For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
TAAVTPSASAVASDDKKSERRLAFWLIAPAVLLMLAVTAYPIGYAVWLSLQRYNLAEPHDTEFIGLANYV TVLTDGYWWTAFAVTLGITVVSVAIEFALGLALALVMHRTIFGKGAVRTAILIPYGIVTVAASYSWYYAW TPGTGYLANLLPEGSAPLTDQLPSLAIVVLAEVWKTTPFMALLLLAGLALVPQDLLNAAQVDGAGPWKRL TKVILPMIKPAILVALLFRTLDAFRIFDNIYILTGGSNDTGSVSILGYDNLFKAFNVGLGSAISVLIFLS VAIIAFIYIKIFGAAAPGSDEEVR*
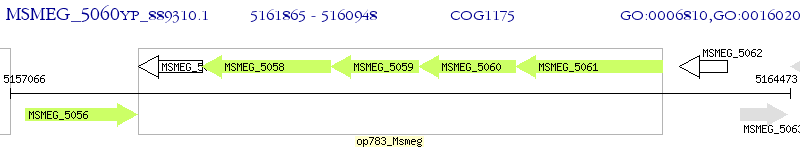
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_5060 | - | - | 100% (305) | ABC transporter, permease protein SugA |
| M. smegmatis MC2 155 | MSMEG_5573 | - | 1e-41 | 33.45% (278) | sugar ABC transporter permease protein |
| M. smegmatis MC2 155 | MSMEG_3110 | - | 5e-39 | 32.97% (273) | binding-protein-dependent transport systems inner membrane |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1268 | sugA | 1e-135 | 78.50% (307) | sugar-transport integral membrane protein ABC transporter |
| M. gilvum PYR-GCK | Mflv_3852 | - | 2e-79 | 51.60% (281) | binding-protein-dependent transport systems inner membrane |
| M. tuberculosis H37Rv | Rv1236 | sugA | 1e-135 | 78.50% (307) | sugar-transport integral membrane protein ABC transporter |
| M. leprae Br4923 | MLBr_01087 | - | 1e-128 | 74.16% (298) | putative ABC-transport protein, inner membrane component |
| M. abscessus ATCC 19977 | MAB_1373 | - | 1e-130 | 78.35% (291) | sugar ABC transporter, permease protein SugA |
| M. marinum M | MMAR_4205 | sugA | 1e-128 | 75.34% (296) | sugar-transport integral membrane protein ABC transporter |
| M. avium 104 | MAV_1375 | sugA | 1e-134 | 79.58% (289) | ABC transporter, permease protein SugA |
| M. thermoresistible (build 8) | TH_4307 | - | 1e-148 | 86.58% (298) | PUTATIVE binding-protein-dependent transport systems inner |
| M. ulcerans Agy99 | MUL_4512 | sugA | 1e-118 | 78.60% (257) | sugar-transport integral membrane protein ABC transporter |
| M. vanbaalenii PYR-1 | Mvan_2543 | - | 6e-79 | 51.04% (288) | binding-protein-dependent transport systems inner membrane |
CLUSTAL 2.0.9 multiple sequence alignment
MSMEG_5060|M.smegmatis_MC2_155 ---MTAAVT-----PSASAVASDDKKSERRLAFWLIAPAVLLMLAVTAYP
TH_4307|M.thermoresistible__bu ---MTAAAEKADPTPAAAVARTDDTRSERRLAYWLIAPAVLLMVAVTGYP
MAB_1373|M.abscessus_ATCC_1997 ---MTDRTS------------SEGKRAERRLGLLLIAPAALLMLAVTAYP
Mb1268|M.bovis_AF2122/97 MTSVEQRTA------TAVFSRTGSRMAERRLAFMLVAPAAMLMVAVTAYP
Rv1236|M.tuberculosis_H37Rv MTSVEQRTA------TAVFSRTGSRMAERRLAFMLVAPAAMLMVAVTAYP
MAV_1375|M.avium_104 -----------------MLGRT----SEQRLALVLVAPAAILMLAVTAYP
MMAR_4205|M.marinum_M ---------------MTALMRSRGRRAERRLAFALVAPATILMLLVIGYP
MUL_4512|M.ulcerans_Agy99 ---------------MTAFLRSRGRRAERRLAFALVAPATILMLLVIGYP
MLBr_01087|M.leprae_Br4923 ---MTAVVG------KSWHVRASSVQPEQRLAFLLVTPAAMLMLVVTAYP
Mflv_3852|M.gilvum_PYR-GCK ---MTAPADAPIQKDPGKPSVSERAKGERRLGLYLTLPSYVVMLLVTAYP
Mvan_2543|M.vanbaalenii_PYR-1 ---MTALHEAPIQKDPGTPAVSERARGERRLGIYLTLPSYVVMLLVTAYP
: *:**. * *: ::*: * .**
MSMEG_5060|M.smegmatis_MC2_155 IGYAVWLSLQRYNLAEPHDTEFIGLANYVTVLTDGYWWTAFAVTLGITVV
TH_4307|M.thermoresistible__bu IAYAVWLSLLRYNLAQPDDVSFVGLENYVTVLTDEYWWTAFAVTLGITVV
MAB_1373|M.abscessus_ATCC_1997 ICYAIWLSVQKYSFASPQDRKFVWFDNYITVLSDRYWWSALVVTLVITVI
Mb1268|M.bovis_AF2122/97 IGYALWLSLQRNNLATPNDTAFIGLGNYHTILIDRYWWTALAVTLAITAV
Rv1236|M.tuberculosis_H37Rv IGYALWLSLQRNNLATPNDTAFIGLGNYHTILIDRYWWTALAVTLAITAV
MAV_1375|M.avium_104 IGYAVWLSLQRNNLAAPHDTAFVGLSNYATILSDRYWWTALAVTLGITVV
MMAR_4205|M.marinum_M IGYAMWLSLQRNNLATPNETRFIGLGNYQTILVDRYWWTALAVTAAITVV
MUL_4512|M.ulcerans_Agy99 IGYAMWLSLQRNNLATPNETRFIGLGNYQTILVDRYWWTALAVTAAITVV
MLBr_01087|M.leprae_Br4923 IGYAVWLSLQRYSLATPNETVFIGLGNYQTILTDPYWWTALAVTLAITVV
Mflv_3852|M.gilvum_PYR-GCK LGYALVLSLYNFRLTDPEGRSFVGLGNYLVILTDPIWWNDFVTTLAITVV
Mvan_2543|M.vanbaalenii_PYR-1 LGYALVLSLYNFRLTDPEGRSFVGLSNYLVILTDPLWWNDFVTTLAITVV
: **: **: . :: *. *: : ** .:* * **. :..* **.:
MSMEG_5060|M.smegmatis_MC2_155 SVAIEFALGLALALVMHRTIFGKGAVRTAILIPYGIVTVAASYSWYYAWT
TH_4307|M.thermoresistible__bu SVAIEFVLGLALALVMHRTIFGKGVVRTAILIPYGIVTVAASYSWYYAWT
MAB_1373|M.abscessus_ATCC_1997 SVLIELVLGLALALVMHRTIFGRGVVRTAVLIPYGIVTVAASYSWYYAWT
Mb1268|M.bovis_AF2122/97 SVTIEFVLGLALALVMHRTLIGKGLVRTAVLIPYGIVTVVASYSWYYAWT
Rv1236|M.tuberculosis_H37Rv SVTIEFVLGLALALVMHRTLIGKGLVRTAVLIPYGIVTVVASYSWYYAWT
MAV_1375|M.avium_104 SVSAEFVLGLALALVMHRTLVGKGLVRTAVLIPYGIVTAVASYSWYYAWT
MMAR_4205|M.marinum_M SVACEFVLGLALALVMHRSLVAKGLVRTAVLVPYGIVTVVASYSWYYAWT
MUL_4512|M.ulcerans_Agy99 SVACEFVLGLALALVMHRSLVAKGLVRTAVLVPYGIVTVVASYSWYYAWT
MLBr_01087|M.leprae_Br4923 SVSIEFILGLMLALVMHRTLLGKSLVRIAVLIPYSIVTVVASYSWYYAWT
Mflv_3852|M.gilvum_PYR-GCK TVAIELVLGFWFAFVMLRIVRGRGPLRTAILIPYGIVTVVSAFIWRYAFA
Mvan_2543|M.vanbaalenii_PYR-1 TVAIELVLGFWFAFVMLRIVRGRGPLRTAILIPYGIVTVVSAFIWRYAFA
:* *: **: :*:** * : .:. :* *:*:**.***..::: * **::
MSMEG_5060|M.smegmatis_MC2_155 PGTGYLANLLPEGSAPLT-DQLPSLAIVVLAEVWKTTPFMALLLLAGLAL
TH_4307|M.thermoresistible__bu PGTGYLANLLPDGSAPLT-QQLPSLAIVVLAEVWKTTPFMALLLLAGLAL
MAB_1373|M.abscessus_ATCC_1997 PGTGYLANLLPAGSAPLT-QQIPSLAVVILAEVWKTTPFMALLLLAGLAL
Mb1268|M.bovis_AF2122/97 PGTGYLANLLPYDSAPLT-QQIPSLGIVVIAEVWKTTPFMSLLLLAGLAL
Rv1236|M.tuberculosis_H37Rv PGTGYLANLLPYDSAPLT-QQIPSLGIVVIAEVWKTTPFMSLLLLAGLAL
MAV_1375|M.avium_104 PGTGYLANLLPHGSAPLT-AQIPSLAIVVLAEVWKTTPFMSLLLLAGLAL
MMAR_4205|M.marinum_M PGTGYLANLLPQGSAPLT-QQIPSLGIVVIAEVWKTTPFMSLLLLAGLAM
MUL_4512|M.ulcerans_Agy99 PGTGYLANLLPQGIAPLT-QQIPSLGIVVIAEVWKTTPFMSLLLLAGLAM
MLBr_01087|M.leprae_Br4923 PGTGYLASLLPQGSVPLT-EQIPSLSIVIIVEVWKTTPFMSLLLLVGLAL
Mflv_3852|M.gilvum_PYR-GCK IDSGFVNQWLNLTDFDWFGDRWSAIFVICLSEIWKTTPFISLLLLAGLVQ
Mvan_2543|M.vanbaalenii_PYR-1 IDSGFVNQWLNLTEFDWFGDRWSAIFVICLSEIWKTTPFISLLLLAGLVQ
.:*:: . * : .:: :: : *:******::****.**.
MSMEG_5060|M.smegmatis_MC2_155 VPQDLLNAAQVDGAGPWKRLTKVILPMIKPAILVALLFRTLDAFRIFDNI
TH_4307|M.thermoresistible__bu VPQELLNAAQMDGANAWQRLVKIILPMIKPAILVALLFRTLDAFRIFDNI
MAB_1373|M.abscessus_ATCC_1997 VPDDLLKAAQVDGANGWSRLIRVTVPIMKPAILVALLFRTLDAFRIFDNI
Mb1268|M.bovis_AF2122/97 VPEDLLRAAQVDGASAWRRLTKVILPMIKPAIVVALLFRTLDAFRIFDNI
Rv1236|M.tuberculosis_H37Rv VPEDLLRAAQVDGASAWRRLTKVILPMIKPAIVVALLFRTLDAFRIFDNI
MAV_1375|M.avium_104 VPEDLLKAAQVDGAGAWRRLTRVTLPIIKPAVVVALLFRTLDAFRIFDNI
MMAR_4205|M.marinum_M VPEDLLQAAQVDGAGAWRRLTKIILPLIKPAVLVALLFRTLDAFRIFDNI
MUL_4512|M.ulcerans_Agy99 VPEDLLQAAQVDGAGAWRRLTKIILPLIKPAVLVALLFRTLDAFRIFDNI
MLBr_01087|M.leprae_Br4923 VPEDLLKAAQMDGAGAWRRLTKIILPIIKPAVMVALLFRTLDAFRIFDNI
Mflv_3852|M.gilvum_PYR-GCK VPEDMQEAAKVDGATAWQRLWKITLPNMKAAIMVALLFRTLDAWRIFDNP
Mvan_2543|M.vanbaalenii_PYR-1 VPEDMQEAAKVDGATAWQRLWKITLPNMKAAIMVALLFRTLDAWRIFDNP
**::: .**::*** * ** :: :* :*.*::**********:*****
MSMEG_5060|M.smegmatis_MC2_155 YILTGGSNDTGSVSILGYDNLFKAFNVGLGSA----------------IS
TH_4307|M.thermoresistible__bu YILTGGANNTGSVSILGYNNLFKAFNLGLGSA----------------IS
MAB_1373|M.abscessus_ATCC_1997 YVLTGGSNNTGSVSILGYDNLFKAFNVGLGSA----------------IS
Mb1268|M.bovis_AF2122/97 YVLTGGSNNTGSVSILGYDNLFKGFNVGLGSA----------------IS
Rv1236|M.tuberculosis_H37Rv YVLTGGSNNTGSVSILGYDNLFKGFNVGLGSA----------------IS
MAV_1375|M.avium_104 YVLTNGANNTGSVSMLGYDNLFKGFNVGLGSA----------------IS
MMAR_4205|M.marinum_M YVLTGGDNDTASVSILGYDNLFKGFNVGLGSA----------------IS
MUL_4512|M.ulcerans_Agy99 YVLTGGDNDTASVSILGYDNLFKGFNVGLAVQGIQCGPRFGDQRVDLRLC
MLBr_01087|M.leprae_Br4923 YVLTRGVNNTDSVSILGYDNLFKGFNVGLGSA----------------IS
Mflv_3852|M.gilvum_PYR-GCK YVMTAGANNTETISFLAYRQNVTLVNLGMGSA----------------VS
Mvan_2543|M.vanbaalenii_PYR-1 YVMTAGANNTETISFLAYRQNVTLVNLGMGSA----------------VS
*::* * *:* ::*:*.* : .. .*:*:. :.
MSMEG_5060|M.smegmatis_MC2_155 VLIFLSVAIIAFIYIKIFGA----AAPGSDEEVR----------------
TH_4307|M.thermoresistible__bu VLIFLCVAVIAFIYIKIFGT----SAPGSDAERS----------------
MAB_1373|M.abscessus_ATCC_1997 VLIFLCVAIIAVIFIKGFGA----SAPTTDGEEARR--------------
Mb1268|M.bovis_AF2122/97 VLIFGCVAVIAFIFIKLFGA----AAPGGEPSGR----------------
Rv1236|M.tuberculosis_H37Rv VLIFGCVAVIAFIFIKLFGA----AAPGGEPSGR----------------
MAV_1375|M.avium_104 VLIFGCVGLIALVFVKVFGA----AAPGGDVDGR----------------
MMAR_4205|M.marinum_M VLIFGCVTLIAVVFITLFGAHQFHSAPGGVTDAR----------------
MUL_4512|M.ulcerans_Agy99 HSHCGCLHHAVRRPSIRFCARWCDRCALRLGPCRRVAGGERSGWPSTPWS
MLBr_01087|M.leprae_Br4923 VLIFGCVALIAVIFIKVFGV----VAPGGDSNGY----------------
Mflv_3852|M.gilvum_PYR-GCK VLLFLSVVAIAWIFIKVFKT----DLSQVRGDQ-----------------
Mvan_2543|M.vanbaalenii_PYR-1 VLLFLSVVGIAWIFIKVFKT----DLSQVRGDQ-----------------
.: . * . .
MSMEG_5060|M.smegmatis_MC2_155 -----------
TH_4307|M.thermoresistible__bu -----------
MAB_1373|M.abscessus_ATCC_1997 -----------
Mb1268|M.bovis_AF2122/97 -----------
Rv1236|M.tuberculosis_H37Rv -----------
MAV_1375|M.avium_104 -----------
MMAR_4205|M.marinum_M -----------
MUL_4512|M.ulcerans_Agy99 WSTPCSRCCGS
MLBr_01087|M.leprae_Br4923 -----------
Mflv_3852|M.gilvum_PYR-GCK -----------
Mvan_2543|M.vanbaalenii_PYR-1 -----------