For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
SQTARRLGPQDMFFLYSESSTTMMHVGALMPFTPPSGAPPDLLRQLVDESKASEVVEPWSLRLSHPELLY HPTQSWVVDDNFDLDYHVRRSALASPGDERELGIPVSRLHSHALDLRRPPWEVHFIEGLEGGRFAIYIKM HHSLIDGYTGQKMLARSLSTDPHDTTHPLFFNIPTPGRSPADTQDSVGGGLIAGAGNVLDGLGDVVRGLG GLVSGVGSVLGSVAGAGRSTFELTKALVNAQLRSDHEYRNLVGSVQAPHCILNTRISRNRRFATQQYPLD RLKAIGAQYDATINDVALAIIGGGLRRFLDELGELPNKSLIVVLPVNVRPKDDEGGGNAVATILATLGTD VADPVQRLAAVTASTRAAKAQLRSMDKDAILAYSAALMAPYGVQLASTLSGVKPPWPYTFNLCVSNVPGP EDVLYLRGSRMEASYPVSLVAHSQALNVTLQSYAGTLNFGFIGCRDTLPHLQRLAVYTGEALDQLAAADG AAGLGS*
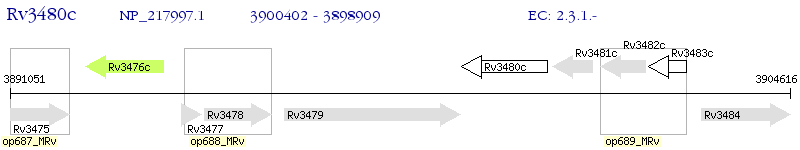
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. tuberculosis H37Rv | Rv3480c | - | - | 100% (497) | hypothetical protein Rv3480c |
| M. tuberculosis H37Rv | Rv3734c | - | 4e-88 | 38.68% (486) | hypothetical protein Rv3734c |
| M. tuberculosis H37Rv | Rv3740c | - | 5e-83 | 38.60% (487) | hypothetical protein Rv3740c |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3509c | - | 1e-154 | 100.00% (274) | hypothetical protein Mb3509c |
| M. gilvum PYR-GCK | Mflv_1058 | - | 1e-174 | 61.15% (489) | hypothetical protein Mflv_1058 |
| M. leprae Br4923 | MLBr_01244 | - | 2e-24 | 27.87% (427) | hypothetical protein MLBr_01244 |
| M. abscessus ATCC 19977 | MAB_4544c | - | 3e-76 | 36.55% (476) | hypothetical protein MAB_4544c |
| M. marinum M | MMAR_4955 | - | 0.0 | 88.07% (486) | hypothetical protein MMAR_4955 |
| M. avium 104 | MAV_0355 | - | 1e-79 | 37.65% (486) | bifunctional wax ester synthase/acyl-coadiacylglycerol |
| M. smegmatis MC2 155 | MSMEG_6322 | - | 1e-84 | 38.98% (490) | bifunctional wax ester synthase/acyl-CoA diacylglycerol |
| M. thermoresistible (build 8) | TH_2030 | - | 6e-83 | 39.59% (490) | CONSERVED HYPOTHETICAL PROTEIN |
| M. ulcerans Agy99 | MUL_4351 | - | 2e-80 | 38.02% (484) | hypothetical protein MUL_4351 |
| M. vanbaalenii PYR-1 | Mvan_5773 | - | 1e-166 | 60.84% (475) | hypothetical protein Mvan_5773 |
CLUSTAL 2.0.9 multiple sequence alignment
Rv3480c|M.tuberculosis_H37Rv MSQTARRLGPQDMF----FLYSESSTTMMHVGALMPFTPP-SGAPPDLLR
Mb3509c|M.bovis_AF2122/97 --------------------------------------------------
MMAR_4955|M.marinum_M MSKTAKRLGPQDMF----FLYSESPSTMMHVGALMPFTPP-DDAPADFLR
Mflv_1058|M.gilvum_PYR-GCK MSANSVRLGLQDLM----FIYGETSSSKMHVGGLLPFTPP-ADAPRDYLR
Mvan_5773|M.vanbaalenii_PYR-1 -------------M----FIYGETSSSKMHVAGLLPFTPP-ADAPRDYLR
MSMEG_6322|M.smegmatis_MC2_155 -MNRMQLMSPTDSM----FLIAESREHPMHVGGLALYDPP-DDAGPEFVR
TH_2030|M.thermoresistible__bu ----MRLISPADSM----FLLAESREHPMHVAGLQLFEPPPGTAGPEFLQ
MAV_0355|M.avium_104 ----MELIMPTDAV----FLLGESREHPMHVGGLQLYQPP-EGAGRNFVR
MUL_4351|M.ulcerans_Agy99 -------MSPVDAL----FLSAESREHPLHVGALQLFEPP-EGAGRAFVH
MAB_4544c|M.abscessus_ATCC_199 ----MPFMPVSDSM----FLIGETREHPFHVGGLQLFKPP-VDAGPDYAE
MLBr_01244|M.leprae_Br4923 MADSVEGIGPFDELGALDYLLHRGEANPRTRAGIMAVELLDTTPDWNRFR
Rv3480c|M.tuberculosis_H37Rv QLVDESKASEVVEPWSLRLSHPELLYHPTQSWVVDDNFDLDYHVRRSALA
Mb3509c|M.bovis_AF2122/97 --------------------------------------------------
MMAR_4955|M.marinum_M QLVDESKANEVVEPWNLKLSHPDLLYSPTQSWVYDENFDLDYHVRRSALA
Mflv_1058|M.gilvum_PYR-GCK GMIDEARRQEVVAPWNRRLAYPRLQFSPLHSWVTDEDFDFDYHVRRSALA
Mvan_5773|M.vanbaalenii_PYR-1 RMVDDARRQEVMPPWNRKLAYPRLQFSPLHTWVTDTEFDFDYHVRRSALA
MSMEG_6322|M.smegmatis_MC2_155 ELYEEMVRHTDFQPVFRKHPATLLGGIANVGWTLDDEVDLDYHLRRSALP
TH_2030|M.thermoresistible__bu GLHQRLAAGRELHPTFRKHPATLFGGIAHLGWAVDDDVDFDYHLRRSALP
MAV_0355|M.avium_104 DFYKQLVAQPDFQPTFRKHPAKFVGGIANLGWSYDADVDVDYHVRRSALP
MUL_4351|M.ulcerans_Agy99 ETHQAMLDCREVAPLFRKRPASFQGVLTNLGWSVDDDIDFGFHVRRSALP
MAB_4544c|M.abscessus_ATCC_199 VFYEQLMSTTEVSSDFRKRPGQPLAVMGNLTWIVDDEVDWEYHVRRSALP
MLBr_01244|M.leprae_Br4923 SRIEDVSQR---VLRLRQKVVVPTLPTAAPRWVVDPDFELDFHVRRVRVP
Rv3480c|M.tuberculosis_H37Rv SPGDERELGIPVSRLHSHALDLRRPPWEVHFIEGLEGGRFAIYIKMHHSL
Mb3509c|M.bovis_AF2122/97 --------------------------------------------------
MMAR_4955|M.marinum_M SPGDERELGILVSRLHSHALDLRRPPWEMHFIEGLEGGRFAIYIKMHHSL
Mflv_1058|M.gilvum_PYR-GCK SPGDERELGVLVSRLHSNHLDLTRPPWELHVIEGLEGGRFALYMKIHHAL
Mvan_5773|M.vanbaalenii_PYR-1 SPGDERELGILVSRLHSNHLDLTRPPWELHVIEGLEGGRFALYMKIHHAL
MSMEG_6322|M.smegmatis_MC2_155 SPGRPRELLELTSRVHGTLLDRHRPLWEAYLIEGMADGRFAVYTKVHHSL
TH_2030|M.thermoresistible__bu GPGRIRELLDLTSRWHGTLLDRHRPLWEVHLVEGLADGRVAIYFKVHHAL
MAV_0355|M.avium_104 SPRRVRELLELTSRWHSSLLDRHRPLWETHIVEGLKDGRFAIYTKVHHAL
MUL_4351|M.ulcerans_Agy99 APGRVRELLELTSRLHAHLLDRHRPLWETHVIEGLNDGRFAIYSKMHHAL
MAB_4544c|M.abscessus_ATCC_199 RPARVRELLTVTSRWHSSPLDRHRPLWEMHVVEGLSDGRLAVYTKMHHAV
