For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
ERMTGQANSRRGELLGLAAVMFAERGLRATTVRDIADGAGILSGSLYHHFASKEEMVDELLREFLDWLFA RYLEIVDGEPNPLERLKGLFLASFEAIEHRHAQVVIYQDEAQRLSSQPRFSYLEDRNKEQRKMWVDVLNQ GIEEGYFRPDLDVDLVYRFIRDTTWVSVRWYKPGGALTAQQVGQKYLAIVLGGIQKEGV*
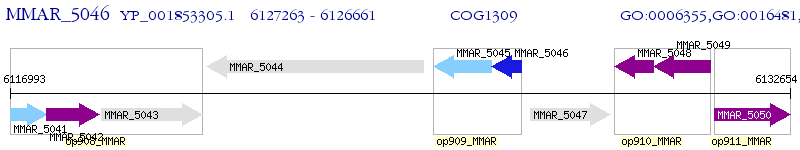
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. marinum M | MMAR_5046 | - | - | 100% (200) | TetR family transcriptional regulator |
| M. marinum M | MMAR_4325 | - | 6e-12 | 52.38% (63) | TetR family transcriptional regulator |
| M. marinum M | MMAR_0381 | - | 3e-10 | 23.71% (194) | TetR family transcriptional regulator |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3587c | - | 1e-101 | 88.50% (200) | TetR family transcriptional regulator |
| M. gilvum PYR-GCK | Mflv_1483 | - | 6e-83 | 77.42% (186) | TetR family transcriptional regulator |
| M. tuberculosis H37Rv | Rv3557c | - | 1e-101 | 88.50% (200) | TetR family transcriptional regulator |
| M. leprae Br4923 | MLBr_00815 | - | 4e-09 | 33.70% (92) | putative TetR-family transcriptional regulator |
| M. abscessus ATCC 19977 | MAB_0599 | - | 4e-74 | 72.04% (186) | TetR family transcriptional regulator |
| M. avium 104 | MAV_0603 | - | 5e-95 | 85.42% (192) | transcriptional regulator, TetR family protein |
| M. smegmatis MC2 155 | MSMEG_6009 | - | 4e-84 | 76.29% (194) | transcriptional regulator, TetR family protein |
| M. thermoresistible (build 8) | TH_2070 | - | 2e-89 | 83.87% (186) | PROBABLE TRANSCRIPTIONAL REGULATORY PROTEIN (PROBABLY |
| M. ulcerans Agy99 | MUL_4120 | - | 1e-104 | 99.46% (185) | TetR family transcriptional regulator |
| M. vanbaalenii PYR-1 | Mvan_5283 | - | 1e-90 | 83.33% (192) | TetR family transcriptional regulator |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_1483|M.gilvum_PYR-GCK ----------------------------MDRALPS----RRDELLQLAAT
MSMEG_6009|M.smegmatis_MC2_155 ----------------------------MAPDTPSQPASRRDELLQLAAT
MMAR_5046|M.marinum_M -----------------------------MERMTGQANSRRGELLGLAAV
MUL_4120|M.ulcerans_Agy99 --------------------------------------------MGLAAV
Mb3587c|M.bovis_AF2122/97 -----------------------------MDRVAGQVNSRRGELLELAAA
Rv3557c|M.tuberculosis_H37Rv -----------------------------MDRVAGQVNSRRGELLELAAA
MAV_0603|M.avium_104 --------------------------------MTGPANSRRDELLELAAA
TH_2070|M.thermoresistible__bu ----------------------------MTNSGPP---TRRDELLELAAQ
Mvan_5283|M.vanbaalenii_PYR-1 ----------------------------MSSGQAPQANTRRDELLQLAAT
MAB_0599|M.abscessus_ATCC_1997 -----------------MDKVSEVRESDALRAGGSPQPSRRDELLSLAAE
MLBr_00815|M.leprae_Br4923 MNNLAKAAQRRTVRPIDRGRLSDDIATPNQRSNRLSRDKRRGQLLIVASD
: :*:
Mflv_1483|M.gilvum_PYR-GCK MFADRGLRATTVRDIADSAGILSGSLYHHFKSKEQMVEEVLKDFLDWLFT
MSMEG_6009|M.smegmatis_MC2_155 MFADRGLKATTVRDIADSAGILSGSLYHHFKSKEQMVEEVLRDFLDWLFG
MMAR_5046|M.marinum_M MFAERGLRATTVRDIADGAGILSGSLYHHFASKEEMVDELLREFLDWLFA
MUL_4120|M.ulcerans_Agy99 MFAERGLRATTVRDIADGAGILSGSLYHHFASKEEMVDELLREFLDWLFA
Mb3587c|M.bovis_AF2122/97 MFAERGLRATTVRDIADGAGILSGSLYHHFASKEEMVDELLRGFLDWLFA
Rv3557c|M.tuberculosis_H37Rv MFAERGLRATTVRDIADGAGILSGSLYHHFASKEEMVDELLRGFLDWLFA
MAV_0603|M.avium_104 MFAERGLRATTVRDIADSAGILSGSLYHHFSSKEEMVDEVLRSFLDWLFA
TH_2070|M.thermoresistible__bu MFAERGLKATTVRDIADSAGILSGSLYHHFASKEEMVDEVLRGFLDWLFD
Mvan_5283|M.vanbaalenii_PYR-1 MFAERGLRATTVRDIADSAGILSGSLYHHFSSKEEMVDEVLRGFLDWLFD
MAB_0599|M.abscessus_ATCC_1997 MFAERGLRATTVRDIADAAGILSGSLYHHFDSKEAIVDELLRDFLEGLFA
MLBr_00815|M.leprae_Br4923 VFVDHGYHAAGMDEIADRAGVSKPVLYQHFSSKLELYLAVLERHMNNLVS
:*.::* :*: : :*** **: . **:** ** : :*. .:: *.
