For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MATPTLPPGFDFTDPDLNRERLPVEELAELRRCAPIWWNEQPDGIGGFGDGGFWVVTKHHDVKEISKRSD VFSSEVKTALPRYPEGSTGEQIETGSLVLLNMDAPRHTHLRKIISRGFTPRAVERLRDDLNARAQNIAKT AASAGSGDFVEQVSCELPLQAIAGLLGVPVDDRKKLFDWSNQMVSDDDPEYAHYDNRNAATELIMYAMQL AAVRAEQPGEDIVTKLIEADVDGHKLSDDEFGFFMVLLAVAGNETTRNSITHGMIAFTEHPDQWELYKRE RPITAVDEIVRWATPVTSFQRTALTDYELSGVQITKGQRVVMSYRSANFDEEVFDDPFTFDILRDPNPHV GFGGTGAHYCIGANLARMTIDLMFNAIADHMPDLSAIGSPDRLRSGWLNGIKHWQVDYSGRGCPVAH
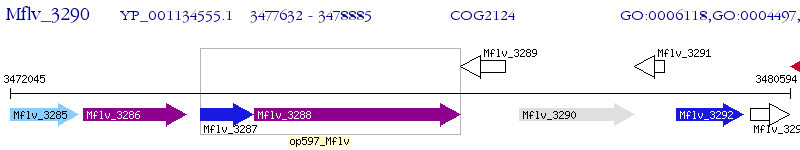
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. gilvum PYR-GCK | Mflv_3290 | - | - | 100% (417) | cytochrome P450 |
| M. gilvum PYR-GCK | Mflv_1508 | - | e-179 | 73.06% (412) | cytochrome P450 |
| M. gilvum PYR-GCK | Mflv_1607 | - | e-138 | 58.27% (405) | cytochrome P450 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3575c | cyp125 | 0.0 | 74.04% (416) | cytochrome P450 125 |
| M. tuberculosis H37Rv | Rv3545c | cyp125 | 0.0 | 74.04% (416) | cytochrome P450 125 |
| M. leprae Br4923 | MLBr_02088 | - | 1e-38 | 31.99% (297) | putative cytochrome p450 |
| M. abscessus ATCC 19977 | MAB_0613 | - | 0.0 | 72.71% (414) | putative cytochrome P450 |
| M. marinum M | MMAR_2783 | cyp125A6 | 0.0 | 78.52% (419) | cytochrome P450 125A6 Cyp125A6 |
| M. avium 104 | MAV_2811 | - | 0.0 | 79.38% (417) | putative cytochrome P450 124 |
| M. smegmatis MC2 155 | MSMEG_3524 | - | 0.0 | 79.01% (405) | linalool 8-monooxygenase |
| M. thermoresistible (build 8) | TH_1956 | - | 0.0 | 75.85% (414) | PROBABLE CYTOCHROME P450 125 CYP125 |
| M. ulcerans Agy99 | MUL_4106 | cyp125A7 | 0.0 | 73.73% (415) | cytochrome P450 125A7 Cyp125A7 |
| M. vanbaalenii PYR-1 | Mvan_3012 | - | 0.0 | 90.93% (419) | cytochrome P450 |
CLUSTAL 2.0.9 multiple sequence alignment
Mb3575c|M.bovis_AF2122/97 MSWNHQSVEIAVRRTTVPSPNLPPGFDFTDPAIYAERLPVAEFAELRSAA
Rv3545c|M.tuberculosis_H37Rv MSWNHQSVEIAVRRTTVPSPNLPPGFDFTDPAIYAERLPVAEFAELRSAA
MUL_4106|M.ulcerans_Agy99 ----------------MPCPNLPPGFDFTDPDIYAERLPVEEFAELRSSE