MLBr_01244|M.leprae_Br4923 DPGTLREVFDLAEVIQQSPMDVSRPLWTATLVEGLAAGRAAMLLQISHAI
Rv3480c|M.tuberculosis_H37Rv IDGYTGQKMLARSLSTDPHDTTHPLFFNIPTPGRSPADTQDSVGGGLIAG
Mb3509c|M.bovis_AF2122/97 --------------------------------------------------
MMAR_4955|M.marinum_M IDGYTGQKMLARSLSADPHDTTHPLFFNIPTPGPAPADTEESAGGGLIGG
Mflv_1058|M.gilvum_PYR-GCK VDGYSAMRMLGRSLSTDPASRDTRMFFNVPSPTRSRRD----------PG
Mvan_5773|M.vanbaalenii_PYR-1 VDGYTAMRMLGRSLSTDPETRDARMFFNVPIPKRSRPT----------GA
MSMEG_6322|M.smegmatis_MC2_155 IDGVSAMKLVERTLSEDPSDTTVRVPWNLPRRESSRRAGSSS--------
TH_2030|M.thermoresistible__bu IDGVAVMKLMQRTLSTDPGDD-ARVPWNLPPPRRNP-AGPVS--------
MAV_0355|M.avium_104 IDGVSAQKLMQRALSSDPDDPEIRAPWTLP--KRSRKAGPSS--------
MUL_4351|M.ulcerans_Agy99 IDGVSGLALMRRSLPADPADTNFRATWSPAPERPRGVQTRPG--------
MAB_4544c|M.abscessus_ATCC_199 IDGVGALRMMMRSLSDDPDARDCQAVWAPAKRRAKSSIVSTTN-------
MLBr_01244|M.leprae_Br4923 TDGVGSVEMFAEMYDLERDPPSRPRSPQPIPQDLSRNDLMLQG-------
Rv3480c|M.tuberculosis_H37Rv AGNVLDGLGDVVRGLGGLVSGVGSVLGSVAGAGRSTFELTKALVNAQLRS
Mb3509c|M.bovis_AF2122/97 ----------------------------MAGAGRSTFELTKALVNAQLRS
MMAR_4955|M.marinum_M AGNVLSGLGDVVKGLGSMVSGVGSVLGSVASAGRSSLDLTRALVNSQLRT
Mflv_1058|M.gilvum_PYR-GCK AAESSNPLTATLRALGGVSS-------AVTGGVSSAVDLTNALVNTQIRR
Mvan_5773|M.vanbaalenii_PYR-1 GAQSSNPVTATLRALGAVGS-------TVSDGVGSAVDLASALVNTQIRR
MSMEG_6322|M.smegmatis_MC2_155 ----------LARTATGAAT-------SLAALAPSTIRLARAALLEQQ--
TH_2030|M.thermoresistible__bu ----------RLRSATGAVG-------SVAALAPSTLSLARAALLEQR--
MAV_0355|M.avium_104 ----------RLSSLVHAAG-------SVAALAPSTVSLARAALVEQQ--
MUL_4351|M.ulcerans_Agy99 ----------PLRQLGGMLG-------SVAGLAPSSLRLARSALLEQQ--
MAB_4544c|M.abscessus_ATCC_199 -----TSAIDLVKGAAGAVR-------QVVSIPPGLLKYGRHALSDPQ--
MLBr_01244|M.leprae_Br4923 ------INHLPVALAGGVVGG----LSGVASVAGRAILRPASTVSGVVGY
: . :
Rv3480c|M.tuberculosis_H37Rv DHEYRNLVGSVQAPHCILNTRISRNRRFATQQYPLDRLKAIGAQYDATIN
Mb3509c|M.bovis_AF2122/97 DHEYRNLVGSVQAPHCILNTRISRNRRFATQQYPLDRLKAIGAQYDATIN
MMAR_4955|M.marinum_M DNEYRDLVSSVQAPHCILNTRISRNRRFATQQYPLARMKAIGAQYDATVN
Mflv_1058|M.gilvum_PYR-GCK DGENAHIAGSVSAPHSILNARISRNRRFATQQYEFDRLKKLSSQHGATLN
Mvan_5773|M.vanbaalenii_PYR-1 DGDFGRISGSASAPHSILNARISRNRRFATQQYEFDRLKKLSSQHGATIN
MSMEG_6322|M.smegmatis_MC2_155 ------LTLPFGAPRTMFNVKIGGARRVAAQSWPLERLRRIKAVTGATIN
TH_2030|M.thermoresistible__bu ------LTLPFGAPRTMFNVRIGGARRVAAQSWPLQRIRRIRQAAGXTLN
MAV_0355|M.avium_104 ------LTLPFGAPRTMLNVKIGGARRCAAQSWPLERIKNVKNAAGVTVN
MUL_4351|M.ulcerans_Agy99 ------LTLPFATPRTMLNVAIGGARRCAAQSWDIDRVETVKNAAGVSLN
MAB_4544c|M.abscessus_ATCC_199 ------LVKPLSAPHTMLNVSIGGARRFAAQTWSMARIKEAGKALGGTVN
MLBr_01244|M.leprae_Br4923 VRSGIRVLSQAAEPSPLLRQRS-LATRTEAIEIQLSDLHKAAKAGDGSIN
: * ::. * : : :. . ::*
Rv3480c|M.tuberculosis_H37Rv DVALAIIGGGLRRFLDELGELPNKSLIVVLPVNVRPKDDE-GGGNAVATI
Mb3509c|M.bovis_AF2122/97 DVALAIIGGGLRRFLDELGELPNKSLIVVLPVNVRPKDDE-GGGNAVATI
MMAR_4955|M.marinum_M DVAMAIIGGGLRRFLDELGELPDKSLIAVLPVNVRPKDDE-GGGNAVATI
Mflv_1058|M.gilvum_PYR-GCK DVALAIIGGGLRKFLSDFDKLPDRSLIAFLPVNVRPKGDE-GGGNAVGAI
Mvan_5773|M.vanbaalenii_PYR-1 DVALAIIGGGLRKFLADLGELPDRSLIAFLPVNVRPKGDE-GGGNAVGAI
MSMEG_6322|M.smegmatis_MC2_155 DIVLAMCAGALRAYLAEQDALPDRPLIAMVPVSMRSEHEADAGGNMVGSI
TH_2030|M.thermoresistible__bu DVALAMCAGALRQYLLEHDALPATPLVAMVPVSLRSADESAGGGNRTGLV
MAV_0355|M.avium_104 DVVLAMCSGALRYYLLEQNALPDTPLIAMVPVSLRTEEEADAGGNLVGAI
MUL_4351|M.ulcerans_Agy99 DVVLAMCAGALRCYLEDNDALPDAPLVAMVRVSLRTERDS-VGGNMVGAV
MAB_4544c|M.abscessus_ATCC_199 DVVLAMCGGALRTYLEEQNALPDKPLVAMCPVSIRAEGDQNAGNSIT-AL
MLBr_01244|M.leprae_Br4923 DAYLAGLVGALRRYHEALGVSIS-TLPMAVPVNVRTEADVVGSNRFVGVT
* :* *.** : . .* *.:*. : .. .
Rv3480c|M.tuberculosis_H37Rv LAT-LGTDVADPVQRLAAVTASTRAAKAQLRSMDKDAILAYSAALMAPYG
Mb3509c|M.bovis_AF2122/97 LAT-LGTDVADPVQRLAAVTASTRAAKAQLRSMDKDAILAYSAALMAPYG
MMAR_4955|M.marinum_M LAT-LGTDIADPVERLRAVTASTRMAKAQLKNMDRDAILAYSAALMVPYG
Mflv_1058|M.gilvum_PYR-GCK LAP-MGTDIEDPVERLDTITTATRAAKGQLQSMSPAAIIAYSAALLAPAG
Mvan_5773|M.vanbaalenii_PYR-1 LAP-MGTDIEDPVERLAVITAATRASKAQLQSMSPAAIIAYSAALLAPAG
MSMEG_6322|M.smegmatis_MC2_155 LCS-LGTDVEDPADRLAVIRRSITDNKRVFSELPRLQALALSALLIAP--
TH_2030|M.thermoresistible__bu LCN-LATDSDDPAERLDRISSSMRSNKRVFSQLPRVQAMALSAAMIAP--
MAV_0355|M.avium_104 LCN-LATDSDDPAQRLLTISESMRSNKKVFSQLPRFQALALSAANLSS--
MUL_4351|M.ulcerans_Agy99 LCN-LATHLDDPAQRLAVIHASMRDNKKVLAQLPRAQALALSLLLLSP--
MAB_4544c|M.abscessus_ATCC_199 LAN-LATDKPDPLERFSAIRDSVQAAKSVLSEMTPLQRMLVGALNGAP--
MLBr_01244|M.leprae_Br4923 LAAPLGTN--DPAARMQKIRSQMTQRR----DEPAMNIIGS----LAPLM
*. :.*. ** *: : : . : .
