For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MTNTMTQIRSDLWQTRTDSPFPGLTTHSYLWKPNGHSSSPGVLFYSPATDADFAAIDALGGVGHQYLSHQ DEAGPMLARIAEHYGARLHAPAAELANIARHAKVHVPLDRRHRDDNGVEVIPTPGHSPGSTSYLVDGADG RYLFTGDTVFVDHAGRWSTFVIPGIGDPADMAESLRLLATLEPDVVISSAFGGAAVSTIDDRPWARCVDE AMGSLAA
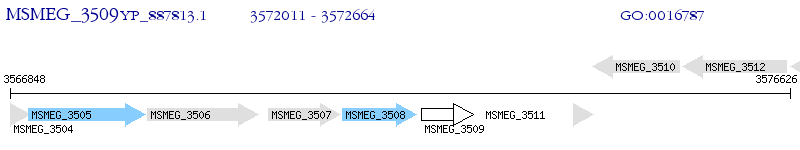
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_3509 | - | - | 100% (217) | hypothetical protein MSMEG_3509 |
| M. smegmatis MC2 155 | MSMEG_1334 | - | 2e-07 | 33.90% (118) | metallo-beta-lactamase family protein |
| M. smegmatis MC2 155 | MSMEG_6679 | - | 8e-05 | 32.67% (101) | metallo-beta-lactamase family protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0651c | - | 5e-09 | 34.17% (120) | glyoxalase II |
| M. gilvum PYR-GCK | Mflv_1248 | - | 8e-60 | 56.74% (215) | hypothetical protein Mflv_1248 |
| M. tuberculosis H37Rv | Rv0634c | - | 5e-09 | 34.17% (120) | glyoxalase II |
| M. leprae Br4923 | MLBr_01912 | - | 7e-08 | 33.04% (112) | putative glyoxylase II |
| M. abscessus ATCC 19977 | - | - | - | - | - |
| M. marinum M | MMAR_0965 | gloB | 6e-08 | 32.20% (118) | glyoxalase II, GloB |
| M. avium 104 | MAV_4532 | - | 8e-08 | 32.20% (118) | metallo-beta-lactamase family protein |
| M. thermoresistible (build 8) | TH_1674 | - | 5e-70 | 60.75% (214) | conserved hypothetical protein |
| M. ulcerans Agy99 | MUL_0715 | gloB | 4e-08 | 32.20% (118) | glyoxalase II, GloB |
| M. vanbaalenii PYR-1 | Mvan_5553 | - | 3e-67 | 60.00% (215) | hypothetical protein Mvan_5553 |
CLUSTAL 2.0.9 multiple sequence alignment
Mb0651c|M.bovis_AF2122/97 MSK-DRLYFRQLLSGRDFAVGDMFATQMRNFAYLIGDRTTGDCVVVDPAY
Rv0634c|M.tuberculosis_H37Rv MSK-DRLYFRQLLSGRDFAVGDMFATQMRNFAYLIGDRTTGDCVVVDPAY
MAV_4532|M.avium_104 MSD-DRLYFRQLLSGRDFAAGDMFATQMRNFAYLIGDRQTGDCVVVDPAY
MMAR_0965|M.marinum_M MSDSDRLYFRQLLAGRDFAVNDMIAAQMRNFAYLIGDRETGDCVVVDPAY
MUL_0715|M.ulcerans_Agy99 MSDSDRLYFRQLLSGRDFAVNDMIAAQMRNFAYLIGDRETGDCVVVDPAY
MLBr_01912|M.leprae_Br4923 MSDSDRLYFRQLLSGRDFAVDDVVATQMRNFAYLIGDRQTGDCVVVDPAY
Mflv_1248|M.gilvum_PYR-GCK --------MRYVLPDLAETRTDSPFPGLTTHAYLWTP---------ANIL
Mvan_5553|M.vanbaalenii_PYR-1 --------MKQVLRDLWETRTDSPFPGLTTHAYLWTP---------ANIL
TH_1674|M.thermoresistible__bu --------MHQVLPTLWQTRTDSPFPGLTTHAFLWTGG-------RRNAL
MSMEG_3509|M.smegmatis_MC2_155 ----MTNTMTQIRSDLWQTRTDSPFPGLTTHSYLWKPNGH---SSSPGVL
: : : * . : ..::*
Mb0651c|M.bovis_AF2122/97 AAGDLLDALESDDMQLSGVLVTHHHPDHVGGSMMGFQLPGLAELLERASV
Rv0634c|M.tuberculosis_H37Rv AAGDLLDALESDDMQLSGVLVTHHHPDHVGGSMMGFQLPGLAELLERASV
MAV_4532|M.avium_104 AASDLLDTLEADGMHLSGVLVTHHHPDHVGGSMMGFTLAGLAELVERAPV
MMAR_0965|M.marinum_M AAGDLVNALEADDMHLSGVLVTHHHPDHVGGSMMGFELKGLAELLERVSV
MUL_0715|M.ulcerans_Agy99 AAGDLVNALEAGDMHLSGVLVTHHHPDHVGGSMMGFELKGLAELLERVSV
MLBr_01912|M.leprae_Br4923 AAGDLVDALEADGMRLSGVLVTHHHPDHVGGVMMGFQLKGLTELLERVTV
Mflv_1248|M.gilvum_PYR-GCK FYSPATDADFDDLAALGGVSDQYLSHRDEAGPML-------RQIAERFGT
Mvan_5553|M.vanbaalenii_PYR-1 FYSPATDAQFNELTELGGVNDQYLSHRDEAGPML-------ARIAERFGS
TH_1674|M.thermoresistible__bu FYSVATDADFDAIEQLGGIADQYLSHRDEAGPML-------ARIARRFGS
MSMEG_3509|M.smegmatis_MC2_155 FYSPATDADFAAIDALGGVGHQYLSHQDEAGPML-------ARIAEHYGA
. :: *.*: : . .* *: .: .:
Mb0651c|M.bovis_AF2122/97 PVHVNTHEALWVSRVTGIPVGDLITHEHGDKVSVGDIDIELLHTPGHTPG
Rv0634c|M.tuberculosis_H37Rv PVHVNTHEALWVSRVTGIPVGDLITHEHGDKVSVGDIDIELLHTPGHTPG
MAV_4532|M.avium_104 PVHVNSHEALWVSRVTGISASDLTAHENHDKVSIGDIEIELLHTPGHTPG
MMAR_0965|M.marinum_M PVHVNTHEAVWVSRVTGIPMSDLTTHEHRDKVSVGDIEIELLHTPGHTPG
MUL_0715|M.ulcerans_Agy99 PVHVNTHEAVWVSRVTGIPMSDLTTHEHRDKVSVGDIEIELLHTPGHTPG
MLBr_01912|M.leprae_Br4923 PVHVNTHEALWISRVTGIDIGHLTTHEHRDKLSVGDIEIELLHTPGHTPG
Mflv_1248|M.gilvum_PYR-GCK TLHAPLGDRADIEKHAHVDVP-------LSGHYVDYNGVEVIPTPGHSPG
Mvan_5553|M.vanbaalenii_PYR-1 TLHAPAADLADIGKHAHVDVP-------LKGRYVDYNGVEIIPTPGHSPG
TH_1674|M.thermoresistible__bu RLHAPAAESAEISAHATIDVP-------LDRRHVDDNGIEVIPTPGHTPG
MSMEG_3509|M.smegmatis_MC2_155 RLHAPAAELANIARHAKVHVP-------LDRRHRDDNGVEVIPTPGHSPG
:*. : : : : . . :*:: ****:**
Mb0651c|M.bovis_AF2122/97 SQCFLLDG-----RLVAGDTLFLEGCG---RTDFPGGD----SDEMYRSL
Rv0634c|M.tuberculosis_H37Rv SQCFLLDG-----RLVAGDTLFLEGCG---RTDFPGGD----SDEMYRSL
MAV_4532|M.avium_104 SQCFLLDG-----RLVAGDTLFLEGCG---RTDFPGGD----SDEMYRSL
MMAR_0965|M.marinum_M SQCFLLDG-----RLVAGDTLFLEGCG---RTDFPGGD----SDEIFRSL
MUL_0715|M.ulcerans_Agy99 SQCFLLDG-----QLVAGDTLFLEGCG---RTDFPGGD----SDEIFRSL
MLBr_01912|M.leprae_Br4923 SQCFLLDG-----LLVAGDTLFLEGCG---RTDFPGGN----SDEMYRSL
Mflv_1248|M.gilvum_PYR-GCK AVCYLVTGAAG-RYLFTGDTLFRSAEGHWYAGYIEGFHHLDDADTIAASL
Mvan_5553|M.vanbaalenii_PYR-1 SVSYLVSGADG-RYLFTGDTLFRNPDGHWWAGYIEGFHQPGDADTIAESL
TH_1674|M.thermoresistible__bu SVCYLVAGSGGARYLFTGDTLYVDASGAWTAGYLPGMS---DARSLDHSL
MSMEG_3509|M.smegmatis_MC2_155 STSYLVDGADG-RYLFTGDTVFVDHAGRWSTFVIPGIG---DPADMAESL
: .:*: * *.:***:: . * : * . : **
Mb0651c|M.bovis_AF2122/97 RQLAELPGDPTVFPGHWYSAEPSASLSEVKRSNYVYRPASLDQWRMLMGG
Rv0634c|M.tuberculosis_H37Rv RQLAELPGDPTVFPGHWYSAEPSASLSEVKRSNYVYRPASLDQWRMLMGG
MAV_4532|M.avium_104 QRLAKLPGDPTVFPGHWYSAEPSASLSEVKRSNYVYRASDLDQWRMLMGG
MMAR_0965|M.marinum_M QQLAALPGDPTVFPGHWYSAEPSASLSEVKRSNFVYRAADLAQWRMLMGG
MUL_0715|M.ulcerans_Agy99 QQLAALPGDPTVFPGHWYSAEPSASLSEVKRSNFVYRAADLAQWRMLMGG
MLBr_01912|M.leprae_Br4923 QQLAQLPGNPTVFPGHWYSTAPSATLSMVKCSNYVYRAADLNQWRMLMDS
Mflv_1248|M.gilvum_PYR-GCK RTLADLTPDFVISSAFQGESAVHRIDADAWRGHIAHAIAGLPNSART---
Mvan_5553|M.vanbaalenii_PYR-1 RTLAEVSPDFVISSAFQGNSAVHRVEPDLWRGHIAHAIAGLPTSARR---
TH_1674|M.thermoresistible__bu ELLATLTPDVVISSAFAGDSAVHPVNRNRWPEHVEQARAGLAT--RR---
MSMEG_3509|M.smegmatis_MC2_155 RLLATLEPDVVISSAFGG-AAVSTIDDRPWARCVDEAMGSLAA-------
. ** : : .: ... : ..*