ALRLVLGR-----------------------------------------
TH_4000|M.thermoresistible__bu ALRLVLGR-----------------------------------------
Mflv_5109|M.gilvum_PYR-GCK ALRLVLR------------------------------------------
Mvan_1244|M.vanbaalenii_PYR-1 ALRLVLR------------------------------------------
MAV_3986|M.avium_104 ASKLVLR------------------------------------------
MAB_3300c|M.abscessus_ATCC_199 LVALVTR---------ANIRADQHEDSGEAAKPGN--------------
MLBr_01733|M.leprae_Br4923 LTIIMMAVIVGTAETLGISMNPDRPIHDAVPLDRSAS------------
::