For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VRRLRPMAVTGDDLRHLRRCVELAREAQDAGDEPFGSLLLDADGVTRVEDRNRVKDGDATRHPEYAIAKW AVENLSPDDRATVYTSGEHCPMCAAAHAWVGLGRVVYAASSVQLAQWLVEWKAPPPPVASLPITTVAPGV PVDGPAPELTEEMKTLYERAFRR
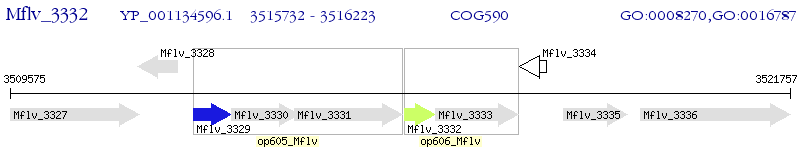
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. gilvum PYR-GCK | Mflv_3332 | - | - | 100% (163) | CMP/dCMP deaminase, zinc-binding |
| M. gilvum PYR-GCK | Mflv_1214 | - | 6e-05 | 31.53% (111) | CMP/dCMP deaminase, zinc-binding |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3141 | - | 2e-06 | 27.72% (101) | hypothetical protein Mb3141 |
| M. tuberculosis H37Rv | Rv3114 | - | 2e-06 | 27.72% (101) | hypothetical protein Rv3114 |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_3459 | - | 2e-58 | 69.33% (150) | putative cytidine/deoxycytidylate deaminase |
| M. marinum M | - | - | - | - | - |
| M. avium 104 | MAV_0777 | - | 3e-05 | 26.42% (106) | cytidine/deoxycytidylate deaminase |
| M. smegmatis MC2 155 | MSMEG_3575 | - | 2e-61 | 69.81% (159) | cytidine/deoxycytidylate deaminase, zinc-binding region |
| M. thermoresistible (build 8) | TH_2296 | - | 2e-64 | 71.52% (158) | cytidine/deoxycytidylate deaminase, zinc-binding region |
| M. ulcerans Agy99 | - | - | - | - | - |
| M. vanbaalenii PYR-1 | Mvan_3058 | - | 4e-68 | 78.48% (158) | CMP/dCMP deaminase, zinc-binding |
CLUSTAL 2.0.9 multiple sequence alignment
Mb3141|M.bovis_AF2122/97 MVAARLPFGWPADSGVTADIIEAAMELAIDTARHATAPFGAALLDVTTLR
Rv3114|M.tuberculosis_H37Rv MVAARLPFGWSADSGVTADIIEAAMELAIDTARHATAPFGAALLDVTTLR
Mflv_3332|M.gilvum_PYR-GCK ----MRRLRPMAVTGDDLRHLRRCVELAREAQDAGDEPFGSLLLDADGVT
Mvan_3058|M.vanbaalenii_PYR-1 ----------MAVTDDDLVFLRRCVSLARTALDGGDEPFGSLLVDHEGSV
MSMEG_3575|M.smegmatis_MC2_155 ----------MAISDADLKYLRRCVDLAREALDDGDEPFGSVLVDHTGTT
TH_2296|M.thermoresistible__bu ----------VAIGDDDLRYLHRCVELARAALAAGDEPFGSILVDADGQI
MAB_3459|M.abscessus_ATCC_1997 ------------MDDNDIQHLRRCVELATEALDAGDEPFGSLLTGPDGAV
MAV_0777|M.avium_104 ----------------MTDFARRTIDLARQNVAEGGRPFATVIVQ-DGEV
. :.** . **.: :
Mb3141|M.bovis_AF2122/97 AFSGGNTYFESGDRFAHAETNVLRAAMSTLPELS--NHVLISTAEPCPMC
Rv3114|M.tuberculosis_H37Rv AFSGGNTYFESGDRFAHAETNVLRAAMSTLPELS--NHVLISTAEPCPMC
Mflv_3332|M.gilvum_PYR-GCK RVEDRNR-VKDGDATRHPEYAIAKWAVENLSPD--DRATVYTSGEHCPMC
Mvan_3058|M.vanbaalenii_PYR-1 RFEDHNR-VKDGDATRHPEFAIARWAVEHLSPAQRNRSTVYTSGEHCPMC
MSMEG_3575|M.smegmatis_MC2_155 LFEDRNR-VKDGDATAHPEFAIARWAARHLTPDRRARATVYTSGEHCPMC
TH_2296|M.thermoresistible__bu RYEDRNR-VSGGDRTRHPEFAIARWAAQHLGPDERGRATVYTSGEHCPMC
MAB_3459|M.abscessus_ATCC_1997 LAEDRNR-VGAGDPTRHPEFELARWSVRNLSPTDRALSTVYTSGEHCPMC
MAV_0777|M.avium_104 LAESANKAAQTNDPTAHAEILAIRAACLKLGTEHLVDATIYVLAHPCPMC
.. * .* *.* : : * .: .. ****
Mb3141|M.bovis_AF2122/97 AAASVLSGVRAIIFGTSIETLIQCG-----WFQIRISASDVVAASTRPTR
Rv3114|M.tuberculosis_H37Rv AAASVLSGVRAIIFGTSIETLIQCG-----WFQIRISASDVVAASTRPTR
Mflv_3332|M.gilvum_PYR-GCK AAAHAWVGLGRVVYAASSVQLAQW------LVEWKAPPPPVASLPITTVA
Mvan_3058|M.vanbaalenii_PYR-1 AAAHAWVGLGRVVYAASSAQLAQW------LREWGVPAPPVATLPIATVA
MSMEG_3575|M.smegmatis_MC2_155 AAAHAWVGLGRIVYATSSAQLGGW------LTEWGAQAPPVATLPINTVA
TH_2296|M.thermoresistible__bu AAAHAWVGLGRIVYAASAAQLSGW------LAEWGAPPGPVAPLPVTVVA
MAB_3459|M.abscessus_ATCC_1997 SAAHAWVGLGRIVYASSSAQLTQW------LTELGAPPSPVASLSINEVA
MAV_0777|M.avium_104 LGALYYCSPREVIFLTTRESYEPYYVDDRKYFELSNFYAEFAKPFDERRL
.* . ::: :: : ..
Mb3141|M.bovis_AF2122/97 PSVYSGFLSHKTDLLYRNSENRRAMNPWTDPSH
Rv3114|M.tuberculosis_H37Rv PSVYSGFLSHKTDLLYRNSENRRAMNPWTDPSH
Mflv_3332|M.gilvum_PYR-GCK PGVPVDGPAPELTEEMKTLYERAFRR-------
Mvan_3058|M.vanbaalenii_PYR-1 PGVDVDGPAAELTEEMKALYARAFRA-------
MSMEG_3575|M.smegmatis_MC2_155 PGVVVDGPAEELAETMHNLYRAKFGR-------
TH_2296|M.thermoresistible__bu PGLVADGPAAELTETMRELYERKFRP-------
MAB_3459|M.abscessus_ATCC_1997 PGVQTDGPAPDLAEDVRALHVRFLGDA------
MAV_0777|M.avium_104 PMRYQPHPDAVDVYRLWNERNGGRRRSTVVPSD
*