Mflv_1483|M.gilvum_PYR-GCK RYDEIVAATTDPLERVSGLFMVSFEAIEHRHAQVVIYQDEAKRLNSLPQF
MSMEG_6009|M.smegmatis_MC2_155 RYQQILDTATSPLEKLTGLFMASFEAIEHRHAQVVIYQDEAKRLSDLPQF
MMAR_5046|M.marinum_M RYLEIVDGEPNPLERLKGLFLASFEAIEHRHAQVVIYQDEAQRLSSQPRF
MUL_4120|M.ulcerans_Agy99 RYLEIVDGEPNPLERLKGLFLASFEAIEHRHAQVVIYQDEAQRLSSQPRF
Mb3587c|M.bovis_AF2122/97 RYRDIVDSTANPLERLQGLFMASFEAIEHHHAQVVIYQDEAQRLASQPRF
Rv3557c|M.tuberculosis_H37Rv RYRDIVDSTANPLERLQGLFMASFEAIEHHHAQVVIYQDEAQRLASQPRF
MAV_0603|M.avium_104 RYREIIDSESNPLERLKGLFMASFEAIEHRHAQVVIYQDEAKRLLSQPRF
TH_2070|M.thermoresistible__bu RYRSIVETEPNPLERFKGLFMASFEAIEHRHAQVVIYQDEAKRLSSQPRF
Mvan_5283|M.vanbaalenii_PYR-1 RYQQIVATEPNPLERLKGLFMASFEAIETRHAEVVIYQDEAKRLSAQGRF
MAB_0599|M.abscessus_ATCC_1997 RYREIAAANLNPVDTLKGLFLASFDAIETKHAQVAIYQAEAPRLLSQERF
MLBr_00815|M.leprae_Br4923 SVQQALRTTTNNRQRLHAAVQAFFDFIEHDGQGYRLIFENDN--ITEPRV
. . : . . . . *: ** : : :.
Mflv_1483|M.gilvum_PYR-GCK GFLDDRNRQQRRMWVEVLKQGIAEGSFRPDIDVDLVYRFIRDTTWVSVRW
MSMEG_6009|M.smegmatis_MC2_155 DFVETRNKEQRKMWVDILQEGVADGSFRPDLDVDLVYRFIRDTTWVSVRW
MMAR_5046|M.marinum_M SYLEDRNKEQRKMWVDVLNQGIEEGYFRPDLDVDLVYRFIRDTTWVSVRW
MUL_4120|M.ulcerans_Agy99 SYLEDRNKEQRKMWLDVLNQGIEEGYFRPDLDVDLVYRFIRDTTWVSVRW
Mb3587c|M.bovis_AF2122/97 SYIEDRNKQQRKMWVDVLNQGIEEGYFRPDLDVDLVYRFIRDTTWVSVRW
Rv3557c|M.tuberculosis_H37Rv SYIEDRNKQQRKMWVDVLNQGIEEGYFRPDLDVDLVYRFIRDTTWVSVRW
MAV_0603|M.avium_104 SYIEDMNRQQRKMWLEVLNQGVDEGYFRPDLDVDLVYRFIRDTTWVSVRW
TH_2070|M.thermoresistible__bu AYVEQRNREQRKMWVDVLTQGIEEGYFRSDIDVDLVYRFIRDTTWVSVRW
Mvan_5283|M.vanbaalenii_PYR-1 AYVEERNKEQRKMWVDVLTQGIDEGYFRPDVDVDLVYRFIRDTTWVSVRW
MAB_0599|M.abscessus_ATCC_1997 AYLNELNTEQRQLWLEVLRDGIAQGYFRPDLDVAVVYRFIRDATWVSVRW
MLBr_00815|M.leprae_Br4923 AAQVRVATESCIDAVFALIS--ADSRLDPHRARMVAVGLVGISVDCARYW
:. : * . :. : .. :. :: :. : *
Mflv_1483|M.gilvum_PYR-GCK YQPGGPLTAEQVGRQYLAIVLGGITE--GSK-----
MSMEG_6009|M.smegmatis_MC2_155 YKPGGPLSAEQVGQQYLAIVLGGITQSQGDKHA---
MMAR_5046|M.marinum_M YKPGGALTAQQVGQKYLAIVLGGIQKEGV-------
MUL_4120|M.ulcerans_Agy99 YKPGGALTAQQVGQKYLAIVLGGIQKEGV-------
Mb3587c|M.bovis_AF2122/97 YRPGGPLTAQQVGQQYLAIVLGGITKEGV-------
Rv3557c|M.tuberculosis_H37Rv YRPGGPLTAQQVGQQYLAIVLGGITKEGV-------
MAV_0603|M.avium_104 YQPGGPLTAEQVGQQYLAIVLGGITVDAKKD-----
TH_2070|M.thermoresistible__bu YRPGGPLTAEEVGRQYLSIVLGGITAGAN-------
Mvan_5283|M.vanbaalenii_PYR-1 YQPGGPLTAQQVGRQYLSIVLGGITAEPTSKESTNG
MAB_0599|M.abscessus_ATCC_1997 FRPGGSRTAQEVAEQYLSIVLGGITVGSFSNSPEK-
MLBr_00815|M.leprae_Br4923 LDTNRPISKSDAVEGTVQFAWGGLSHVPLTRL----
.. . : .:. . : :. **: