For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
NFSVLPPEINSALIFAGAGPEPMAAAATAWDGLAMELASAAASFGSVTSGLVGGAWQGASSSAMAAAAAP YAAWLAAAAVQAEQTAAQAAAMIAEFEAVKTAVVQPMLVAANRADLVSLVMSNLFGQNAPAIAAIEATYE QMWAADVSAMSAYHAGASAIASALSPFSKPLQNLAGLPAWLASGAPAAAMTAAAGIPALAGGPTAINLGI ANVGGGNVGNANNGLANIGNANLGNYNFGSGNFGNSNIGSASLGNNNIGFGNLGSNNVGVGNLGNLNTGF ANTGLGNFGFGNTGNNNIGIGLTGNNQIGIGGLNSGTGNFGLFNSGSGNVGFFNSGNGNFGIGNSGNFNT GGWNSGHGNTGFFNAGSFNTGMLDVGNANTGSLNTGSYNMGDFNPGSSNTGTFNTGNANTGFLNAGNINT GVFNIGHMNNGLFNTGDMNNGVFYRGVGQGSLQFSITTPDLTLPPLQIPGISVPAFSLPAITLPSLNIPA ATTPANITVGAFSLPGLTLPSLNIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVGAFSLPGLT LPSLNIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVS GFQLPPLSIPSVAIPPVTVPPITVGAFNLPPLQIPEVTIPQLTIPAGITIGGFSLPAIHTQPITVGQIGV GQFGLPSIGWDVFLSTPRITVPAFGIPFTLQFQTNVPALQPPGGGLSTFTNGALIFGEFDLPQLVVHPYT LTGPIVIGSFFLPAFNIPGIDVPAINVDGFTLPQITTPAITTPEFAIPPIGVGGFTLPQITTQEIITPEL TINSIGVGGFTLPQITTPPITTPPLTIDPINLTGFTLPQITTPPITTPPLTIDPINLTGFTLPQITTPPI TTPPLTIEPIGVGGFTTPPLTVPGIHLPSTTIGAFAIPGGPGYFNSSTAPSSGFFNSGAGGNSGFGNNGS GLSGWFNTNPAGLLGGSGYQNFGGLSSGFSNLGSGVSGFANRGILPFSVASVVSGFANIGTNLAGFFQGT TS*
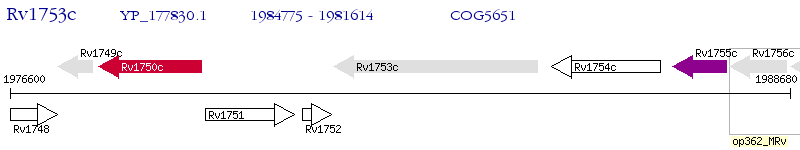
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. tuberculosis H37Rv | Rv1753c | PPE24 | - | 100% (1053) | PPE FAMILY PROTEIN |
| M. tuberculosis H37Rv | Rv1918c | PPE35 | 0.0 | 57.33% (1092) | PPE family protein |
| M. tuberculosis H37Rv | Rv0355c | PPE8 | e-173 | 39.09% (1100) | PPE family protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1782c | PPE24 | 0.0 | 83.33% (1068) | PPE family protein |
| M. gilvum PYR-GCK | Mflv_0325 | - | 7e-26 | 35.29% (204) | PPE protein |
| M. leprae Br4923 | MLBr_01182 | - | 2e-29 | 39.41% (170) | PPE-family protein |
| M. abscessus ATCC 19977 | MAB_0669 | - | 2e-26 | 37.91% (182) | PPE-family protein |
| M. marinum M | MMAR_2822 | - | 0.0 | 64.38% (1064) | PPE family protein |
| M. avium 104 | MAV_4704 | - | 4e-54 | 54.46% (202) | PPE family protein |
| M. smegmatis MC2 155 | MSMEG_0619 | - | 1e-24 | 35.86% (198) | PPE family protein |
| M. thermoresistible (build 8) | TH_3569 | - | 3e-26 | 37.09% (213) | PUTATIVE PPE protein |
| M. ulcerans Agy99 | MUL_0782 | - | 0.0 | 70.70% (488) | PPE family protein |
| M. vanbaalenii PYR-1 | Mvan_0415 | - | 6e-26 | 36.71% (207) | PPE protein |
CLUSTAL 2.0.9 multiple sequence alignment
MSMEG_0619|M.smegmatis_MC2_155 ----------MTAPIWMALPPEVHSSLLSSGPGPGSLLAAAGAWQSLSAE
TH_3569|M.thermoresistible__bu ---------MMTAPIWMALPPEVHSALLSSGPGPGSLLAAASAWHALSAE
Mflv_0325|M.gilvum_PYR-GCK MTSPVASLAPVTSPVWMALPPEVHSTMLSSGPGPGSLLAAGVAWQSLSAE
Mvan_0415|M.vanbaalenii_PYR-1 ----------MSSPVWMALPPEVHSALLSSGPGPGSMLAAAATWQSLSAE
MAB_0669|M.abscessus_ATCC_1997 ----------MTAPIWMAEPPEVHSALLSAGPGPGAVLAAAVQWQEIGNN
Rv1753c|M.tuberculosis_H37Rv ----------MN---FSVLPPEINSALIFAGAGPEPMAAAATAWDGLAME
Mb1782c|M.bovis_AF2122/97 ----------MN---FSVLPPEINSALIFAGAGPEPMAAAATAWDGLAME
MUL_0782|M.ulcerans_Agy99 ----------MN---FSVLPPEVNSALIFTGPGAEPMRTAATAWGGMAVE
MMAR_2822|M.marinum_M ----------MN---FSVLPPEINSALIFAGAGSEPMLTAAAAWDGLADE
MAV_4704|M.avium_104 ----------MTNPHFAWLPPEVNSALIYSGPGPGPLLAAAAAWDGLAEE
MLBr_01182|M.leprae_Br4923 ------------MFDFAALSPETNSTRMYLGPGSSPILTAAAAWVVLAKE
: .** :*: : *.*. .: :*. * :. :
MSMEG_0619|M.smegmatis_MC2_155 YAAAAAELTSVLSAVQAGSWEGPSSEQYVAAHAPYLQWLAQQSANSAAAA
TH_3569|M.thermoresistible__bu YASAAAELSSVLANVQAGSWEGPSAERYVAAHAPYLAWLTQQSANSAATA
Mflv_0325|M.gilvum_PYR-GCK YASAAAELTSILAGVQSGAWEGPSSQQYVAAHTPYLAWLAQQSAVSAATA
Mvan_0415|M.vanbaalenii_PYR-1 YAAAAAELSSILADVQAGAWEGPSSEQYVAAHTPYLAWLAQQSAAGAAAA
MAB_0669|M.abscessus_ATCC_1997 YANTAVELAQVLAGVRAGSWQGPSASEYVAAHLPYLAWLEQAAAESSVIA
Rv1753c|M.tuberculosis_H37Rv LASAAASFGSVTSGLVGGAWQGASSSAMAAAAAPYAAWLAAAAVQAEQTA
Mb1782c|M.bovis_AF2122/97 LASAAASFGSVTSGLVGGAWQGASSSAMAAAAAPYAAWLAAAAVQAEQTA
MUL_0782|M.ulcerans_Agy99 LESAADSFASVTSGLVGGAWQGASSVAMGTAAASYLGWLNAAAAHAEQAA
MMAR_2822|M.marinum_M LSMAALSFESITSGILGGSWQGAASLAMAGAAAPYAGWLATAAGQAAQAG
MAV_4704|M.avium_104 LASSAQSFSSVTSDLASGSWQGASSAAMMTVANQYVSWLSAAAAQAEEVS
MLBr_01182|M.leprae_Br4923 LTAAAQGLQSAVEALLT-TFEGESAAALAERVTPYEKWLTQNAASAELTA
:* : . : :::* :: * ** : . .
MSMEG_0619|M.smegmatis_MC2_155 VQHETAAAAYSTALATMPTMAELALNHTMHGVLVATNFFGINTIPIALNE
TH_3569|M.thermoresistible__bu GRHETAAAAYSTALATMPTMAELTLNKTTHAVLVATNFLGINTIPIALNE
Mflv_0325|M.gilvum_PYR-GCK VQHDTAAAAYTTALATMPTLPELALNHTTHAVLVGTNFLGINTIPIALNE
Mvan_0415|M.vanbaalenii_PYR-1 AQHEAAAAAYSTALATMPTLPELALNHTTHAVLVATNFLGINTIPIAMTE
MAB_0669|M.abscessus_ATCC_1997 AQHETTAAAYNSALAAMPTLAELAANHITHGVLVGTNFFGINTIPIALNE
Rv1753c|M.tuberculosis_H37Rv AQAAAMIAEFEAVKTAVVQPMLVAANRADLVSLVMSNLFGQNAPAIAAIE
Mb1782c|M.bovis_AF2122/97 AQAAAMIAEFEAVKTAVVQPMLVAANRADLVSLVMSNLFGQNAPAIAAIE
MUL_0782|M.ulcerans_Agy99 AQAEALVAEFAAAQSAMVQPFLLAANRSSLVSLVLSNLFGQNAPAIEAVE
MMAR_2822|M.marinum_M AQASVMAAEFEAVRTAMVPPALVAANRSNLVSLVLSNLFGQNAPAIATVE
MAV_4704|M.avium_104 HQASAIATAFEVALAATVQPAVVAANRALVQALAATNWLGQNTPAIADIE
MLBr_01182|M.leprae_Br4923 TQLTVAANAYETAFTMTVPPLMVFVNRAQACLLIMSNIFGQNSTAIAEKE
: . : . : : *: * :* :* *: .* *
MSMEG_0619|M.smegmatis_MC2_155 ADYVRMWIQAATTMSTYQAVSGAAVAAAPVSAPAPFLLKPGV--------
TH_3569|M.thermoresistible__bu ADYVRMWIQAATSMSTYQAVAGTAVMSAPASSPAPLLLAPGV--------
Mflv_0325|M.gilvum_PYR-GCK ADYVRMWIQAATTMATYQSVSGAALAATPTSAPAPSVLLPGV--------
Mvan_0415|M.vanbaalenii_PYR-1 ADYVRMWIQAATTMATYQAVSGAALAATPTATPAPFVLAPGV--------
MAB_0669|M.abscessus_ATCC_1997 ADYVRMWVQAAATMSAYQATTEAMTSAIPPTQQPPAILSPG---------
Rv1753c|M.tuberculosis_H37Rv ATYEQMWAADVSAMSAYHAGASAIASALSPFSKPLQNLAGLP----AWLA
Mb1782c|M.bovis_AF2122/97 ATYEQMWAADVSAMSAYHAGASAIASALSPFSKPLQNLAGLP----AWLA
MUL_0782|M.ulcerans_Agy99 AEYEQMWALDVSTMATYHAGASAAAASLTPLAQPLQNLSELPGRVIAAFG
MMAR_2822|M.marinum_M AAYEEIWALDVSVMSAYHSGASAVVTALTPFA-PLQSLAGLQ----SGLS
MAV_4704|M.avium_104 AAYEQMWASDVAAMFGYHADASAAVAKLPPWNEVLQNLGFSN--------
MLBr_01182|M.leprae_Br4923 AEYTEMWIQDAAAMTSYQASVLEAVGATKAFTAPPLGVNEVG--------
* * .:* .: * *:: :
MSMEG_0619|M.smegmatis_MC2_155 -------GEAGSASASVQQMSAQATAAESGSSLDLSSIISDLLNQYLQGV
TH_3569|M.thermoresistible__bu -------SEAADAMATTQQIGSQATAADSGTALNLSDLIANLLQYYQDYL
Mflv_0325|M.gilvum_PYR-GCK -------GEAGRAAADMSAFAAQPMAAQAGAAFDVSNIIADIIRLYGEVL
Mvan_0415|M.vanbaalenii_PYR-1 -------GEAGRAAADVTAFAAQAQAAEAGSALDLSNIIADLIRAYGELL
MAB_0669|M.abscessus_ATCC_1997 -------GEARSDQSDVPGSPDQ---------------IMKMLQGFQQWF
Rv1753c|M.tuberculosis_H37Rv SGAPAAAMTAAAGIPALAGGPTAINLGIANVGGGNVGNANNGLANIGNAN
Mb1782c|M.bovis_AF2122/97 SGAPAAAMTAAAGIPALAGGPTAINLGIANVGGGNVGNANNGLANIGNAN
MUL_0782|M.ulcerans_Agy99 NGAPATTVTAAVTALRLPADASTVNFGLANIGSGNVGNANNGIGNLGSGN
MMAR_2822|M.marinum_M T-AMSGGASVAAAAPAVPDRVTTLNLGLANVGGGNVGNGNNGFGNFGSGN
MAV_4704|M.avium_104 ---ASTAVTRPAGSGAVARGYTSRIAGFLAPRAPQ---------------
MLBr_01182|M.leprae_Br4923 ----LAQEVVEEVVEEVVEEVVEAEQAISQAALDQAVNEGMEATVVPQVD
MSMEG_0619|M.smegmatis_MC2_155 PGGQ-AILDFLKDPLGNLGRLLTDFMTNPAGALATWWPFLFGVAYQVFFN
TH_3569|M.thermoresistible__bu DQLFGPIVDFLRDPIGNLFQLIDDFLTNPAGALVTWWPLLFAVAYQVFTN
Mflv_0325|M.gilvum_PYR-GCK RFLFEPIFNFLRDPIGNTFALITDFLTNPVQALITWGPFLLAVVYQAVSW
Mvan_0415|M.vanbaalenii_PYR-1 RFLFEPIFDFLRDPLGNTIKLITDFLTNPAQALITWGPFLAAVAYQAVSW
MAB_0669|M.abscessus_ATCC_1997 EKMG--FNPATSAVLAVIMLFLYDLLWYPY-----YASYLLPFLIPALSG
Rv1753c|M.tuberculosis_H37Rv LGNYNFGSGNFGNSNIGSASLGNNNIGFGNLGSNNVGVGNLGNLNTGFAN
Mb1782c|M.bovis_AF2122/97 LGNYNFGSGNFGNSNIGSASLGNNNIGFGNLGSNNVGVGNLGNLNTGFAN
MUL_0782|M.ulcerans_Agy99 LGNFNLGSGNLGSSNAGWANLGSNNIGGANSGGNNLGWGNLGGLNTGFAN
MMAR_2822|M.marinum_M LGSGNFGSGNFGSS----------NIGASNLGSNNFGFGNLGSFNNGFAN
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 QQVN-------------------VDVATPQTAVPDSSSAAAPQLWGGFAQ
MSMEG_0619|M.smegmatis_MC2_155 LVGWPTWGMIFASPFLIPALAALGITGLVLLIDHVLNNPVDVPAEDEAPQ
TH_3569|M.thermoresistible__bu LVGWPTWAMILASPVLIPLAITLGVAGLVMIADQAQP-----PAEEAAPE
Mflv_0325|M.gilvum_PYR-GCK VGASILYPSLFLLPLLATTLAIVAGVGQHLLSLLNTP-PAEEAAEEPAPA
Mvan_0415|M.vanbaalenii_PYR-1 VGASILYPSLLLLPLVATTLAIVLGVGAYLLENLPAP-AEDAPAEEPAAS
MAB_0669|M.abscessus_ATCC_1997 LSG-LAALMHLRPAAPGPVPAAALSPGDRPIRRPTQHVEPNLAALPLPTS
Rv1753c|M.tuberculosis_H37Rv TGLGNFGFGNTGNNNIGIGLTGNNQIGIGGLNSGTGNFGLFNSGSGNVGF
Mb1782c|M.bovis_AF2122/97 TGLGNFGFGNTGNNNIGIGLTGNNQIGIGGLNSGTGNFGLFNSGSGNVGF
MUL_0782|M.ulcerans_Agy99 AGSGNFRFANTGNNNIGIGLTGDNQVGIGGLNSGVGNFGLFNSGTGNVGL
MMAR_2822|M.marinum_M IGAGNFGFGNNGNNNIGIGLNGNNQVGIGLLNSGTGNVGLFNSGDGNIGF
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 HLSPINDTLSMINNHAGMANAG---------------LSLVNGMGSAMKS
MSMEG_0619|M.smegmatis_MC2_155 EVAAAPRTDQPLAVGISPAGVPGSAATAAAPAPAPAGTTVAAPAPAAATF
TH_3569|M.thermoresistible__bu EDVAAP-------------------VRAEPQNPLPLTXXX----------
Mflv_0325|M.gilvum_PYR-GCK QSAPSRPDQQSFPATMSAPPPPSGAPIAANTVSAGSPPAPGAPAAAAATF
Mvan_0415|M.vanbaalenii_PYR-1 SPAPTRADQPSPAIAVSAPPPPSSAAATVGTVATGTAPAPGAPAAATASF
MAB_0669|M.abscessus_ATCC_1997 SVVPSGG---------------SSVPVSTPTAPAHTVSSVPAPSAAISYA
Rv1753c|M.tuberculosis_H37Rv FNSGNGNFGIGNSGNFNTGGWNSGHGNTGFFNAGSFNTGMLDVGNANTGS
Mb1782c|M.bovis_AF2122/97 FNSGNGNFGIGNSGNFNTGGWNSGHGNTGFFNAGSFNTGMLDVGNANTGS
MUL_0782|M.ulcerans_Agy99 FNSGTGNFGIGNSGAYGTGLFNSGQANTGLLNAGSFNTGLLDVGNANTGS
MMAR_2822|M.marinum_M FNSGSGNFGIGNTGNFGTGIFNSGNGSTGLFNAGSFNTGVLNVGDGNTGI
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 LAPTTTKAAESAFKAMGSAVQSTGRGLLGSSSGGHVTAQLG--RAASIGS
MSMEG_0619|M.smegmatis_MC2_155 VPYAVANRDPGEGFTPTLGNKSQVAAPASGVPAAASGVAASLASRRKRRR
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK VPYVVAGRDPYEGFSPTVRDSTGAKAPAGTIPAAASGIAASAAERRKRRR
Mvan_0415|M.vanbaalenii_PYR-1 VPYAVAGRDPGTGFSPTVRDSTSAKAPASGIPAAASGVAASAAERRKRRR
MAB_0669|M.abscessus_ATCC_1997 IPGLVP---PGVSFGPKSTEKSSESAVDTVAAVAAARGAVAAQARRRLRG
Rv1753c|M.tuberculosis_H37Rv LNTGSYNMGDFNPGSSNTGTFNTGNANTGFLNAGNINTGVFNIGHMNNGL
Mb1782c|M.bovis_AF2122/97 LNTGSYNMGDFNPGSSNTGTFNTGNANTGFLNAGNINTGVFNIGHMNNGL
MUL_0782|M.ulcerans_Agy99 LNTGSFNMGSFNPGPSNTGALNTGTANTGWFNTGSLNTGPFNIGDMNNGL
MMAR_2822|M.marinum_M LNTGSYNMGGFNPGSSNTGSFNIGGANTGWLNAGSINTGILNSGDMNNGL
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 LRVPQTWTTASQPVTAATRALSPARVAVATESE----SAPLLGGGLPMAP
MSMEG_0619|M.smegmatis_MC2_155 KKGAQIEERGYADAYMDYEE-PDEDPAPQPEPRVRASERGSDAMGFTGTV
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK RQKEEIKGRGYADAYADYEELPDDGPPEQPAPRVTASTRGAGPMGFAGTV
Mvan_0415|M.vanbaalenii_PYR-1 RQKDEIAGRAYADAYADYEPEPDDEPPVRQEPRIAATERGAGPMGFAGTV
MAB_0669|M.abscessus_ATCC_1997 KNKAEAGGR--RHEYLEETAGMGAT-ATADVPAYAATGRGTGPLGFAGTA
Rv1753c|M.tuberculosis_H37Rv FNTGDMNNGVFYRGVGQGSLQFSITTPDLTLPPLQIPGISVPAFSLPAIT
Mb1782c|M.bovis_AF2122/97 FNTGDMNNGVFYRGVGQGSLQFSITTPDLTLPPLQIPGISVPAFSLPAIT
MUL_0782|M.ulcerans_Agy99 FNTGEMNNGIFYRGVGQGNFYFSITTPDLTLPPLEIPGITIPNLSLPPIT
MMAR_2822|M.marinum_M FNTGDMNNGIFFRGVGQGRLYFGIGLPELTLPPLDVPGITVPGFNLPALT
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 MVPGGGS------GTGGVNTALRLQPRAFVMPRNPAAG------------
MSMEG_0619|M.smegmatis_MC2_155 TRHDVTAAGLTTLTGDTFDDRPVTPMLPGTWDGDENPAPSPDDPEGGPRR
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK SRGDEQAGGLTTLAGDSFGGGPREPMLPGTWDPDATDDRHTDGRDGHDGH
Mvan_0415|M.vanbaalenii_PYR-1 SRDAAQAGGLTTLPGDPFGGGPKAPMLPGTWDPDTEPD------ERHNHH
MAB_0669|M.abscessus_ATCC_1997 PAAASTHVAATVRLS-SDEARQVIPMLPSTWDADGGDQN-----------
Rv1753c|M.tuberculosis_H37Rv LPSLNIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVGAFSLPG
Mb1782c|M.bovis_AF2122/97 LPSLTIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVGAFSLP-
MUL_0782|M.ulcerans_Agy99 MRR-----------------------------------------------
MMAR_2822|M.marinum_M LPSMSLPAITTPANITVGAFDLPGLTLPPLTIPAATTPANITVGAFDLP-
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK DGKEKP--------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 DGKDSQ--------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv LTLPSLNIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVGAFSL
Mb1782c|M.bovis_AF2122/97 --------------------------------------------------
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M --------------------------------------------------
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv PGLTLPSLNIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVSGF
Mb1782c|M.bovis_AF2122/97 -GLTLPSLNIPAATTPANITVGAFSLPGLTLPSLNIPAATTPANITVSGF
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M -GLTLPPLTIPAATTPANITVGAFDLPGLTLPPLTIPAATTPANITVGAF
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv QLPPLSIPSVAIPPVTVPP-ITVGAFNLPPLQIPEVTIPQLTIPAGITIG
Mb1782c|M.bovis_AF2122/97 QLPPLSIPSVAIPPVTVPP-ITVGAFNLPPLQIPEVTIPQLTIPAGITIG
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M DLPGLTLPPLTIPAATTPANITVGAFNLPGLAIPSLTIPSISIPAGTAIG
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv GFSLPAIHTQPITVGQIGVGQFGLPSIGWDVFLSTPRITVPAFGIPFTLQ
Mb1782c|M.bovis_AF2122/97 GFSLPAIHTQPITVGQIGVGQFGLPSIGWDVFLSTPRITVPAFGIPFTLQ
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M SFDLPGITTQSIFVNNITTGSFTTPTIVTPTIAG-PVVTIPGFHLDFKAL
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv FQTNVPALQ-------PPGGGLSTFTNGALIFGEFDLPQLVVHPYTLTGP
Mb1782c|M.bovis_AF2122/97 FQTNVPALQ-------PPGGGLSTFTNGALIFGEFDLPQLVVHPYTLTGP
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M IPNNIIALQTGIAGQYPAIGNFYGMEGAYLETGTLTLGPIGLSGFTIS-P
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv IVIGSFFLPAFNIPGIDVPAINVDGFTLPQITTPAITTPEFAIPPIGVGG
Mb1782c|M.bovis_AF2122/97 IVIGSFFLPAFNIPGIDVPAINVDGFTLPQITTPAITTPEFAIPPIGVGG
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M ISISGFSLPNFTIPGINVPPIHLADFTLPAITTPEFTTPPLSIDPIALDG
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv FTLPQITTQEIITPELTINSIGVG--------------------------
Mb1782c|M.bovis_AF2122/97 FTLPQITTQEIITPELTINSIGVGGFTLPQITTPPITTPPLTIDPINLTG
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M FTLPQISTPQITTEPFTIDPIAVGG-------------------------
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv ------------------------GFTLPQITTPPITTPPLTIDPINLTG
Mb1782c|M.bovis_AF2122/97 FTLPQITTPPITTPPLTIDPINLTGFTLPQITTPPITTPPLTIDPINLTG
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M FTLPQISTPQITTEPFTIDPIAVGGFTLPQISTPQITTEPFTIDPIAVGG
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv FTLPQITTPPITTPPLTIDPINLTGFTLPQITTPPITTPPLTIEPIGVGG
Mb1782c|M.bovis_AF2122/97 FTLPQITTPPITTPPLTIDPINLTGFTLPQITTPPITTPPLTIEPIGVGG
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M FTLPQISTPQITTEPFTIDPIAVGGFTLPQINIPPTTTGELTIPPIGLAS
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv FTTPPLTVPGIHLPSTTIGAFAIPGGPGYFNSSTAPSSGFFNSGAGGNSG
Mb1782c|M.bovis_AF2122/97 FTTPPLTVPGIHLPSTTIGAFAIPGGPGYFNSSTAPSSGFFNSGAGGNSG
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M FTTPALTIPSIHVPAGTLAQVSIPAGSGFLNSSTAPSSGFFNAGAGGSSG
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 --------------------------------------------------
TH_3569|M.thermoresistible__bu --------------------------------------------------
Mflv_0325|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_0415|M.vanbaalenii_PYR-1 --------------------------------------------------
MAB_0669|M.abscessus_ATCC_1997 --------------------------------------------------
Rv1753c|M.tuberculosis_H37Rv FGNNGSGLSGWFNTNPAGLLGGSGYQNFGGLSSGFSNLGSGVSGFANRGI
Mb1782c|M.bovis_AF2122/97 FGNNGSGLSGWFNTNPAGLLGGSGYQNFGGLSSGFSNLGSGVSGFANRGI
MUL_0782|M.ulcerans_Agy99 --------------------------------------------------
MMAR_2822|M.marinum_M FGNNGSGLSGWFNTNATGLIGGSGYQNYGSVFSGFSNLGTGVSGFANTGT
MAV_4704|M.avium_104 --------------------------------------------------
MLBr_01182|M.leprae_Br4923 --------------------------------------------------
MSMEG_0619|M.smegmatis_MC2_155 -----------------------------
TH_3569|M.thermoresistible__bu -----------------------------
Mflv_0325|M.gilvum_PYR-GCK -----------------------------
Mvan_0415|M.vanbaalenii_PYR-1 -----------------------------
MAB_0669|M.abscessus_ATCC_1997 -----------------------------
Rv1753c|M.tuberculosis_H37Rv LPFSVASVVSGFANIGTNLAGFFQGTTS-
Mb1782c|M.bovis_AF2122/97 LPFSVASVVSGFANIGTNLAGFFQGTTS-
MUL_0782|M.ulcerans_Agy99 -----------------------------
MMAR_2822|M.marinum_M LPLAMTSLVSGFANVGSSLSGLFFQSVTP
MAV_4704|M.avium_104 -----------------------------
MLBr_01182|M.leprae_Br4923 -----------------------------