MAB_0613|M.abscessus_ATCC_1997 ----------------MVHPSLPAGFDFTDPEIYAERLPVEELKELRKTA
Mflv_3290|M.gilvum_PYR-GCK ----------------MATPTLPPGFDFTDPDLNRERLPVEELAELRRCA
Mvan_3012|M.vanbaalenii_PYR-1 ----------------MATPTLPPGFDFTDPDLNLERLPVEELAELRRCA
MMAR_2783|M.marinum_M --------MPAAEPTATSVPNLPPGFDFTDPDIYAERLPVAELAEMRRSA
MAV_2811|M.avium_104 --------MATVEPTTKPVPNLPPGFDFTDPDIYAERLPVEELAEMRRVA
MSMEG_3524|M.smegmatis_MC2_155 ------------MVMSDSALHLPAGFDFTDPDIYAERLPVDELAELRRVA
TH_1956|M.thermoresistible__bu ----------------MPTPNIPPGFDFTDPDIYAVRLPVEELAELRRSA
MLBr_02088|M.leprae_Br4923 --MRTCVPTRTCVYAFIEYLSHNRPMGTNPPSLVEAQMLLLRLIDPGTRA
:. . * : :: : .: :
Mb3575c|M.bovis_AF2122/97 ---PIWWNGQDPGKGGGFHDGGFWAITKLNDVKEISRHSDVFSSYENGVI
Rv3545c|M.tuberculosis_H37Rv ---PIWWNGQDPGKGGGFHDGGFWAITKLNDVKEISRHSDVFSSYENGVI
MUL_4106|M.ulcerans_Agy99 ---PIWWDEQFPGQGGGFHDGGFWAITKLKDVKEVSRRSDVFSSYENGVI
MAB_0613|M.abscessus_ATCC_1997 ---PIWWQEQPDGVGG-FNDGGYWVVTKHKDVKEVSLRSDVFSSWENTAI
Mflv_3290|M.gilvum_PYR-GCK ---PIWWNEQPDG-IGGFGDGGFWVVTKHHDVKEISKRSDVFSSEVKTAL
Mvan_3012|M.vanbaalenii_PYR-1 ---PIWWNEQTSGGAGPFGDGGYWVVTKHRDVKEISKHSEVFSSQQKTAL
MMAR_2783|M.marinum_M ---PIWWNEQPTG-CGGFDDGGFWVVTKHKDVKEISLRSDVFSSLQKTAL
MAV_2811|M.avium_104 ---PIWWNEQPIG-AGGFDDGGFWVVTKHKDVKEVSLRSDVFSSLQKTAL
MSMEG_3524|M.smegmatis_MC2_155 ---PIWWNAQPIG-AGGFDDGGFWVVTKHKDVKEISLRSDVFSSLEKTAL
TH_1956|M.thermoresistible__bu ---PIWWNEQPMG-AGGFDDGGFWLVTRHSDVKEVSRRSDVFSSQRKTAL
MLBr_02088|M.leprae_Br4923 DPFPVYRALIDYG-PMQLPGMPLTVFSSFSDCDEALRHPLSASDRLKATL
*:: * : . .: * .* :. *. : .:
Mb3575c|M.bovis_AF2122/97 PRFKNDIAREDIEVQRFVMLNMDAPHHTRLRKIISRGFTPRAVGRLHDEL
Rv3545c|M.tuberculosis_H37Rv PRFKNDIAREDIEVQRFVMLNMDAPHHTRLRKIISRGFTPRAVGRLHDEL
MUL_4106|M.ulcerans_Agy99 PRFKNDIAREDIDVQRFVMLNMDAPHHTRLRKIISRGFTPRAIGRLHDEL
MAB_0613|M.abscessus_ATCC_1997 PRFQDDITREAIELQRYVMLNMDAPHHTRLRKIISRGFTPRAIGRLRDEL
Mflv_3290|M.gilvum_PYR-GCK PRYPEGSTGEQIETGSLVLLNMDAPRHTHLRKIISRGFTPRAVERLRDDL
Mvan_3012|M.vanbaalenii_PYR-1 PRYPEGSTTEQVETGSLVLLNMDAPRHTHLRKIISRGFTPRAVERLREDL
MMAR_2783|M.marinum_M PRYKDGTVDEQIERGKFVLLNMDAPQHTRLRKIVSRAFTPRAVERLRDDL
MAV_2811|M.avium_104 PRYKDGTVAEQVERGKFVLLNMDAPQHTRLRKIISRAFTPRAVERLRDDL
MSMEG_3524|M.smegmatis_MC2_155 PRYPEGTVQDQIEQGRFVLLNMDAPHHTHLRKIISRAFTPRAVERLRDDL
TH_1956|M.thermoresistible__bu PRYRDGTVAEQIDRGKYVLLNMDAPHHTRLRKIISRAFTPRAVERLRDEL
MLBr_02088|M.leprae_Br4923 AQQAIAAGAEPRPFYASSFMFLDPPDHTRLRKLVSKAFAPKVVQALEGDI
.: : :: :*.* **:***::*:.*:*:.: *. ::
Mb3575c|M.bovis_AF2122/97 QERAQKIAAEAAAAGSGDFVEQVSCELPLQAIAGLLGVPQEDRGKLFHWS
Rv3545c|M.tuberculosis_H37Rv QERAQKIAAEAAAAGSGDFVEQVSCELPLQAIAGLLGVPQEDRGKLFHWS
MUL_4106|M.ulcerans_Agy99 NDRAQNIAKAAAAAGSGDFVEQVSCELPLQAIAGLLGIPQEDRGKLFDWS
MAB_0613|M.abscessus_ATCC_1997 NERAQEIAKAAAASGTGDFVEQVSCELPLQAIAGLLGVPIEDRGKLFNWS
Mflv_3290|M.gilvum_PYR-GCK NARAQNIAKTAASAGSGDFVEQVSCELPLQAIAGLLGVPVDDRKKLFDWS
Mvan_3012|M.vanbaalenii_PYR-1 AQRAHNIAKSAAAAGAGDFVEQVSCELPLQAIAGLLGVPLEDRKKLFDWS
MMAR_2783|M.marinum_M RERARRIVEAAAAEGRGDFVEQVSCELPLQAIAGLMGVPQEDRKKLFHWS
MAV_2811|M.avium_104 RERARRIVEAAAAEGSGDFVEQVSCELPLQAIASLMGVPQEDRKKLFHWS
MSMEG_3524|M.smegmatis_MC2_155 AERARAIVRAAAEEGSGDFVEQVACELPLQAIAGLMGVPQEDRRKLFDWS
TH_1956|M.thermoresistible__bu NERAHRIVMEAAAAGSGDFVEQVACELPLQAIAGLMGVPQEDRKKLFDWS
MLBr_02088|M.leprae_Br4923 AALVDSLLDKGAAAGQFDVIADLAFPLAVAVICRLLGVPYEDAPEFGRVS
. : .* * *.: ::: *.: .*. *:*:* :* :: *
Mb3575c|M.bovis_AF2122/97 NEMTGNED--------PEYAHIDPKASSAELIGYAMKMAEEKAKNPADDI
Rv3545c|M.tuberculosis_H37Rv NEMTGNED--------PEYAHIDPKASSAELIGYAMKMAEEKAKNPADDI
MUL_4106|M.ulcerans_Agy99 NEMTGTED--------PEFAHIDAKASSVELIGYAMKMAEEKAKNPGDDI
MAB_0613|M.abscessus_ATCC_1997 NEMTSYDD--------PEYADIDPAASSMEILAYSMEMAKQKAENPGEDI
Mflv_3290|M.gilvum_PYR-GCK NQMVSDDD--------PEYAHYDNRNAATELIMYAMQLAAVRAEQPGEDI
Mvan_3012|M.vanbaalenii_PYR-1 NQMVSDDD--------PEFAHYDNRNAATELIMYAMQLAALRAEQPGEDI
MMAR_2783|M.marinum_M NEMVGDQD--------PEFATNDALTASVELIMYGMQMAADRAKNPGQDL
MAV_2811|M.avium_104 NEMVGDQD--------PEFASNDAITASVELIMYGMQMAADRAKNPGEDL
MSMEG_3524|M.smegmatis_MC2_155 NQMVGNQD--------PEFVANDGASAAVELITYGMQLAAQRAAAPGDDL
TH_1956|M.thermoresistible__bu NQMVGDQD--------PEFADNDAIGASIELIGYGMQLAADRQRNPRDDL
MLBr_02088|M.leprae_Br4923 ALLVQSVDPFITITGEPPEATEERLRAGVWLRDYLEQLVKCRRGTPGEDL
:. * * . : :. : * ::. : * :*:
Mb3575c|M.bovis_AF2122/97 VTQLIQADIDGE-KLSDDEFGFFVVMLAVAGNETTRNSITQGMMAFAEHP
Rv3545c|M.tuberculosis_H37Rv VTQLIQADIDGE-KLSDDEFGFFVVMLAVAGNETTRNSITQGMMAFAEHP
MUL_4106|M.ulcerans_Agy99 VTQLIQADIDGE-KLSDDEFGFFVVMLAVAGNETTRNSITQGMMAFADNP
MAB_0613|M.abscessus_ATCC_1997 VTTLINAEVEGEGKLSDDEFGFFVIMLAVAGNETSRNSITQGMMAFTQFP
Mflv_3290|M.gilvum_PYR-GCK VTKLIEADVDGH-KLSDDEFGFFMVLLAVAGNETTRNSITHGMIAFTEHP
Mvan_3012|M.vanbaalenii_PYR-1 VTKLIEADVDGH-KLTDDEFGFFMVLLAVAGNETTRNSITHGMIAFTEHP
MMAR_2783|M.marinum_M VTKLVEADIDGH-KLSDDEFGFFVILLAVAGNETTRNSITQGMMAFTDFP
MAV_2811|M.avium_104 VTKLVQADIDGH-KLSDDEFGFFVILLAVAGNETTRNSITQGMMAFTDFP
MSMEG_3524|M.smegmatis_MC2_155 VTKLVQADVEGH-KLSDDEFGFFVVLLAVAGNETTRNSITQGMMAFTDHR
TH_1956|M.thermoresistible__bu VTTLVQADVDGH-KLSDDEFGFFVILLAVAGNETTRNSITQGMMAFTEHP
MLBr_02088|M.leprae_Br4923 ISRLIELDESGD-QLTEEEIIATCGLLLVAGHETTVNLIANAVLAMLRNP
:: *:: : .*. :*:::*: :* ***:**: * *::.::*:
Mb3575c|M.bovis_AF2122/97 DQWELYKK--VRPETAADEIVRWATPVTAFQRTALRDYELSGVQIKKGQR
Rv3545c|M.tuberculosis_H37Rv DQWELYKK--VRPETAADEIVRWATPVTAFQRTALRDYELSGVQIKKGQR
MUL_4106|M.ulcerans_Agy99 EQWELYKR--ERPGTAADEIVRWATPVTSFQRTALEDYELSGVQIKKGQR
MAB_0613|M.abscessus_ATCC_1997 EQWELYKK--ERPETAADEIVRWATPVTSFQRTALEDTELDGVKIKKGQR
Mflv_3290|M.gilvum_PYR-GCK DQWELYKR--ERPITAVDEIVRWATPVTSFQRTALTDYELSGVQITKGQR
Mvan_3012|M.vanbaalenii_PYR-1 DQWELFKR--ERPATAVDEIVRWATPVTSFQRTALRDYELSGVQIKKGQR
MMAR_2783|M.marinum_M DQWELYKR--ERPVTTADEIVRWATPVTSFQRTALQDYELSGVRIKKGQR
MAV_2811|M.avium_104 DQWELFKR--ERPATAADEIVRWATPVTSFQRTALQDYELSGVKIKKGQR
MSMEG_3524|M.smegmatis_MC2_155 DQWELFKR--ERPVTTADEIVRWATPVTSFQRTALADTEVSGVRIKKGQR
TH_1956|M.thermoresistible__bu DQWELFKE--QRPPTTADEIVRWATPVTSFQRTALADVELSGVQIKAGQR
MLBr_02088|M.leprae_Br4923 SQWKALSSNPQRAPLVVEETLRYDPAIHLIGRVAAKDMTIGQTTLTEGDT
.**: . *. ..:* :*: ..: : *.* * :. . :. *:
Mb3575c|M.bovis_AF2122/97 VVMFYRSANFDEEVFQDPFTFNILRNPNPHVGFGGTGAHYCIGANLARMT
Rv3545c|M.tuberculosis_H37Rv VVMFYRSANFDEEVFQDPFTFNILRNPNPHVGFGGTGAHYCIGANLARMT
