For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MLEAILSKPSNRRAPIEMGEGEPVLLLHPFMLSQSVWETVAEQLADAGYEVFAPTYPGHNGGGPMPKLPP FSQRSVDVYLDQIEQIMDQKGWETAHLVGNSLGGWIALGLALRGRARSVTAIAPAGGWTKYSPLKAEIVV KFAAMLPWLAFSRLLGRRAAGLPFAKFATARMLCGETEALSLPDFHSIFEDITHCPAYFASLTSILPAPG LENLDTISVPAQLVLCEKDEIVPPGRFGKLIGEGLPGNARRITLAGMGHVPMLEAPGQVTEVITDLIDPL VSAKRSAASDAG
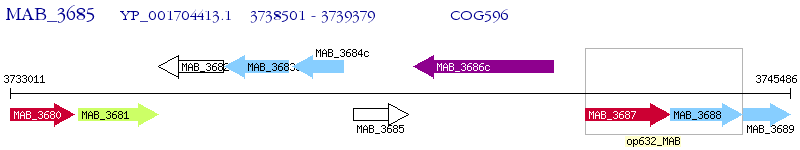
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. abscessus ATCC 19977 | MAB_3685 | - | - | 100% (292) | hypothetical protein MAB_3685 |
| M. abscessus ATCC 19977 | MAB_2946c | - | 5e-14 | 27.86% (280) | Alpha/beta hydrolase fold |
| M. abscessus ATCC 19977 | MAB_3034 | - | 2e-11 | 28.21% (280) | hydrolase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3370 | - | 1e-76 | 51.30% (269) | hypothetical protein Mb3370 |
| M. gilvum PYR-GCK | Mflv_4874 | - | 3e-68 | 50.19% (267) | alpha/beta hydrolase fold |
| M. tuberculosis H37Rv | Rv3338 | - | 5e-56 | 48.76% (201) | hypothetical protein Rv3338 |
| M. leprae Br4923 | MLBr_00685 | - | 6e-71 | 50.36% (280) | putative hydrolase |
| M. marinum M | MMAR_1187 | - | 3e-63 | 46.84% (269) | hypothetical protein MMAR_1187 |
| M. avium 104 | MAV_4312 | - | 2e-65 | 48.85% (262) | hydrolase, alpha/beta fold family protein, putative |
| M. smegmatis MC2 155 | MSMEG_1655 | - | 3e-68 | 48.21% (280) | hydrolase, alpha/beta fold family protein, putative |
| M. thermoresistible (build 8) | TH_1972 | - | 7e-68 | 47.16% (282) | CONSERVED HYPOTHETICAL PROTEIN |
| M. ulcerans Agy99 | MUL_1440 | - | 1e-47 | 43.46% (214) | hypothetical protein MUL_1440 |
| M. vanbaalenii PYR-1 | Mvan_1555 | - | 3e-64 | 44.60% (287) | alpha/beta hydrolase fold |
CLUSTAL 2.0.9 multiple sequence alignment
TH_1972|M.thermoresistible__bu ------------------------MSERAPIHLGSG----EPVLLLHPFM
Mvan_1555|M.vanbaalenii_PYR-1 ------------------------MNLREPIHLGSG----EPVLLLHPFM
MSMEG_1655|M.smegmatis_MC2_155 ------------------------MVERAPIHLGSRDGNTEPILLLHPFL
Mb3370|M.bovis_AF2122/97 --MPSPSTTGHHAACGTGGTGFSVGSMRSPIRVGSG----EPVLLLHPFL
Rv3338|M.tuberculosis_H37Rv --------------------------------------------------
MMAR_1187|M.marinum_M ------------------------------------------MLLLHPFL
MUL_1440|M.ulcerans_Agy99 --------------------------------------------------
MAV_4312|M.avium_104 -------------------------------------------MLLHGFL
MLBr_00685|M.leprae_Br4923 MPACQHQRHRPVVACGTRGTTFRVGSMRQPIHLGSG----EPVLLLHPFM
Mflv_4874|M.gilvum_PYR-GCK ------------------------MAPREPIHRGTG----EPIVLLHPFL
