For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
SSAKPISAGPVSGNVCLMERRVLEAVIYEREPPIARIVLNRVDKANTKDAVLVTEVDDCLHEADRDKDIK VVIIKANGKGFCGGHVARWGPDENPYPDFGETFEDLYKGTADLFLWPTLYLWEFPKPTISQIHGYCMGGG IYLGLLTDFCVASEDAYFQMPLAQSLGEPGGHTMIEPWLMMNWHRTMDWLLLAPTLSAQQALDWGLLNKV VARDDLEDTVEEMARKIAQIPLTTLMAVKNNVKRAWELMGMRVHLQVSHILTNMVGAASDVQARRAELMR SGMKPREFVDKPDEKGSPPT*
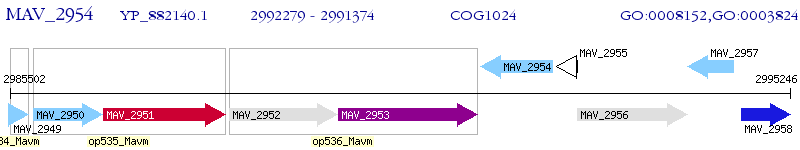
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. avium 104 | MAV_2954 | - | - | 100% (301) | enoyl-CoA hydratase/isomerase family protein |
| M. avium 104 | MAV_1848 | - | 9e-75 | 52.22% (270) | enoyl-CoA hydratase/isomerase |
| M. avium 104 | MAV_2950 | - | 4e-72 | 50.94% (267) | enoyl-CoA hydratase/isomerase family protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0464c | echA2 | 2e-24 | 27.72% (303) | enoyl-CoA hydratase |
| M. gilvum PYR-GCK | Mflv_2433 | - | 2e-24 | 32.33% (232) | enoyl-CoA hydratase |
| M. tuberculosis H37Rv | Rv0456c | echA2 | 2e-24 | 27.72% (303) | enoyl-CoA hydratase |
| M. leprae Br4923 | MLBr_02402 | - | 1e-13 | 25.33% (225) | enoyl-CoA hydratase |
| M. abscessus ATCC 19977 | MAB_3859c | - | 1e-19 | 27.27% (242) | enoyl-CoA hydratase |
| M. marinum M | MMAR_0778 | echA2 | 2e-23 | 30.77% (260) | enoyl-CoA hydratase EchA2 |
| M. smegmatis MC2 155 | MSMEG_4846 | - | 9e-25 | 33.63% (226) | enoyl-CoA hydratase |
| M. thermoresistible (build 8) | TH_3498 | - | 7e-74 | 50.55% (275) | PUTATIVE enoyl-CoA hydratase/isomerase |
| M. ulcerans Agy99 | MUL_1414 | echA2 | 1e-23 | 30.77% (260) | enoyl-CoA hydratase |
| M. vanbaalenii PYR-1 | Mvan_5190 | - | 6e-26 | 33.20% (244) | enoyl-CoA hydratase |
CLUSTAL 2.0.9 multiple sequence alignment
MAV_2954|M.avium_104 MSSAKPISAGPVSGNVCLMERRVLEAVIYEREP--PIARIVLNRVDKANT
TH_3498|M.thermoresistible__bu ------------------LETRTLEHVIYEKDG--PIARIILNNPDRANA
Mflv_2433|M.gilvum_PYR-GCK ------------------------MYIDYEVSD--RIATITLNRPEAANA
MSMEG_4846|M.smegmatis_MC2_155 ------------------------MYIDYGVAD--SIATITLNRPEAANA
Mvan_5190|M.vanbaalenii_PYR-1 -----------------MQRVATLEQVTYETLDEGRIARIWLNRPDAQNA
Mb0464c|M.bovis_AF2122/97 ------------------MPTPDFQTLLYTTAG--PVATITLNRPEQLNT
Rv0456c|M.tuberculosis_H37Rv ------------------MPTPDFQTLLYTTAG--PVATITLNRPEQLNT
MMAR_0778|M.marinum_M ------------------MPTPEFQTLLYSTAG--PVATITLNRPEHLNT
MUL_1414|M.ulcerans_Agy99 ------------------MPTPEFQTLLYSTAG--PVATITLNRPEHLNT
MAB_3859c|M.abscessus_ATCC_199 -------------------MVDTLKTMTYEVTD--RVARITFNRPEQGNA
MLBr_02402|M.leprae_Br4923 -----------------------MKYDTILVDGYQRVGIITLNRPQALNA
:. * :*. : *:
MAV_2954|M.avium_104 KDAVLVTEVDDCLHEADRDKDIKVVIIKANGKGFCGGHVARW---GPDEN
TH_3498|M.thermoresistible__bu QTSEMVHSVNTALDDAQYDYDIKVVIIKANGKGFCSGHIP--------DG
Mflv_2433|M.gilvum_PYR-GCK QNPELLDELDAAWTRAAEDNDVSVIVLRANGKHFSAGHDLR------GGG
MSMEG_4846|M.smegmatis_MC2_155 QNPELLDELDAAWTRAAEDNEVKVIILRANGKHFSAGHDLR------GGG
Mvan_5190|M.vanbaalenii_PYR-1 QSRTLLVQLDEAFGRAEADDEVRVVILAARGKNFSAGHDLG------SEE
Mb0464c|M.bovis_AF2122/97 IVPPMPDEIEAAIGLAERDQDIKVIVLRGAGRAFSGGYDFG--------G
Rv0456c|M.tuberculosis_H37Rv IVPPMPDEIEAAIGLAERDQDIKVIVLRGAGRAFSGGYDFG--------G
MMAR_0778|M.marinum_M IVPPMPDEIEAAVGLAERDQGIKVIVLRGAGRAFSGGYDFG--------G
MUL_1414|M.ulcerans_Agy99 IVPPMPDEIEAAVGLAERDQGIKVIVLRGAGRAFSGGYDFG--------G
MAB_3859c|M.abscessus_ATCC_199 IIADTPLELAALVERADLDPQVHVILVSGRGDGFCGGFDLSAYAEANDEG
MLBr_02402|M.leprae_Br4923 LNSQMMNEITNAAKELDIDPDVGAILITGSPKVFAAGADIK---------
.: * : .::: . *..*
MAV_2954|M.avium_104 PYPDFGETFED--------------------LYKGTADLFLWPTLYLWEF
TH_3498|M.thermoresistible__bu SYPEFKAEIEAS-----------------GKVWRSAAQLFLWPVLKLWEF
Mflv_2433|M.gilvum_PYR-GCK PVPDKIT---------------------LEFIIQHEAKRYLEYTLRWRNV
MSMEG_4846|M.smegmatis_MC2_155 EVPEKIS---------------------LEFIIQHEARRYLDYTLRWRNV
Mvan_5190|M.vanbaalenii_PYR-1 AIAERTPGPDQHPTF---TFNGGTRDAISERTYLQEWHYFFNNTSRWRDL
Mb0464c|M.bovis_AF2122/97 GFQHWGDAM---------MTDGRWDPGKDFAMVTARETGPTQKFMAIWRA
Rv0456c|M.tuberculosis_H37Rv GFQHWGDAM---------MTDGRWDPGKDFAMVTARETGPTQKFMAIWRA
MMAR_0778|M.marinum_M GFQHWGETM---------MTDGQWDPGKDFAMVTARETAPTQKFMAIWRA
MUL_1414|M.ulcerans_Agy99 GFQHWGETM---------MTDGQWDPGKDFAMVTARETAPTQKFMAIWRA
MAB_3859c|M.abscessus_ATCC_199 SVRYSGTALDGQVQLRNHLPNINWDPMLDYQMMSR----FTRGFSSLLHA
MLBr_02402|M.leprae_Br4923 ---EMASLT----------------------FTDAFDADFFSAWGKLAAV
MAV_2954|M.avium_104 PKPTISQIHGYCMGGGIYLGLLTDFCVASEDAYFQMPLAQSLGEPGGHTM
TH_3498|M.thermoresistible__bu PKPVIAQVHGYAIGGGTTWALIPEITVCSEDAWFQMPLVPGFGLPGSETM
Mflv_2433|M.gilvum_PYR-GCK PKPSIAAVQGRCISGGLLLCWPCDLIVAADDAQFSDPVVL-MGIGGVEYH
MSMEG_4846|M.smegmatis_MC2_155 PKPSIAAVQGRCISGGLLLCWPCDLILASDDALFSDPVAL-MGIGGVEYH
Mvan_5190|M.vanbaalenii_PYR-1 RKITIAQVQGNAISAGLMLIWACDLIVASDDARFSDVVAVRMGMPGVEYY
Mb0464c|M.bovis_AF2122/97 SKPVIAQVHGWCVGGASDYALCADIVIASEDAVIGTPYSRMWGAYLTGMW
Rv0456c|M.tuberculosis_H37Rv SKPVIAQVHGWCVGGASDYALCADIVIASEDAVIGTPYSRMWGAYLTGMW
MMAR_0778|M.marinum_M SKPVIAQIHGWCVGGASDYALCADIVIASDDAVIGTPYSRMWGAYLSGMW
MUL_1414|M.ulcerans_Agy99 SKPVIAQIHGWCVGGASDYALCADIVIASDDAVIGTPYSRMWGAYLSGMW
MAB_3859c|M.abscessus_ATCC_199 NKPTVVKVHGYCVAGGTDIALHADQLIVAADAKIGYPPTRVWGVPATGLW
MLBr_02402|M.leprae_Br4923 RTPMIAAVAGYALGGGCELAMMCDLLIAADTAKFGQPEIKLGVLPGMGGS
. : : * .:... : : : * :
MAV_2954|M.avium_104 IEPWLMMNWHRTMDWLLLAPTLSAQQALDWGLLNKVVARDDLEDTVEEMA
TH_3498|M.thermoresistible__bu FEPWVFMNYKRAAEYLYTAQKITAQQALEFGLVNRVVPADELESTVEELA
Mflv_2433|M.gilvum_PYR-GCK GHTWELG-PRKAKEILFTGRAMTADEVAATGMVNKVVPRAELDAETRALA
MSMEG_4846|M.smegmatis_MC2_155 GHTWELG-PRKAKEILFTGRALTAEEAERTGMVNRVVARDELDAQTRELA
Mvan_5190|M.vanbaalenii_PYR-1 AHPWEFG-ARKAKELLLTGDAIDADEAHRLGMVSKVFPRAELEDKTLEFA
Mb0464c|M.bovis_AF2122/97 LYRLSLA---KVKWHSLTGRPLTGVQAAEAELINEAVPFERLEARVAEIA
Rv0456c|M.tuberculosis_H37Rv LYRLSLA---KVKWHSLTGRPLTGVQAAEAELINEAVPFERLEARVAEIA
MMAR_0778|M.marinum_M LYRLSFA---KAKWHSLTGRPLSGVQAAQIELINEAVPFERLEARVAEVA
MUL_1414|M.ulcerans_Agy99 LYRLSFA---KAKWHSLTGRPLSGVQAAQIELINEAVPFERLEARVAEVA
MAB_3859c|M.abscessus_ATCC_199 AHKLGDQ---RAKRLLLTGDCLSGRQAYDWGLAVEAPEPDELDERTERLV
MLBr_02402|M.leprae_Br4923 QRLTRAIGKAKAMDLILTGRTIDAAEAERSGLVSRVVLADDLLPEAKAVA
:. . : . :. : .. * . ..
MAV_2954|M.avium_104 RKIAQIPLTTLMAVKNNVKRAWELMGMRVHLQVSHILTNMVGA-------
TH_3498|M.thermoresistible__bu AKIAKAPLITLQATKAGILRAWETMGFRTHQQASNDLQAVVTG-------
Mflv_2433|M.gilvum_PYR-GCK EQIAKMPPFALRQAKRAVNQTLDVQGFYAAIQSVFDVHQTGHG-------
MSMEG_4846|M.smegmatis_MC2_155 EQIATMPPFALRQAKRAVNQTLDVQGFYAAIQSVFDIHQTGHG-------
Mvan_5190|M.vanbaalenii_PYR-1 RRIAERPTMAALLVKDSVNAASDAMGFTEALRHAFHIHELGHAH--WAAY
Mb0464c|M.bovis_AF2122/97 TELARIPLSQLQAQKLIVNQAYENMGLASTQLLGGILDGLMRNTPDALEF
Rv0456c|M.tuberculosis_H37Rv TELARIPLSQLQAQKLIVNQAYENMGLASTQLLGGILDGLMRNTPDALEF
MMAR_0778|M.marinum_M AELAQIPLSQLQAQKLIVNQAYENMGLASTQTLGGILDGLMRNTPDALDF
MUL_1414|M.ulcerans_Agy99 AELAQIPLSQLQAQKLIVNQAYENMGLASTQTLGGILDGLMRNTPDALDF
MAB_3859c|M.abscessus_ATCC_199 ERIAAMPLNQLMMIKLACNSVLINQGIANSAMIGTVFDGISRHTPEAHAF
MLBr_02402|M.leprae_Br4923 TTISQMSRSATRMAKEAVNRSFESTLAEGLLHERRLFHSTFVT-------
:: . * .
MAV_2954|M.avium_104 --ASDVQARRAELMRSGMKPREFVDKPDEKGSPPT---------
TH_3498|M.thermoresistible__bu --SREFQEYMAELMKRASKPADRV--------------------
Mflv_2433|M.gilvum_PYR-GCK --NALSVGGWPVLVNLEEMKANIK--------------------
MSMEG_4846|M.smegmatis_MC2_155 --NALSVSGWPVLVDIEEMKANIK--------------------
Mvan_5190|M.vanbaalenii_PYR-1 NENRYPVGLPPNVPDWRTLGAPKLSRRDQP--------------
Mb0464c|M.bovis_AF2122/97 IRTAQTQGVRAAVERRDGPFGDYSQAPPELRPDPTHVITPDGSM
Rv0456c|M.tuberculosis_H37Rv IRTAQTQGVRAAVERRDGPFGDYSQAPPELRPDPTHVITPDGSM
MMAR_0778|M.marinum_M IKTAENQGVRAAVERRDGPFGDYSQAPPELRPNPNHVIVPDRDR
MUL_1414|M.ulcerans_Agy99 IKTAENQGVRAAVERRDGPFGDYSQAPPELRPNPNHVIVPDRDR
MAB_3859c|M.abscessus_ATCC_199 TADAIANGYRQAVRHRDEPFGDYGRKPTEL--------------
MLBr_02402|M.leprae_Br4923 --DDQSEGMAAFIEKRAPQFTHR---------------------
: