For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
LVLRALIAVVAVAFAAVGLVAPAGAAPEDGQWDPTLPKLISAGAPGDPLAIANASLEATAQATQVTMGLG RKFLASLGLVEDTPAAAGVAPGRVRGPQAIEYVIRRGGTQMGVPYSWGGGKPNGPSRGIDSGANTVGFDC SGFTQFSFAGVGVLIPKYSGDQYNTGRKVPVNQAKRGDLLFWGPGGSQHVALYLGGGKMLEASGSAGKVT VSPVRMSGLQPYAARIIES
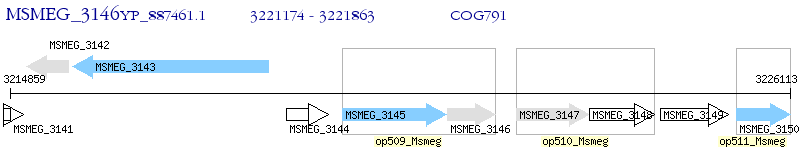
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_3146 | - | - | 100% (229) | invasin 1 |
| M. smegmatis MC2 155 | MSMEG_3145 | - | 2e-63 | 55.56% (207) | secreted cell wall-associated hydrolase |
| M. smegmatis MC2 155 | MSMEG_3477 | - | 1e-25 | 34.91% (232) | inv protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1514 | - | 5e-83 | 66.22% (225) | invasion protein |
| M. gilvum PYR-GCK | Mflv_3662 | - | 5e-91 | 71.81% (227) | NLP/P60 protein |
| M. tuberculosis H37Rv | Rv1478 | - | 5e-83 | 66.22% (225) | invasion protein |
| M. leprae Br4923 | MLBr_01811 | - | 2e-75 | 60.43% (235) | putative exported p60 protein homologue |
| M. abscessus ATCC 19977 | MAB_2727c | - | 2e-61 | 58.33% (204) | invasion protein Inv2 |
| M. marinum M | MMAR_2285 | iipB | 2e-81 | 64.47% (228) | invasion and intracellular persistence protein, IipB |
| M. avium 104 | MAV_3300 | - | 2e-79 | 63.91% (230) | invasin 1 |
| M. thermoresistible (build 8) | TH_4058 | - | 1e-103 | 79.82% (228) | PUTATIVE NLP/P60 protein |
| M. ulcerans Agy99 | MUL_1487 | inv2 | 1e-81 | 64.47% (228) | invasion protein Inv2 |
| M. vanbaalenii PYR-1 | Mvan_2748 | - | 1e-107 | 84.14% (227) | NLP/P60 protein |
CLUSTAL 2.0.9 multiple sequence alignment
Mb1514|M.bovis_AF2122/97 -------MRHTRFHPIKLAWITAVVAGLMVGVATP--ADAEPGQWDP---
Rv1478|M.tuberculosis_H37Rv -------MRHTRFHPIKLAWITAVVAGLMVGVATP--ADAEPGQWDP---
MMAR_2285|M.marinum_M MSSYRGPMRYKRFRLLGLAWITLAVAALLVVVSAP--ADADPGGWDP---
MUL_1487|M.ulcerans_Agy99 -------MRYKRFRLLGLAWITLAVAALMVVVSAP--ADADPGGWDP---
MLBr_01811|M.leprae_Br4923 -------MRHKNFRLINLAGLTAMVAGLIIVVTIP--ATADPGQWDP---
MAV_3300|M.avium_104 -------MRRNRFRLIVFAWITAMVTGLMFSVAPTPAALADPGEWDP---
MSMEG_3146|M.smegmatis_MC2_155 -------------MVLRALIAVVAVAFAAVGLVAPAGAAPEDGQWDP---
TH_4058|M.thermoresistible__bu ---VVILLRSKTFRLLRPLIPAILGLGVLIAAAGPATAAPDDGQWDP---
Mflv_3662|M.gilvum_PYR-GCK ------------MALRRVLVALLAAAALLAGAVAPASAQPGGGAWDP---
Mvan_2748|M.vanbaalenii_PYR-1 -----------MIALRRFVIALVTAAALLAGAVAPASAQPESGQWDP---
MAB_2727c|M.abscessus_ATCC_199 ---------MQGLFLLRSAIAVTIAIALVLFMGVPR-AAADDNPLGPNVG
. * . . .*
Mb1514|M.bovis_AF2122/97 -----TLPALVSAGAPGDPLAVANASLQAT-AQATQTTLDLGRQFLGGLG
Rv1478|M.tuberculosis_H37Rv -----TLPALVSAGAPGDPLAVANASLQAT-AQATQTTLDLGRQFLGGLG
MMAR_2285|M.marinum_M -----TLPATASAGAPGDALAMANASLQAT-AQATQTTLDLGRQFLGGLG
MUL_1487|M.ulcerans_Agy99 -----TLPATASAGAPGDALAMANASLQAT-AQATQTTLDLGRQFLGGLG
MLBr_01811|M.leprae_Br4923 -----TLPAAVSAGAPGDPLAVANASLQVT-AQATQTTLDLGKQFLGRLG
MAV_3300|M.avium_104 -----TLPAQISAGAPGDPLAVANASLQAT-AQATQTTLNLGKQFLGGLG
MSMEG_3146|M.smegmatis_MC2_155 -----TLPKLISAGAPGDPLAIANASLEAT-AQATQVTMGLGRKFLASLG
TH_4058|M.thermoresistible__bu -----TLPKLISAGAPGDPLAIANASLAAT-AQATQVTMDLGRKFLASIG
Mflv_3662|M.gilvum_PYR-GCK -----LLPMRPSAGAPGDALAIANASLQVT-AQATQATMDMGRKFLQTLG
Mvan_2748|M.vanbaalenii_PYR-1 -----TLPKLISAGAPGDPLAIANASLAAT-AQATQVTMDLGRKFLQTIG
MAB_2727c|M.abscessus_ATCC_199 TAFLNALGLNPTGGQYDPTLPFAGPSQGADPAKIMNGVMGVGQTALGALG
* :.* . .*..*..* . *: : .:.:*: * :*
Mb1514|M.bovis_AF2122/97 INLGGPAASAPSAATTG-ASRIPRANARQAVEYVIRRAGSQMGVPYSWGG
Rv1478|M.tuberculosis_H37Rv INLGGPAASAPSAATTG-ASRIPRANARQAVEYVIRRAGSQMGVPYSWGG
MMAR_2285|M.marinum_M INLGG--SPAPSAANTG-ANRIPRVYGQQALEYVIRRGGSQMGVPYSWGG
MUL_1487|M.ulcerans_Agy99 INLGG--SPAPSAANTG-ANRIPRVYGRQALEYVIRRGGSQMGVPYSWGG
MLBr_01811|M.leprae_Br4923 INFGG-DPSNAATTSSNPGGRIPRVYGGQAIEYVIKRGGAQIGVPYSWGG
MAV_3300|M.avium_104 INLGGNDAPAAAATPSNPGGKIPRVYGRQAIEYVIKRMGSQMGVPYSWGG
MSMEG_3146|M.smegmatis_MC2_155 LVEDTPAAAGVAPGRVR---------GPQAIEYVIRRGGTQMGVPYSWGG
TH_4058|M.thermoresistible__bu LGGSQPVSS-VAPGRVR---------GPQAIEYVIRRGASQLGVPYSWGG
Mflv_3662|M.gilvum_PYR-GCK LVP-ADAPT-VAPGSVR---------GPAAVEYVIKRGAARLGTPYSWGG
Mvan_2748|M.vanbaalenii_PYR-1 LAP-KDAPAGVAPGVVR---------GPAAIEYAIRRGGTQIGTPYSWGG
MAB_2727c|M.abscessus_ATCC_199 IGGNSASAGAGRPLVY----------GRAAVERVIQRGGTQLGVPYSWGG
: . . . *:* .*:* .:::*.******
Mb1514|M.bovis_AF2122/97 GSLQGPSKGVDSGANTVGFDCSGLVRYAFAGVGVLIPRFSGDQYNAGRHV
Rv1478|M.tuberculosis_H37Rv GSLQGPSKGVDSGANTVGFDCSGLVRYAFAGVGVLIPRFSGDQYNAGRHV
MMAR_2285|M.marinum_M GSLQGPSKGVDSGANTVGFDCSGLMRYAFAGVGVLIPRYSGDQYNAGRHI
MUL_1487|M.ulcerans_Agy99 GSLQGPSKGVDSGANTVGFDCSGLMRYAFAGVGVLIPRYSGDQYNAGRHI
MLBr_01811|M.leprae_Br4923 GSLQGPSKGVGDGANITGFDCSGLMRYAFAGVGVLIPRFSGDQYNAGRHL
MAV_3300|M.avium_104 GSLDGPSKGVGDGANITGFDCSGLMRYGFAGVGVLIPRFSGDQYNAGRHI
MSMEG_3146|M.smegmatis_MC2_155 GKPNGPSRGIDSGANTVGFDCSGFTQFSFAGVGVLIPKYSGDQYNTGRKV
TH_4058|M.thermoresistible__bu GKPSGPSTGVGSGANTVGFDCSGFTQFAFAGVGVLIPKYSGDQYNTGRKV
Mflv_3662|M.gilvum_PYR-GCK GKPTGPSLGVERGANTVGFDCSGFTQYSFAGVGVLIPKYSGDQFNTGRRV
Mvan_2748|M.vanbaalenii_PYR-1 GKPNGPSRGIDSGANTVGYDCSGFTQFSYAGVGVLIPKYSGDQYNTGRKV
MAB_2727c|M.abscessus_ATCC_199 GTVRGPSGGVDYDSGKVGYDCSGFTMFSYAAAGVKLPKYSGDQYNAGQKV
*. *** *: .:. .*:****: :.:*..** :*::****:*:*:::
Mb1514|M.bovis_AF2122/97 PPAEAKRGDLIFYGPGGGQHVTLYLGNGQMLEASGSAGKVTVSPVRKAGM
Rv1478|M.tuberculosis_H37Rv PPAEAKRGDLIFYGPGGGQHVTLYLGNGQMLEASGSAGKVTVSPVRKAGM
MMAR_2285|M.marinum_M SPDQARRGDLIFYGPGGSQHVTMYLGNGQMLEASGSAGKVTVSPVRKAGM
MUL_1487|M.ulcerans_Agy99 SPDQARRGDLIFYGPGGSQHVTMYLGNGQMLEASGSAGKVTVSPVRKAGM
MLBr_01811|M.leprae_Br4923 TPDQAKRGDLIFYGPGGGQHVTMYLGNGQMLEASSSVGKVTVSSVRKAGM
MAV_3300|M.avium_104 PQDQARRGDLIFYGPGGSQHVTMYLGNGQMLEASGSAGKVTVSPVRKPGM
MSMEG_3146|M.smegmatis_MC2_155 PVNQAKRGDLLFWGPGGSQHVALYLGGGKMLEASGSAGKVTVSPVRMSGL
TH_4058|M.thermoresistible__bu PVSQAKRGDLLFWGPGGSQHVALYLGNGQMLEASGSAGKVVAGPVRTAGL
Mflv_3662|M.gilvum_PYR-GCK PVQQAKRGDLLFWGPGGSQHVAIYLGNGQMLESGGTADKVVVSKVRMSGL
Mvan_2748|M.vanbaalenii_PYR-1 PVQQAKRGDLLFWGPGGSQHVAIYLGGGKMLESSGSAGKVTVSPVRMSGL
MAB_2727c|M.abscessus_ATCC_199 PVAQAKRGDLLFYGPGGSQHVVIYLGNGQMLEASGSAGKVTVSPVRTGGM
. :*:****:*:****.***.:***.*:***:..:..**... ** *:
Mb1514|M.bovis_AF2122/97 TPFVTRIIEY
Rv1478|M.tuberculosis_H37Rv TPFVTRIIEY
MMAR_2285|M.marinum_M TPFVTRIIEY
MUL_1487|M.ulcerans_Agy99 TPFVTRIIEY
MLBr_01811|M.leprae_Br4923 TPFVTRIIES
MAV_3300|M.avium_104 TPFLTRIIEY
MSMEG_3146|M.smegmatis_MC2_155 QPYAARIIES
TH_4058|M.thermoresistible__bu QPYAARIIET
Mflv_3662|M.gilvum_PYR-GCK QPTVQRIIES
Mvan_2748|M.vanbaalenii_PYR-1 QPFVARIIES
MAB_2727c|M.abscessus_ATCC_199 TPYAVRIIAW
* ***