For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
TIEVSNESGIDVSEEELISVARFVIGKMDVNPAAELSMLLLDTAAMADLHMRWMDLPGPTDVMSFPMDEL EPGGRPDAPEPGPSMLGDIVLCPEFAAKQAADAGHTLGQELALLTVHGVLHLLGYDHAEPDEEKEMFALQ RELLEEWVADQVQAYHQDRQSQKDQRLLDKSRYFDELNHGDTP*
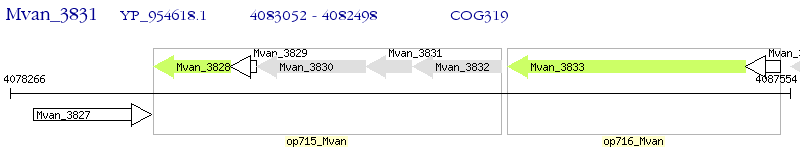
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_3831 | - | - | 100% (184) | putative metalloprotease |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb2388c | - | 4e-90 | 89.20% (176) | putative metalloprotease |
| M. gilvum PYR-GCK | Mflv_2707 | - | 4e-94 | 93.18% (176) | putative metalloprotease |
| M. tuberculosis H37Rv | Rv2367c | - | 4e-90 | 89.20% (176) | putative metalloprotease |
| M. leprae Br4923 | MLBr_00628 | - | 6e-84 | 83.71% (178) | putative metalloprotease |
| M. abscessus ATCC 19977 | MAB_1669 | - | 2e-76 | 77.40% (177) | putative metalloprotease |
| M. marinum M | MMAR_3677 | - | 2e-90 | 88.70% (177) | metal-dependent hydrolase |
| M. avium 104 | MAV_2026 | - | 1e-90 | 89.77% (176) | putative metalloprotease |
| M. smegmatis MC2 155 | MSMEG_4496 | - | 8e-91 | 89.83% (177) | putative metalloprotease |
| M. thermoresistible (build 8) | TH_1055 | - | 7e-90 | 88.76% (178) | UNKNOWN |
| M. ulcerans Agy99 | MUL_3620 | - | 1e-90 | 88.70% (177) | putative metalloprotease |
CLUSTAL 2.0.9 multiple sequence alignment
Mvan_3831|M.vanbaalenii_PYR-1 -----MTIEVSNESGIDVSEEELISVARFVIGKMDVNPAAELSMLLLDTA
Mflv_2707|M.gilvum_PYR-GCK -----MSIEVSNESGIDVSESELVSVARFVIRKMDVNPAAELSMVLLDTA
MMAR_3677|M.marinum_M -----MSIEVSNESGIDVSETELVSVARFVIGKMDVNPGAELSMVLLDTA
MUL_3620|M.ulcerans_Agy99 -----MSIEVSNESGIDVSETELVSVARFVIGKMDVNPGAELSMVLLDTA
MAV_2026|M.avium_104 -----MSIEVSNESGIDVSEAELISVARFVIAKMDVNPAAELSMVLLDTA
Mb2388c|M.bovis_AF2122/97 MREHLMSIEVANESGIDVSEAELVSVARFVIAKMDVNPCAELSMLLLDTA
Rv2367c|M.tuberculosis_H37Rv MREHLMSIEVANESGIDVSEAELVSVARFVIAKMDVNPCAELSMLLLDTA
MLBr_00628|M.leprae_Br4923 -----MSVEVSNESGFDVSEVELVSVARFVIIKMDVNPAAELSMVLLDTA
MSMEG_4496|M.smegmatis_MC2_155 -----MSIEVSNESGYDVSEPELISVARFVIEKMDVHPAAELSMVLLDSA
TH_1055|M.thermoresistible__bu -----MSIEVSNESGIDVCEEELISVARFVIEKMNVHPAAELSMVLLDPA
MAB_1669|M.abscessus_ATCC_1997 -----MSIEVVNESGIDVAEGELVSVARFAIAAMDVHPAAELSMMLVDLA
*::** **** **.* **:*****.* *:*:* *****:*:* *
Mvan_3831|M.vanbaalenii_PYR-1 AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPSMLGDIVLCPE
Mflv_2707|M.gilvum_PYR-GCK AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDTPEPGPSMLGDIVLCPE
MMAR_3677|M.marinum_M AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPAMLGDIVLCPE
MUL_3620|M.ulcerans_Agy99 AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPAMLGDIVLCPE
MAV_2026|M.avium_104 AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPAMLGDIVLCPE
Mb2388c|M.bovis_AF2122/97 AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPSMLGDIVLCPE
Rv2367c|M.tuberculosis_H37Rv AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPSMLGDIVLCPE
MLBr_00628|M.leprae_Br4923 AMADLHMRWMDLPGPTDVMSFPMDEFEPGGRPDAAEPGPSMLGDIVLCPE
MSMEG_4496|M.smegmatis_MC2_155 AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPAMLGDIVLCPE
TH_1055|M.thermoresistible__bu AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAAEPGPAMLGDIVLCPE
MAB_1669|M.abscessus_ATCC_1997 AMADLHMRWMDLPGPTDVMSFPMDELEPGGRPDAPEPGPSMLGDIVLCPQ
*************************:*******:.****:*********:
Mvan_3831|M.vanbaalenii_PYR-1 FAAKQAADAGHTLGQELALLTVHGVLHLLGYDHAEPDEEKEMFALQRELL
Mflv_2707|M.gilvum_PYR-GCK FAAKQAADAGHTLGQELALLTVHGVLHLLGYDHAEPDEEKEMFALQRELL
MMAR_3677|M.marinum_M FAAEQAAAAGHSLGHELALLTIHGVLHLLGYDHGEPDEEKEMFALQDRLL
MUL_3620|M.ulcerans_Agy99 FAAEQAAAAGHSLGHELALLTIHGVLHLLGYDHGEPDEEKEMFALQDRLL
MAV_2026|M.avium_104 FAADQAAAAGHSLGHELALLTIHGVLHLLGYDHGEPDEEKEMFALQDRLL
Mb2388c|M.bovis_AF2122/97 FAAEQAAAAGHSLGHELALLTIHGVLHLLGYDHAEPDEEKEMFALQDRLL
Rv2367c|M.tuberculosis_H37Rv FAAEQAAAAGHSLGHELALLTIHGVLHLLGYDHAEPDEEKEMFALQDRLL
MLBr_00628|M.leprae_Br4923 FAAQQAAAEGHSLGHELALLTIHGVLHLLGYDHGEPDEEKEMFALQGRLL
MSMEG_4496|M.smegmatis_MC2_155 FAEQQAAKAGHSLGHELALLTVHGVLHLLGYDHAEPDEEKEMFALQRQLL
TH_1055|M.thermoresistible__bu FAAEQAAAAGHSLGQELALLTVHGVLHLLGYDHAEPEEEKEMFALQRRLL
MAB_1669|M.abscessus_ATCC_1997 FAAEQAEAAGHSLAHELALLTVHGVLHLLGYDHAEPEEEREMFALQNQLL
** .** **:*.:******:***********.**:**:****** .**
Mvan_3831|M.vanbaalenii_PYR-1 EEWVADQVQAYHQDRQSQKDQRLLDKSRYFDELNHGDTP
Mflv_2707|M.gilvum_PYR-GCK EEWVAEQVEAYHLDRQNEKDRRLLDKSRYF------DTP
MMAR_3677|M.marinum_M EEWVAEQVQAYQQDRQDERDRRLLDKSRYFDEP------
MUL_3620|M.ulcerans_Agy99 EEWVAEQVQAYQQDRQDERDRRLLDKSRYFDEP------
MAV_2026|M.avium_104 EEWVADQVEAYHQDRQQERDRRLLDKSRYFDN-------
Mb2388c|M.bovis_AF2122/97 EEWVADQVEAYQHDRQDEKDRRLLDKSRYFDL-------
Rv2367c|M.tuberculosis_H37Rv EEWVADQVEAYQHDRQDEKDRRLLDKSRYFDL-------
MLBr_00628|M.leprae_Br4923 EEWVAEQVRAYQHDRQNEKDCRLLYKSGYFGYL------
MSMEG_4496|M.smegmatis_MC2_155 EEWVADQVEAYHADRQSEKDRRLLDKSRYFDEP------
TH_1055|M.thermoresistible__bu EEWVADQVEAYHQELQSEKDRRLLDKSRYFDQL------
MAB_1669|M.abscessus_ATCC_1997 QDWYEQQAQLY-------RDSRLLDKSRNFDE-------
::* :*.. * :* *** ** *