MAB_3685|M.abscessus_ATCC_1997 ---------------MLEAILSKPSNRRAPIEMGEG----EPVLLLHPFM
TH_1972|M.thermoresistible__bu MSQSVWKDVAPRIAG----------TGRYEVYAPTMPGHNGGV-KGRFF-
Mvan_1555|M.vanbaalenii_PYR-1 MSQNVWKDVAPQLAGVDVANPSAPATGRYEVLAPTMPGHNGGV-KGRFF-
MSMEG_1655|M.smegmatis_MC2_155 LSQSVWKYVAPQLAD----------TGRYEVFAPTMADHNGGP-RGPFL-
Mb3370|M.bovis_AF2122/97 MSQTVWEKVAQQLAD----------TGRFEVFAPTMAGHNGGPASGTRF-
Rv3338|M.tuberculosis_H37Rv --------------------------------------------------
MMAR_1187|M.marinum_M LSQAVWEQVAQRLAD----------TGRYEVFAPTMASHNGGPRAGTWF-
MUL_1440|M.ulcerans_Agy99 -----------------------------------MASHNGRPRAGTWF-
MAV_4312|M.avium_104 MSQTVWDPVAPRLAD----------TGRYEVFAPTMAGHNGGPYAGTWL-
MLBr_00685|M.leprae_Br4923 MSQTVWEIVAPQLAN----------TERYEVFAPTMAAHHGGPSASSWM-
Mflv_4874|M.gilvum_PYR-GCK CSQNVWRTVADQLAD----------TERFEVFAPTMVGHHGGPKSPTWL-
MAB_3685|M.abscessus_ATCC_1997 LSQSVWETVAEQLAD-----------AGYEVFAPTYPGHNGGGPMPKLPP
TH_1972|M.thermoresistible__bu ---LDTAELADDMERRLDELGWDTAHIVGNSLGGWVAFELERRGRARTLT
Mvan_1555|M.vanbaalenii_PYR-1 ---LDTAELADDMERRMDALGWDTAHVVGNSLGGWVAFELERRGRVRSLT
MSMEG_1655|M.smegmatis_MC2_155 ---LDVAHLADDIERRMDEVGWTTAHIVGNSLGGWVAFELERRGRARTLT
Mb3370|M.bovis_AF2122/97 ---LSSAVLADHVERQLDELGWETSHIVGNSLGGWVAFELERRGRARSVT
Rv3338|M.tuberculosis_H37Rv ---MSSAVLADHVERQLDELGWETSHIVGNSLGGWVAFELERRGRARSVT
MMAR_1187|M.marinum_M ---LSSAVLADHVERQMDELGWDSAHIVGNSLGGWVAFELERRGRARSVT
MUL_1440|M.ulcerans_Agy99 ---LSSAVLADHVERQMDELGWDSAHIVGNSLGGWVAFELERRGRARSVT
MAV_4312|M.avium_104 ---LSSSVLADHVERQLDRLGWDSAHLVGNSLGGWVAFELERRGRARSVT
MLBr_00685|M.leprae_Br4923 ---MSVSTLADHVEQQLDELGWDTAHLVGNSLGGWVVFELERRGRARSVT
Mflv_4874|M.gilvum_PYR-GCK ---LHTEVLVDDIEHRMDQLGWDTAHIVGNSLGGWVAFELERRGRARTLT
MAB_3685|M.abscessus_ATCC_1997 FSQRSVDVYLDQIEQIMDQKGWETAHLVGNSLGGWIALGLALRGRARSVT
*.:*: :* ** ::*:********:.: * ***.*::*
TH_1972|M.thermoresistible__bu GIAPAGGWHRWSPAKYEIVGKFVLGFPVWAVAKLLGRRAARLPIARQLAF
Mvan_1555|M.vanbaalenii_PYR-1 AIAPAGGWHRWSLPKYEIVFKFVMGFPVWLTAKALGPRAARLPGARAAAS
MSMEG_1655|M.smegmatis_MC2_155 GIAPAGGWTRFTPAKYEIVAKFVGGLPIVLATKLLGQRVHRLPFTRHISY
Mb3370|M.bovis_AF2122/97 GIAPAGGWTRWSPVKFEVIAKFIAGAPILAVAHILGQRALRLPFSRLLAT
Rv3338|M.tuberculosis_H37Rv GIAPAGGWTRWSPVKFEVIAKFIAGAPILAVAHILGQRALRLPFSRLLAT
MMAR_1187|M.marinum_M GIAPAGGWTRWSPAKFEVIIKFVMGIPLLALAWLLGPRVMRLPFSRRLAT
MUL_1440|M.ulcerans_Agy99 GIAPAGGWTRWSPAKFEVIIKFVMGIPLLALAWLFGPRVMRLPFSRRLAT
MAV_4312|M.avium_104 GIGAAGGWTRWSPVKFEVIGKFIGGMPLLLLARLLGQRILQLPLSRRLAT
MLBr_00685|M.leprae_Br4923 GIAPAGGWTRWSLAKFEVIAKFVAGMPLVAVARGLGSRALSLPFSHRLAS
Mflv_4874|M.gilvum_PYR-GCK AIAPAGGWPHHSLSKYETVLKFIMGGPALIAARLLGPRILNVPGARRLAT
MAB_3685|M.abscessus_ATCC_1997 AIAPAGGWTKYSPLKAEIVVKFAAMLPWLAFSRLLGRRAAGLPFAKFATA
.*..**** : : * * : ** * : :* * :* :: :
TH_1972|M.thermoresistible__bu VAVSAYPDGISDDDLVQIIDDVTHCAAYYQLLVKALLLPGLMELAETSVP
Mvan_1555|M.vanbaalenii_PYR-1 VPVSATPDGISDADMLDIIDDVTHCAAYYQLLVKALLLPGLMELADTRIP
MSMEG_1655|M.smegmatis_MC2_155 VAVAATADSLTDDDRRDLIDDLAHCPAYHKLLLKSLTTAGLMELPNSCTP
Mb3370|M.bovis_AF2122/97 LPISATPDGVSERELSGIIDDAAHCPAYFQLLVKALVLPGLQELEHTAVP
Rv3338|M.tuberculosis_H37Rv LPISATPDGVSERELSGIIDDAAHCPAYFQLLVKALVLPGLQELEHTAVP
MMAR_1187|M.marinum_M YAISASPDGVSDAELLGCIDDVAHCPAYLQLLVKALLLHGLRELAENAVP
MUL_1440|M.ulcerans_Agy99 YAISASPDGVSDAELLGCIDDVAHCPAYLQLLVKALLLRGLRELAENAVP
MAV_4312|M.avium_104 LPLSATPSGVSERQLSGVVEDAAHCPAYFQLLVKSLLQPGLLELAQTAVP
MLBr_00685|M.leprae_Br4923 LPLTGSPAGLSEHQLRNIVEDAGHCPAYFLLLIKTLLQPGLRELADITVP
Mflv_4874|M.gilvum_PYR-GCK LPVSGPADGPSQSDLAALVEDATHCSAYLQLLVKTLRMPGLLELASLGAP
MAB_3685|M.abscessus_ATCC_1997 RMLCGETEALSLPDFHSIFEDITHCPAYFASLTSILPAPGLENLDTISVP
: . . . : : .:* **.** * . * ** :* *
TH_1972|M.thermoresistible__bu THLVVCEKDRVLPNPRFVDHFHKNLREITRYTELDGVGHIPMFEAPQRVA
Mvan_1555|M.vanbaalenii_PYR-1 THMVICEKDRVLPHPRFVNHFRKHLPESTRFTALDGVGHIPMFEAPGRIA
MSMEG_1655|M.smegmatis_MC2_155 THLVICEKDRVLPHPRFTRHFTRHLPANAEVTHLDGVGHVPMFEAPGRVT
Mb3370|M.bovis_AF2122/97 SHVVLCEQDRVVPPSRFSRHFTDSLPAGHRLTVLDGVGHVPMFEAPGRIT
Rv3338|M.tuberculosis_H37Rv SHVVLCEQDRVVPPSRFSRHFTDSLPAGHRLTVLDGVGHVPMFEAPGRIT
MMAR_1187|M.marinum_M AHLVICGKDRIIPAPRYSRHFTTHLPDDHRVTVLDNVGHVPMFEAPARVT
MUL_1440|M.ulcerans_Agy99 AHLVICGKDRIIPAPRYSRHFTTHLPDDHRVTVLDNVGHVPMFEAPARVT
MAV_4312|M.avium_104 VQLVLCEKDRVVPPSRFSRHFTDHLPPETRITRLDGVGHVPMFEAPERVT
MLBr_00685|M.leprae_Br4923 VLLVLCEKDRVLPPPKYSRHFTDHLPERTQVTKFDGMGHVPMFEAPEVIT
Mflv_4874|M.gilvum_PYR-GCK TQLVLCEKDRVFPTPRGNRYFLTHLPDEARVVRLDGVGHIPMLEAPGAIT
MAB_3685|M.abscessus_ATCC_1997 AQLVLCEKDEIVPPGRFGKLIGEGLPGNARRITLAGMGHVPMLEAPGQVT
:*:* :*.:.* : : * . : .:**:**:*** ::
TH_1972|M.thermoresistible__bu DLIVDFLDDHTGKRRQASPGAVG
Mvan_1555|M.vanbaalenii_PYR-1 DLIRDFLDEQIDGQRTTG-----
MSMEG_1655|M.smegmatis_MC2_155 ELITEFVDRHTERGRAAG-----
Mb3370|M.bovis_AF2122/97 ELITSFIEECCPHVRAS------
Rv3338|M.tuberculosis_H37Rv ELITSFIEECCPHVRAS------
MMAR_1187|M.marinum_M ALISGFIEECCPHLRSAEPPAS-
MUL_1440|M.ulcerans_Agy99 ALISGFIEECCPHLRSAEPPAS-
MAV_4312|M.avium_104 TVLTDFIDDCVARSQRQQPPAS-
MLBr_00685|M.leprae_Br4923 KLITNFINQCSKPAEAASPAS--
Mflv_4874|M.gilvum_PYR-GCK ELIADFVDDHSTPSGKAIG----
MAB_3685|M.abscessus_ATCC_1997 EVITDLIDPLVSAKRSAASDAG-
:: :::