For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MAAVLIAVLSPVGHVEPLFAVAEDLVSRGDQVTVMTGAAHTDSIRAIGAHPHPLPRYADFDDAPFDTGQR AATSGIDALSRAVIRLFLTPMPLQAAELRRALADRHYDAVIVDYLFLGILPLLLDDTVRTPPVLHYTPTP MMLSSRDTAPVGLGLPPARTAARRALYRALTLLSQKVLLRNAQRTADRMLTSMGVPRLPVPLLDAGRLAD RLIVPTVPSFEYPRSDLPEGVRFVGLVRPRSADRFVTPPWWEVLDADRPVVHVTQGTVDNRDLSRLIEPT ITALADTDVTVVVSTGGRTRDSIRVPIPANTHVCEYIPHDRLLPKVDVMVTNGGYGGVQRALAAGVPLVV AGSTEDKPEVAARVAWSGAGINLDTGTPSADAIRSAVALLRSDDRYLRNARRLEAAFARQDGIAEIAALI DELIGVRRTAAT
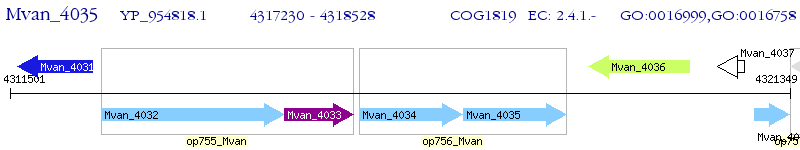
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_4035 | - | - | 100% (432) | glycosyl transferase family protein |
| M. vanbaalenii PYR-1 | Mvan_4034 | - | e-132 | 56.64% (429) | glycosyl transferase family protein |
| M. vanbaalenii PYR-1 | Mvan_4886 | - | 3e-79 | 43.20% (419) | UDP-glucuronosyl/UDP-glucosyltransferase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb2986c | - | 1e-22 | 35.82% (201) | glycosyl transferase |
| M. gilvum PYR-GCK | Mflv_2592 | - | 1e-133 | 58.45% (426) | glycosyl transferase family protein |
| M. tuberculosis H37Rv | Rv2962c | - | 1e-22 | 35.82% (201) | glycosyl transferase |
| M. leprae Br4923 | MLBr_00125 | - | 4e-20 | 31.80% (217) | putative glycosyl transferase |
| M. abscessus ATCC 19977 | MAB_3062c | - | 2e-19 | 28.68% (408) | hypothetical protein MAB_3062c |
| M. marinum M | MMAR_1755 | - | 4e-23 | 31.25% (224) | glycosyl transferase |
| M. avium 104 | MAV_3631 | - | 2e-19 | 29.18% (425) | hypothetical protein MAV_3631 |
| M. smegmatis MC2 155 | MSMEG_4670 | - | 1e-137 | 59.53% (430) | N-glycosylation |
| M. thermoresistible (build 8) | TH_3465 | - | 5e-78 | 40.91% (440) | PUTATIVE UDP-glucuronosyl/UDP-glucosyltransferase |
| M. ulcerans Agy99 | MUL_3383 | - | 3e-16 | 27.29% (414) | glycosyl transferase |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_2592|M.gilvum_PYR-GCK ---------------------------------------MNAMPDILITA
MSMEG_4670|M.smegmatis_MC2_155 ------------------------------------------MADILIAS
Mvan_4035|M.vanbaalenii_PYR-1 ------------------------------------------MAAVLIAV
TH_3465|M.thermoresistible__bu ------------------VGGYGLTRTRLLGRRLSSGSEDEFLARYLLAA
MAV_3631|M.avium_104 -------------------------------------------MRVAVVA
MUL_3383|M.ulcerans_Agy99 -------------------------------------------MRVAVVA
MAB_3062c|M.abscessus_ATCC_199 -------------------------------------------MRVAVVA
Mb2986c|M.bovis_AF2122/97 MRVSCVYATASRWGGPPVASEVRGDAAISTTPDAAPGLAARRRRILFVAE
Rv2962c|M.tuberculosis_H37Rv MRVSCVYATASRWGGPPVASEVRGDAAISTTPDAAPGLAARRRRILFVAE
MMAR_1755|M.marinum_M -------------------------MSIATN---AP-QPVRRQRVLFVAE
MLBr_00125|M.leprae_Br4923 ---------------------MGKDAGMSTAPDTVPECQSRRMRILFIAE
:.
Mflv_2592|M.gilvum_PYR-GCK CPPVGHIGPLLNVAHGLVAHGHRVTVLTSARHADKIRAVGADPRPLPYGA
MSMEG_4670|M.smegmatis_MC2_155 HSPIGHIGPLLNVAYGLVGRGDRVTVLTSAQHTDAIRASGAAPVALPDAG
Mvan_4035|M.vanbaalenii_PYR-1 LSPVGHVEPLFAVAEDLVSRGDQVTVMTGAAHTDSIRAIGAHPHPLPRYA
TH_3465|M.thermoresistible__bu TPLSGHVAPLVQIGATLRSRGHTVTVLTTTEFADAVTRAGLRFTALPAGT
MAV_3631|M.avium_104 GPDPGHSFPAIALCRRFADAGDTPTLFTGAEWLDTARGAGVDAVELDGLA
MUL_3383|M.ulcerans_Agy99 GPDPGHSFPAIALCRRFLAAGDIPTLLTGVEWLDTARAAGVDAIELAGLV
MAB_3062c|M.abscessus_ATCC_199 GPDPGHAFPALALCLKFAAAGDDPVLLTGTRWLDQARAAGVESVELLGLQ
Mb2986c|M.bovis_AF2122/97 AVTLAHVVRPFALAQSLDPSRYEVHFACDPRYNQLLGPLPFRHHAIHTIP
Rv2962c|M.tuberculosis_H37Rv AVTLAHVVRPFALAQSLDPSRYEVHFACDPRYNQLLGPLPFRHHAIHTIP
MMAR_1755|M.marinum_M ATTLAHVVRPFVVALSLDPSRYEVHFACDPRFNKLLGPIPFPHHPIYTIP
MLBr_00125|M.leprae_Br4923 AVTLAHVVRPFAVARLLDPSRYEVHFACDPRYNNLLGPLPFPHHPIHTVP
.* . : : . . :
Mflv_2592|M.gilvum_PYR-GCK D-YDDARLDADLPGRAHTSGIARIN--FDVEHVFVRPMPYQFSAVQELLA
MSMEG_4670|M.smegmatis_MC2_155 D-PDADRLDIENPKRARTSGIARIN--VDIVAFFVAPMAHQADALAKLLA
Mvan_4035|M.vanbaalenii_PYR-1 D-FDDAPFDTG--QRAATSGIDALS--RAVIRLFLTPMPLQAAELRRALA
TH_3465|M.thermoresistible__bu A-IDPPAAPPWWARYLPSQARRFLLGRAELDSVYIRPLGHQAAALRSVLD
MAV_3631|M.avium_104 A--TDEDVDAGARIHRRAARMAVLN----------------VPALRDLAP
MUL_3383|M.ulcerans_Agy99 A--TDEDLDAGARIHRRAAQMALLN----------------APALSDLAP
MAB_3062c|M.abscessus_ATCC_199 PRVDDDDADSGAKIHQRAAHMALAN----------------LDQLRALAP
Mb2986c|M.bovis_AF2122/97 S---ERFFGNLTQGR--FYAMRTLR--------------KYVEADLRVLD
Rv2962c|M.tuberculosis_H37Rv S---ERFFGNLTQGR--FYAMRTLR--------------KYVEADLRVLD
MMAR_1755|M.marinum_M S---ERFLGNLVQGR--FYTTRTLR--------------QYVEEDSRVLA
MLBr_00125|M.leprae_Br4923 S---KRFLDNMAQGRLFLYGPRTLR--------------KYVEEDSKLLS
Mflv_2592|M.gilvum_PYR-GCK ANSFDAVITDAFFLGVIPLLLDPTLARPPIMAYSTTPLFLSSRDTAPAGP
MSMEG_4670|M.smegmatis_MC2_155 QHRFDAVIADYGFFGILPFLLGNRADRPPFLTYSTTPLMLSSRDTAPAGL
Mvan_4035|M.vanbaalenii_PYR-1 DRHYDAVIVDYLFLGILPLLLDDTVRTPPVLHYTPTPMMLSSRDTAPVGL
TH_3465|M.thermoresistible__bu REPADAVLVDMTFTGAAALLLDDRP-RPPLLVCSVGPLTLSSADTPPFGM
MAV_3631|M.avium_104 DLVVSDVITAGGGMAAELLGI-------PWIELSPHPLYLPSKGLPPIGS
MUL_3383|M.ulcerans_Agy99 DLVVSDVITACGGMAAERLGI-------PWIELNPHPLYRPSKGLPPIGS
MAB_3062c|M.abscessus_ATCC_199 DLIVSDVITACGGMAGELLGV-------PWVELNPHPLYLPSKGLPPLGS
Mb2986c|M.bovis_AF2122/97 EIAPDLVVGDLRISLSVSARLAG------IPYIAIANAYWSPYAQRRFPL
Rv2962c|M.tuberculosis_H37Rv EIAPDLVVGDLRISLSVSARLAG------IPYIAIANAYWSPYAQRRFPL
MMAR_1755|M.marinum_M EVAPDLVVGDLRISLSASARLAG------IPYVAIANAYWSPYSRRRFPL
MLBr_00125|M.leprae_Br4923 EIEPDLVVGDLRWSLSVSARLAS------IPYIAIANAYWSSHARRRFPL
. *: : ..
Mflv_2592|M.gilvum_PYR-GCK GILPMPGAVGRLRNRLLGLLSRAVLLRPAHRSAERMLRAMGLPDLPVFVL
MSMEG_4670|M.smegmatis_MC2_155 GLPPPRHAVERLRNRALETLTHKVLLRASQRAVQEQLHRMNCRPLPVFLF
Mvan_4035|M.vanbaalenii_PYR-1 GLPPARTAARRALYRALTLLSQKVLLRNAQRTADRMLTSMGVPRLPVPLL
TH_3465|M.thermoresistible__bu AWQPRP----GVDYRGMTAVAHRVIMAGSRRRFDRALRTAGAGPAPVFLS
MAV_3631|M.avium_104 GLATGTGLRGRLRDATMRALTARSWRAG--------LRQRAAARAEIGLP
MUL_3383|M.ulcerans_Agy99 GLAPGTGIRGRLRDTTMRALTARSWRAG--------LRQRAAVRAEIGLA
MAB_3062c|M.abscessus_ATCC_199 GLAPGVGLRGRLRDGLMRRLTQKSIDQG--------LRQRSQARVEIGLP
Mb2986c|M.bovis_AF2122/97 PDVIWTRLFGVRLVKLLYRLERPLLFALQCMPLNWVRRRHGLSSLGWNLC
Rv2962c|M.tuberculosis_H37Rv PDVIWTRLFGVRLVKLLYRLERPLLFALQCMPLNWVRRRHGLSSLGWNLC
MMAR_1755|M.marinum_M PEVFWSRLFGVRLVKRAYRLERPLFFALQCMPLNRVRRKHGLPSLGLNLC
MLBr_00125|M.leprae_Br4923 PDVLYTHLLGVRLVKLLYRLERPLFFAFQCLPLNWVRRKHGLPSLGFNLC
: :
Mflv_2592|M.gilvum_PYR-GCK ECAILADRVIVPTVPDFEYPRRDLPAHVRFVGAVSPLPGDDFIAPAWWGE
MSMEG_4670|M.smegmatis_MC2_155 DGAVLSERVIAPTVPQFDYHRSDLPKNVRYVGAVHPMPNQGFRRPMWWRL
Mvan_4035|M.vanbaalenii_PYR-1 DAGRLADRLIVPTVPSFEYPRSDLPEGVRFVGLVRPRSADRFVTPPWWEV
TH_3465|M.thermoresistible__bu DWPALADRLIQLGVPGFEYHRGDLPSTVDFAGPVLPTGLDGVTLPAWWDD
MAV_3631|M.avium_104 ARDPGPLRRLIATLPALEVPRPDWPDEAVLVGPLHFEPTDRVLD------
MUL_3383|M.ulcerans_Agy99 EHDPGPLRRLIATLPALEVPRPDWPAEAIVVGPLHFEPTDQLLK------
MAB_3062c|M.abscessus_ATCC_199 ARDPGPMRRLIATLPALEVPRPDWPAEAVVVGPLHWEPTDVVLQ------
Mb2986c|M.bovis_AF2122/97 RIFTDGDHTLYADVPELMP-TYDLPANHEYLGPVLWSPAGKPPT--WWDS
Rv2962c|M.tuberculosis_H37Rv RIFTDGDHTLYADVPELMP-TYDLPANHEYLGPVLWSPAGKPPT--WWDS
MMAR_1755|M.marinum_M RIFTDGDQTLYADIPELQP-TYNRPDNHQYLGPVLWSPSGETPD--WWDS
MLBr_00125|M.leprae_Br4923 RIFTDGDYTLYADVPELVP-TYDLPANHQYLGPVLWSPAGELPR--WWDS
: :* : : * * :
Mflv_2592|M.gilvum_PYR-GCK LYRGRPVVHVTQGTVDNADLTRLIEPTIEALAG-EDVTVVVSTGGRPVSQ
MSMEG_4670|M.smegmatis_MC2_155 LDGDRPVVHVTQGTVDNTDLTRLLEPTLEALAD-EDVVVIASTGGRDLDV
Mvan_4035|M.vanbaalenii_PYR-1 LDADRPVVHVTQGTVDNRDLSRLIEPTITALAD-TDVTVVVSTGGRTRDS
TH_3465|M.thermoresistible__bu IRSAATVVHVTQGTFDNTDLDQLVVPTLEALTDRPDVLVVATTGGRPGQR
MAV_3631|M.avium_104 IPAGSGPVVVVAPSTAQTGARGLAEVALSCLTPGETLPAGARLVVSRLAG
MUL_3383|M.ulcerans_Agy99 VPAGSGPVLVIAPSTALSGTAGLAEVALESLIPGGTLSSGSRLVVSRLGG
MAB_3062c|M.abscessus_ATCC_199 PPSGSGPVVVIAPSTAFTGAEGMTELAVQALES-----AGARVVISALEP
Mb2986c|M.bovis_AF2122/97 LPTDRPIVYATLGTS---GGRNLLQLVLNALAELPVTVIAATAGRSDLKT
Rv2962c|M.tuberculosis_H37Rv LPTDRPIVYATLGTS---GGRNLLQLVLNALAELPVTVIAATAGRSDLKT
MMAR_1755|M.marinum_M LPADRPIVYATLGTS---GGRNLLQVVLNGLADLPVTVIAATADRSDLDD
MLBr_00125|M.leprae_Br4923 LPTDRPIVYATLGTS---GGKNLLQVVLDALADLPVTVIAATAGRSELQN
* . : . : .: *
Mflv_2592|M.gilvum_PYR-GCK IRTPLPANTFVAEFLPHDVLLPLVDVLVTNGGYGAVQRALAEGVPLVVAG
MSMEG_4670|M.smegmatis_MC2_155 LRTPVPANTYVAKYIPHDLLMPKVDVLVTNGGYGTVQRALSCGVPLVVAG
Mvan_4035|M.vanbaalenii_PYR-1 IRVPIPANTHVCEYIPHDRLLPKVDVMVTNGGYGGVQRALAAGVPLVVAG
TH_3465|M.thermoresistible__bu SGFRVPGNARVTGWLPYRMLMPHVDVMITNGGYGGVQYALSHGVPLIVAG
MAV_3631|M.avium_104 PELAAPP-WAVVGLGSQAELLRHADVVVCGGGHGMVAKTLLAGVPLVAVP
MUL_3383|M.ulcerans_Agy99 DDLVTPS-WAAVGLGSQAELLTHADVVICGGGHGMVSKTLLAGVPLVVVP
MAB_3062c|M.abscessus_ATCC_199 PSSGLPS-WAIAGLARQDLLLSGADLAVCGGGHGFLSKALLAGVPTVVVP
Mb2986c|M.bovis_AF2122/97 ----VPANAFVADYLPGEAAAARSAVVVCNGGSLTTQQALVAGVPVIGVA
Rv2962c|M.tuberculosis_H37Rv ----VPANAFVADYLPGEAAAARSAVVVCNGGSLTTQQALVAGVPVIGVA
MMAR_1755|M.marinum_M ----IPANAYVADYLPGEAAAARSDVVICNGGSLTTQQALVAGVPVLGIA
MLBr_00125|M.leprae_Br4923 ----VPANAFVADYLPGEAAAARSAVVICNGGSLTTQQAFVAGVPVVGIA
* : : .** :: *** :
Mflv_2592|M.gilvum_PYR-GCK DTEDKPEVAARVEYFGVGVNLRTGTPAPQDLRDAVNAVLSDAGYRTRAGR
MSMEG_4670|M.smegmatis_MC2_155 KTEDKPEVAARVAWSGAGIDLRTGTPTPAAIRDAVREVLTDKRYLSRART
Mvan_4035|M.vanbaalenii_PYR-1 STEDKPEVAARVAWSGAGINLDTGTPSADAIRSAVALLRSDDRYLRNARR
TH_3465|M.thermoresistible__bu ETSDKAEIAARVEFCGAGIDLGTATPTVRALRAGFDRIRDQDRYRTAARR
MAV_3631|M.avium_104 GGGDQWEIANRVVRQGSARLIRP--LSADALVAAVNEVLSSPGYRAAAQR
MUL_3383|M.ulcerans_Agy99 GGGDQWEIANRVVRQGSARLIRP--LGADALAAAVNEVLSSPGYREAAQR
MAB_3062c|M.abscessus_ATCC_199 GGGDQWELANRVVRQGSAELVRP--LSRDSLASAIGRVLGEPRYREAARR
Mb2986c|M.bovis_AF2122/97 GNLDQHLNMEAVERAGAGVLLRTERLKSQRVAGAVMQVISRSEYRQAAAR
Rv2962c|M.tuberculosis_H37Rv GNLDQHLNMEAVERAGAGVLLRTERLKSQRVAGAVMQVISRSEYRQAAAR
MMAR_1755|M.marinum_M SNLDQHLNMQAIEAAGAGLLLRTERLKARSVSAAVCQLLDRPGYRQSAEE
MLBr_00125|M.leprae_Br4923 GNLDQHLNMEAVEAAGAGILLRSERLKVRRVADAVNRVLGQSEYRQAAQR
*: : * . : . : .. : * *
Mflv_2592|M.gilvum_PYR-GCK LQAAYASRDSVTAIAALLDEVIPEQAAAR---------------------
MSMEG_4670|M.smegmatis_MC2_155 LEAAFAQRDGVAEIIALVDEVIAERRAEVGGELACEVTHEQR--------
Mvan_4035|M.vanbaalenii_PYR-1 LEAAFARQDGIAEIAALIDELIGVRRTAAT--------------------
TH_3465|M.thermoresistible__bu LAREIRSAAPLDSIAATLERIVAADALNRSARTGSAGAPDPDSTDDRPAP
MAV_3631|M.avium_104 AAAGIAD---VADPVRVCREALAG--------------------------
MUL_3383|M.ulcerans_Agy99 AAASIAD---VADPVQVCHEALALAG------------------------
MAB_3062c|M.abscessus_ATCC_199 AAAGAIA---VEDPVRVCQDVLAALR------------------------
Mb2986c|M.bovis_AF2122/97 LADAFGRD--RVGFPQHVENALRLMPENRPRTWLAS--------------
Rv2962c|M.tuberculosis_H37Rv LADAFGRD--RVGFPQHVENALRLMPENRPRTWLAS--------------
MMAR_1755|M.marinum_M LAEVFGRE--LARFPQHVESALRRVSEGAVAKSAAG--------------
MLBr_00125|M.leprae_Br4923 LAEVLERH--VAGLPQHMENVLHTVSQNPPPTSLARQKRPLRFG------
:
Mflv_2592|M.gilvum_PYR-GCK --------
MSMEG_4670|M.smegmatis_MC2_155 --------
Mvan_4035|M.vanbaalenii_PYR-1 --------
TH_3465|M.thermoresistible__bu RPARPRVH
MAV_3631|M.avium_104 --------
MUL_3383|M.ulcerans_Agy99 --------
MAB_3062c|M.abscessus_ATCC_199 --------
Mb2986c|M.bovis_AF2122/97 --------
Rv2962c|M.tuberculosis_H37Rv --------
MMAR_1755|M.marinum_M --------
MLBr_00125|M.leprae_Br4923 --------