For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
NVRKVVSKGLMAAAISGALAVVPMAMDGTTATAHADSVNWDAIAQCESGGNWAINTGNGHFGGLQFKQAT WSANGGVGNPAAAPRHEQIRVAENVLRTQGLKAWPKCGAKGMAAQVWGNTAPTAPVPATVPAATPGRANG CATMPTSVFGGVLDLRKMCTALFTPRSAR*
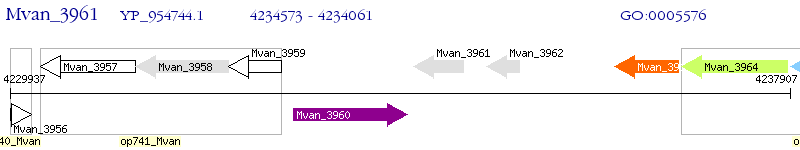
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_3961 | - | - | 100% (170) | transglycosylase domain-containing protein |
| M. vanbaalenii PYR-1 | Mvan_3962 | - | 2e-36 | 64.55% (110) | transglycosylase domain-containing protein |
| M. vanbaalenii PYR-1 | Mvan_5049 | - | 2e-22 | 47.33% (131) | FHA domain-containing protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1916c | rpfC | 1e-33 | 57.63% (118) | resuscitation-promoting factor RpfC |
| M. gilvum PYR-GCK | Mflv_2620 | - | 5e-92 | 91.76% (170) | transglycosylase domain-containing protein |
| M. tuberculosis H37Rv | Rv1884c | rpfC | 1e-33 | 57.63% (118) | resuscitation-promoting factor RpfC |
| M. leprae Br4923 | MLBr_02030 | - | 7e-30 | 49.61% (127) | hypothetical protein MLBr_02030 |
| M. abscessus ATCC 19977 | MAB_4080c | - | 4e-35 | 48.92% (139) | hypothetical protein MAB_4080c |
| M. marinum M | MMAR_2772 | - | 2e-31 | 49.21% (126) | resuscitation-promoting factor-like protein |
| M. avium 104 | MAV_1722 | - | 8e-31 | 74.03% (77) | secreted protein |
| M. smegmatis MC2 155 | MSMEG_4640 | - | 3e-47 | 54.60% (163) | secreted protein |
| M. thermoresistible (build 8) | TH_0358 | - | 4e-48 | 55.03% (169) | PROBABLE RESUSCITATION-PROMOTING FACTOR RPFE |
| M. ulcerans Agy99 | MUL_2975 | - | 5e-32 | 49.21% (126) | resuscitation-promoting factor-like protein |
CLUSTAL 2.0.9 multiple sequence alignment
Mvan_3961|M.vanbaalenii_PYR-1 ----------------------------MN-----VRKVVSKGLMAAAIS
Mflv_2620|M.gilvum_PYR-GCK ----------------------------MN-----VRKLAGKGLMAAAIS
MSMEG_4640|M.smegmatis_MC2_155 ---------------------------------------MKRAFGLTALA
TH_0358|M.thermoresistible__bu ----------------------------MNNIRKALTSSLKRALLLIAGT
Mb1916c|M.bovis_AF2122/97 MHPLPADHGRSRCNRHPISPLSLIGNASATSGDMSSMTRIAKPLIKSAMA
Rv1884c|M.tuberculosis_H37Rv MHPLPADHGRSRCNRHPISPLSLIGNASATSGDMSSMTRIAKPLIKSAMA
MLBr_02030|M.leprae_Br4923 ---------------------------MDMPGEMLDVRKLCKLFVKSAVV
MMAR_2772|M.marinum_M ------------------------------------MTHIAKPLTKSALA
MUL_2975|M.ulcerans_Agy99 ---------------------------MATSAEKSDMTHIAKPLTKSALA
MAB_4080c|M.abscessus_ATCC_199 ---------------------------------MDRARWVRDRRAVCLIF
MAV_1722|M.avium_104 -----MGFDPNVPAAGPAPDAPPPPAPDAPPAPAPDAPPAPDAPPPPAPD
Mvan_3961|M.vanbaalenii_PYR-1 GALAVVPMAMDGT------TATAHADS--VNWDAIAQCESGGNWAINTGN
Mflv_2620|M.gilvum_PYR-GCK GALAVVPMALDGG------TATAHADS--VNWDAIAQCESGGNWAINTGN
MSMEG_4640|M.smegmatis_MC2_155 GALAMGPMVLS--------PATAAADS--VNWDAIAACESGGNWSTNTGN
TH_0358|M.thermoresistible__bu AALALVPMVLS--------TATAQADT--VNWDAIAQCESGGNWASDTGN
Mb1916c|M.bovis_AF2122/97 AGLVTASMSLST--------AVAHAGP-SPNWDAVAQCESGGNWAANTGN
Rv1884c|M.tuberculosis_H37Rv AGLVTASMSLST--------AVAHAGP-SPNWDAVAQCESGGNWAANTGN
MLBr_02030|M.leprae_Br4923 SGIVTASMALSTS------TGMANAVPREPNWDAVAQCESGRNWRANTGN
MMAR_2772|M.marinum_M GGFVVASMSLST--------GVAQADL--MNWEAVAQCESGGNWSANTGN
MUL_2975|M.ulcerans_Agy99 GGFVVASMSLST--------GVAQADL--MNWEAVAQCESGGNWSVNTGN
MAB_4080c|M.abscessus_ATCC_199 TVLFMMSLSMTA--------SVASADS--MNWDAVAECESGGNWSANTGN
MAV_1722|M.avium_104 APPPPAPDAPPAPGPDVPPPPAPKAYS--VNWDAIAQCESGGNWGINTGN
. . * **:*:* **** ** :***
Mvan_3961|M.vanbaalenii_PYR-1 GHFGGLQFKQATWSANGGVGNPAAAPRHEQIRVAENVLRTQGLKAWPKCG
Mflv_2620|M.gilvum_PYR-GCK GHFGGLQFKQATWSANGGVGNPARAPRHEQIRVAENVLRTQGIKAWPKCG
MSMEG_4640|M.smegmatis_MC2_155 GHYGGLQFKQSTWAAHGGVGSPATASREQQIAVAENVLRTQGLAAWPKCG
TH_0358|M.thermoresistible__bu GFYGGLQFKPSTWAAHGGTGSPARASREEQIRVAERVLASQGLKAWPKCG
Mb1916c|M.bovis_AF2122/97 GKYGGLQFKPATWAAFGGVGNPAAASREQQIAVANRVLAEQGLDAWPTCG
Rv1884c|M.tuberculosis_H37Rv GKYGGLQFKPATWAAFGGVGNPAAASREQQIAVANRVLAEQGLDAWPTCG
MLBr_02030|M.leprae_Br4923 GFYGGLQFKPTIWARYGGVGNPAGASREQQITVANRVLADQGLDAWPKCG
MMAR_2772|M.marinum_M GAYGGLQFKQATWEEYGGVGNPASASKQQQIAIANRVLAGQGPAAWPKCS
MUL_2975|M.ulcerans_Agy99 GAYGGLQFKQATWEEYGGVGNPASASKQQQIAIANRVLAGQGPAAWPKCS
MAB_4080c|M.abscessus_ATCC_199 GFYGGLQFKPSTWAAHGGVGNPAEASREEQIAVAEDVLRTQGPGAWPKCG
MAV_1722|M.avium_104 GYAGGLQFTSSTWHANGGSGSPAGASREEQIRVAENVLHSQGIGAWPVCG
* *****. : * ** *.** *.:.:** :*: ** ** *** *.
Mvan_3961|M.vanbaalenii_PYR-1 AKGMAAQVWGNTAPTAPVPATVPAATPGRANGCATMPTSVFGGVLDLRKM
Mflv_2620|M.gilvum_PYR-GCK AKGMAAQVWGNPAPSAPVPAVTPAATPGRANGCATMPTSVFGGIVDLRKM
MSMEG_4640|M.smegmatis_MC2_155 ARGGVPAVWGG-GAGAPAAG----------SGCSAVRPGAVLGILDLRRL
TH_0358|M.thermoresistible__bu AHGTRPAVFR--TPGAPAAGGVPASTPTATTGCGAIRPGAVLGLVDLRQV
Mb1916c|M.bovis_AF2122/97 AASGLP--IALWSKPA---------------------QGIKQIINEIIWA
Rv1884c|M.tuberculosis_H37Rv AASGLP--IALWSKPA---------------------QGIKQIINEIIWA
MLBr_02030|M.leprae_Br4923 AASDLP--ITLWSHPA---------------------QGVKQIINDIIQM
MMAR_2772|M.marinum_M SAGGLPPVPVVLPKPV---------------------RQLEQTVTQIISI
MUL_2975|M.ulcerans_Agy99 SAGGLPPVPVVLPKPV---------------------RQLEQTVTQVISI
MAB_4080c|M.abscessus_ATCC_199 GG-GIGGGAGCQFVPG---------------------GSIFGIVN-FRQM
MAV_1722|M.avium_104 RRG-----------------------------------------------
Mvan_3961|M.vanbaalenii_PYR-1 CTALFTPRSAR--
Mflv_2620|M.gilvum_PYR-GCK CTALFVPRSAR--
MSMEG_4640|M.smegmatis_MC2_155 CSTLLDPLGAFGR
TH_0358|M.thermoresistible__bu CAAFLTPLGVG--
Mb1916c|M.bovis_AF2122/97 G--IQASIPR---
Rv1884c|M.tuberculosis_H37Rv G--IQASIPR---
MLBr_02030|M.leprae_Br4923 GDTTLAAIALNGL
MMAR_2772|M.marinum_M F------TPR---
MUL_2975|M.ulcerans_Agy99 F------TPR---
MAB_4080c|M.abscessus_ATCC_199 CNGVMGLIPPAG-
MAV_1722|M.avium_104 -------------