MUL_4106|M.ulcerans_Agy99 VLMFYRSANFDEEVFEDPFSFNILRNPNPHVGFGGTGAHYCIGANLARMT
MAB_0613|M.abscessus_ATCC_1997 VVMMYRSANFDEDVFEDPFSFNIMRNPNPHMGFGGSGAHYCIGANLARLT
Mflv_3290|M.gilvum_PYR-GCK VVMSYRSANFDEEVFDDPFTFDILRDPNPHVGFGGTGAHYCIGANLARMT
Mvan_3012|M.vanbaalenii_PYR-1 VVMSYRSANFDEEVFDDPFTFDIMRDPNPHVGFGGTGAHYCIGANLARMT
MMAR_2783|M.marinum_M VVMFYRSANFDEDVFDDPYTFNILRDPNPHVGFGGTGAHYCIGANLARMT
MAV_2811|M.avium_104 VVMFYRSANFDEDVFDDPFTFNILRDPNPHVGFGGTGAHYCIGANLARMT
MSMEG_3524|M.smegmatis_MC2_155 VVMFYRSANFDEDVFTDPYRFDILRDPNPHVGFGGTGAHYCIGANLARMT
TH_1956|M.thermoresistible__bu VVMCYRSANFDDTVFDDPYTFNILRDPNPHLGFGGTGAHYCIGANLARLT
MLBr_02088|M.leprae_Br4923 MVLLLAAANRDPAVYSRPDEFDPDRPSSRHLAFA-VGSHFCLGAALARLE
::: :** * *: * *: * .. *:.*. *:*:*:** ***:
Mb3575c|M.bovis_AF2122/97 INLIFNAVADHMPDLKPISAPERLRSGWLNGIKHWQVDYTGR----CPVA
Rv3545c|M.tuberculosis_H37Rv INLIFNAVADHMPDLKPISAPERLRSGWLNGIKHWQVDYTGR----CPVA
MUL_4106|M.ulcerans_Agy99 INLIFNAVADHMPDLKPIAAPERLRSGWLNGIKHWQVDYTGK----CPVS
MAB_0613|M.abscessus_ATCC_1997 INLMFNAIADHMPNLAPAGDPKRLQSGWLNGIKHWQVDFTGAS--GCPVL
Mflv_3290|M.gilvum_PYR-GCK IDLMFNAIADHMPDLSAIGSPDRLRSGWLNGIKHWQVDYSG---RGCPVA
Mvan_3012|M.vanbaalenii_PYR-1 IDLMFNAIADHLPDLSSAGTPDRLRSGWLNGIKHWQVDYTGP--SGCPVA
MMAR_2783|M.marinum_M IDLMFNAIADVMPDLESISQPERLRSGWLNGIKHWQVDYHSDSSGKCPVA
MAV_2811|M.avium_104 IDLMFNAIADAMPDLESIGKPERLRSGWLNGIKHWQVDYHTNGSSKCPVA
MSMEG_3524|M.smegmatis_MC2_155 IDLIFNAIADEMPDLTPISEPVRLRSGWLNGIKHWQVDYRGDAAKHRAAQ
TH_1956|M.thermoresistible__bu IDLIFAAIAEHLPDLRPLGPPERLRSGWLNGIKHWPVDYGAPRVPTGAAG
MLBr_02088|M.leprae_Br4923 ATVTLSAISARFPQVQLAGELVYKPNVAMRGMSALPVQV-----------
: : *:: :*:: . . :.*:. *:
Mb3575c|M.bovis_AF2122/97 H------
Rv3545c|M.tuberculosis_H37Rv H------
MUL_4106|M.ulcerans_Agy99 H------
MAB_0613|M.abscessus_ATCC_1997 Q------
Mflv_3290|M.gilvum_PYR-GCK H------
Mvan_3012|M.vanbaalenii_PYR-1 H------
MMAR_2783|M.marinum_M H------
MAV_2811|M.avium_104 H------
MSMEG_3524|M.smegmatis_MC2_155 ASSQADR
TH_1956|M.thermoresistible__bu PQRCG--
MLBr_02088|M.leprae_Br4923 -------