Rv3480c|M.tuberculosis_H37Rv VQLASTLSGVKPPWPYTFNLCVSNVPGPEDVLYLRGSRMEASYPVS----
Mb3509c|M.bovis_AF2122/97 VQLASTLSGVKPPWPYTFNLCVSNVPGPEDVLYLRGSRMEASYPVS----
MMAR_4955|M.marinum_M IQLASTLTGVKPPLPYTFNLCVSNVPGPQEVLYLRGSRMEASYPVS----
Mflv_1058|M.gilvum_PYR-GCK SQIAGALTGVQPPWPYTFNLCVSNVPGPREPLYFNGSRLEATYPVS----
Mvan_5773|M.vanbaalenii_PYR-1 GQIAGALTGVNPPWPHTFNLCVSNVPGPREPLYFNGSRLEATFPVS----
MSMEG_6322|M.smegmatis_MC2_155 LGLTTAFPGFVDATAPPFNLVISNVPGPKKPLYWRGARLAGNYPLS----
TH_2030|M.thermoresistible__bu LGLG-AVPGVVSSAPPPFNLVISNVPGAPEPRYWMGARLAGNYPLS----
MAV_0355|M.avium_104 LALA-AVPGWVSATSPPFNIVISNVPGPTAPLYYGGARLDGNYPLS----
MUL_4351|M.ulcerans_Agy99 AALN-TLPGLSTVTPPPFNVCISNVPGVREPRYFNGARLVGNYPLS----
MAB_4544c|M.abscessus_ATCC_199 VLTG-AVPGLVDRTAPAFNVIISNVPGPRTDMFWNGFEMDGCYPAS----
MLBr_01244|M.leprae_Br4923 TVLPASVLDFIVDSVASSDVNASNIPAYPGDTYFAGAKILRQYGIGPRPG
:. . . :: **:*. : * .: : .
Rv3480c|M.tuberculosis_H37Rv ------LVAHSQALNVTLQSYAGTLNFGFIGCRDTLPHLQRLAVYTGEAL
Mb3509c|M.bovis_AF2122/97 ------LVAHSQALNVTLQSYAGTLNFGFIGCRDTLPHLQRLAVYTGEAL
MMAR_4955|M.marinum_M ------LVAHSQALNVTLQSYAGTLNFGFIGCRDTLPHLQRLAVYTGEAL
Mflv_1058|M.gilvum_PYR-GCK ------IPIHGMALNITLQSYADTMNLGFVGCRDRLPHLQRLAVYTGDAL
Mvan_5773|M.vanbaalenii_PYR-1 ------IPIHGMALNITLQSYADTMNLGFVGCRDRLPRLQRLAVYTGDAL
MSMEG_6322|M.smegmatis_MC2_155 ------IALDGQALNMTVVSNAHNLDFGLVGCRRSVPHLQRLLGHLETSL
TH_2030|M.thermoresistible__bu ------IVLDGQALNITLVTNADNLDFGLVGARGSVPHLQRLLGHLEDSL
MAV_0355|M.avium_104 ------IVLDGQALNITLASNAGNLDFGLVGCRRSVPHLQRLLMHLESSL
MUL_4351|M.ulcerans_Agy99 ------LPINGQALNITLTSTADRLDFGLVGCRSSVPHLQRMLGHLETSL
MAB_4544c|M.abscessus_ATCC_199 ------IPIDGVALNITCTSSGSSMHFGLTGCRTSVPHLQRILAHLEDAL
MLBr_01244|M.leprae_Br4923 VAMMAVLMSRGGFCTVTVRYDRASVKSEALFARCLLEGFDEVLALAGDPT
: . .:* :. .* : ::.: .
Rv3480c|M.tuberculosis_H37Rv DQLAAADGAAGLGS---------
Mb3509c|M.bovis_AF2122/97 DQLAAADGAAGLGS---------
MMAR_4955|M.marinum_M DELDEPAAASAVN----------
Mflv_1058|M.gilvum_PYR-GCK AELEAASQI--------------
Mvan_5773|M.vanbaalenii_PYR-1 GELEAASS---------------
MSMEG_6322|M.smegmatis_MC2_155 KDLETATGA--------------
TH_2030|M.thermoresistible__bu QQLERAVGG--------------
MAV_0355|M.avium_104 KDLERAVGE--------------
MUL_4351|M.ulcerans_Agy99 KELERAVGL--------------
MAB_4544c|M.abscessus_ATCC_199 TQLEESVA---------------
MLBr_01244|M.leprae_Br4923 PHAVPASFAARSSGSPAGWLSSS
. .