For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MNVEQVKSIDEAMGLAIEHSYQVKGTTYPKPPVGAVIVDPNGRIVGAGGTEPAGGDHAEVVALRRAGGLA AGAIVVVTMEPCNHYGKTPPCVNALIEARVGTVVYAVADPNGIAGGGAGRLSAAGLQVRSGVLAEQVAAG PLREWLHKQRTGLPHVTWKYATSIDGRSAAADGSSQWISSEAARLDLHRRRAIADAILVGTGTVLADDPA LTARLADGSLAPQQPLRVVVGKRDIPPEARVLNDEARTMMIRTHEPMEVLRALSDRTDVLLEGGPTLAGA FLRAGAINRILAYVAPILLGGPVTAVDDVGVSNITNALRWQFDSVEKVGPDLLLSLVAR
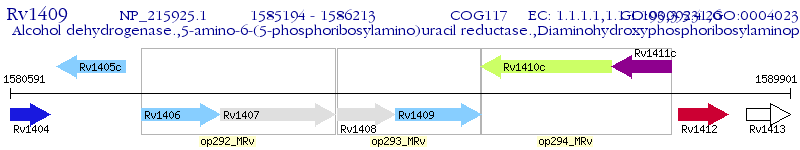
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. tuberculosis H37Rv | Rv1409 | ribG | - | 100% (339) | PROBABLE BIFUNCTIONAL riboflavin biosynthesis protein RIBG : Diaminohydroxyphosphoribosylaminopyrimidine deaminase (Riboflavin-specific deaminase) + 5-amino-6-(5-phosphoribosylamino) uracil reductase (HTP reductase) |
| M. tuberculosis H37Rv | Rv3752c | - | 2e-09 | 34.21% (152) | cytidine/deoxycytidylate deaminase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1444 | ribG | 0.0 | 99.41% (339) | bifunctional diaminohydroxyphosphoribosylaminopyrimidine |
| M. gilvum PYR-GCK | Mflv_3727 | - | 1e-138 | 72.12% (330) | riboflavin biosynthesis protein RibD |
| M. leprae Br4923 | MLBr_00555 | ribG | 1e-155 | 78.17% (339) | putative bifunctional riboflavin-specific |
| M. abscessus ATCC 19977 | MAB_2808c | - | 1e-128 | 65.56% (331) | bifunctional riboflavin biosynthesis protein RibG |
| M. marinum M | MMAR_2218 | ribG | 1e-172 | 88.20% (339) | bifunctional riboflavin biosynthesis protein RibG |
| M. avium 104 | MAV_3368 | ribD | 1e-151 | 77.95% (331) | riboflavin biosynthesis protein RibD |
| M. smegmatis MC2 155 | MSMEG_3067 | ribD | 1e-139 | 69.70% (330) | riboflavin biosynthesis protein RibD |
| M. thermoresistible (build 8) | TH_1007 | ribG | 2e-90 | 76.92% (208) | PROBABLE BIFUNCTIONAL riboflavin biosynthesis protein RIBG : |
| M. ulcerans Agy99 | MUL_1805 | ribG | 1e-172 | 87.91% (339) | bifunctional riboflavin biosynthesis protein RibG |
| M. vanbaalenii PYR-1 | Mvan_2690 | - | 1e-137 | 71.34% (328) | riboflavin biosynthesis protein RibD |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_3727|M.gilvum_PYR-GCK ------MNLDAAMAAAIVQSERVKGRTYPNPPVGAVILDSDGEIAGVGAT
Mvan_2690|M.vanbaalenii_PYR-1 ------MKLDAAMAAAIEQSERVKGRTYPNPPVGAVILDRDGEIAGVGAT
Rv1409|M.tuberculosis_H37Rv MNVEQVKSIDEAMGLAIEHSYQVKGTTYPKPPVGAVIVDPNGRIVGAGGT
Mb1444|M.bovis_AF2122/97 MNVEQVKSIDEAMGLAIEHSYQVKGTTYPNPPVGAVIVDPNGRIVGAGGT
MMAR_2218|M.marinum_M MTVEQVKSLEEAMRLAIEQSMLVKGTTYPNPPVGAVIVDQEGRVVGIGGT
MUL_1805|M.ulcerans_Agy99 MTVEQVKSLEEAMRLAIEQSMLVKGTTYPNPPVGAVIVDQEGWVVGIGGT
MLBr_00555|M.leprae_Br4923 MNVQTVDSLDAAMRLAVEQSYLVKGATYPNPPVGAVIVDPDGRVVGVGAT
MAV_3368|M.avium_104 MTASLGDRLDAAMRLAIEQSNQVKGNTYPNPPVGAVVLDGRGEVVGVGGT
TH_1007|M.thermoresistible__bu --------------------------------------------------
MSMEG_3067|M.smegmatis_MC2_155 ----MSISVEAAMRLAIDQAEQVKGATYPNPPVGAVILDRDGQVAGVGGT
MAB_2808c|M.abscessus_ATCC_199 --MTNPIGLDEAMDLAIRAAEEVKGSTYPNPPVGAVILDAAGQVAGVGGT
Mflv_3727|M.gilvum_PYR-GCK APTGGPHAEVVALRRAGERAAGGTAVVTLEPCNHFGRTPPCVDALIDAGV
Mvan_2690|M.vanbaalenii_PYR-1 APPGGPHAEVVALRRAGERAVGGTAVVTLEPCNHFGRTPPCVDALLDAGI
Rv1409|M.tuberculosis_H37Rv EPAGGDHAEVVALRRAGGLAAGAIVVVTMEPCNHYGKTPPCVNALIEARV
Mb1444|M.bovis_AF2122/97 EPAGGDHAEVVALRRAGGLATGAIVVVTMEPCNHYGKTPPCVNALIEARV
MMAR_2218|M.marinum_M QPSGGDHAEVVALRKAGGLAAGAIAVVTMEPCNHFGKTPPCVNALIEARV
MUL_1805|M.ulcerans_Agy99 QPSGGDHAEVVALRKAGGLAAGAIAVVTMEPCNHFGKTPPCVNALIEARV
MLBr_00555|M.leprae_Br4923 EPAGGDHAEVVALRSAGVTAAGAIAVVTLEPCNHYGKTPPCVNALLEAKV
MAV_3368|M.avium_104 EPAGGDHAEILALRRAGDLAAGGTVVVTLEPCNHHGKTPPCVDALLEAGV
TH_1007|M.thermoresistible__bu --------------------------------------------------
MSMEG_3067|M.smegmatis_MC2_155 QPTGGPHAEVMALRAAAERAEGGTAVVTLEPCNHHGRTPPCVDGLVAAGI
MAB_2808c|M.abscessus_ATCC_199 RPPGQEHAEVVALAGAGAAARGGTAVVTLEPCNHQGRTGPCVDALAAAGV
Mflv_3727|M.gilvum_PYR-GCK AAVAFAVADPNPAAAGGAARLAAAGVRVSAGVVADAVTGGPLREWLHKQR
Mvan_2690|M.vanbaalenii_PYR-1 AEVAFAVADPDPVAAGGAARLAESGVRVSAGVLAADVTGGPLREWLHKQR
Rv1409|M.tuberculosis_H37Rv GTVVYAVADPNGIAGGGAGRLSAAGLQVRSGVLAEQVAAGPLREWLHKQR
Mb1444|M.bovis_AF2122/97 GTVVYAVADPNGIAGGGAGRLSAAGLQVRSGVLAEQVAAGPLREWLHKQR
MMAR_2218|M.marinum_M GTVIYGVADPNGIAGGGAGRLSAAGLQVRSGVLADHVAAGPLREWLHKQR
MUL_1805|M.ulcerans_Agy99 GTVIYGVADPNGIAGGGAGRLSAAGLQVRSGVLADHVAAGPLREWLHKQR
MLBr_00555|M.leprae_Br4923 DKVAYAIADPNASAGGGAARLTAAGVHVQSGVLADLVTAGPLREWLHKQR
MAV_3368|M.avium_104 STVVYAVADPNPQAAGGAGRLQEAGVTVRSGLLADEVSGGPLREWLHKQR
TH_1007|M.thermoresistible__bu -------------------------------VLADEVAAGPLREWLHKTR
MSMEG_3067|M.smegmatis_MC2_155 SRVVYAVADPNPVAAGGSARMASSGIEVTSGVLSDEVAGGPLREWLHKQR
MAB_2808c|M.abscessus_ATCC_199 SRVVYAVADPNPQAAGGAARLRTLGIEVLDGLASDRVSAGPLREWLHKQR
: : *:.********* *
Mflv_3727|M.gilvum_PYR-GCK TGKPHVTWKFAASVDGRSAAADGTSQWITSEASRADVHRRRAAADAIVAG
Mvan_2690|M.vanbaalenii_PYR-1 TGMPHVTWKFATSVDGRSAAADGSSQWITSEAARADVHRRRATADAIIVG
Rv1409|M.tuberculosis_H37Rv TGLPHVTWKYATSIDGRSAAADGSSQWISSEAARLDLHRRRAIADAILVG
Mb1444|M.bovis_AF2122/97 TGLPHVTWKYATSIDGRSAAADGSSQWISSEAARLDLHRRRAIADAILVG
MMAR_2218|M.marinum_M TGLPHVTWKYATSIDGRSAAADGSSQWISSEAARADLHRRRAISDAIVVG
MUL_1805|M.ulcerans_Agy99 TGLPHVTWKYATSIDGRSAAADGSSQWISSEAARADLHRRRAISDAIVVG
MLBr_00555|M.leprae_Br4923 TGLPHVTWKYASSIDGRSAAADGSSRWLSSEASRADLHRRRATADAIVVG
MAV_3368|M.avium_104 TGLPHLTWKYASSVDGRSAAADGSSRWISSEASRLDLHRRRAAADAIVVG
TH_1007|M.thermoresistible__bu TGRPHVTWKFAASVDGRSAAADGTSQWITSEEARADVHRRRAAADAIVVG
MSMEG_3067|M.smegmatis_MC2_155 TGLPHVTWKFATSVDGRSAAADGSSQWITSAAARADVHRRRAAADAIVVG
MAB_2808c|M.abscessus_ATCC_199 TGRPHVTWKYGASVDGRSAANDGTSQWITSEQSRADVHRRRAFADAVLVG
** **:***:.:*:****** **:*:*::* :* *:***** :**::.*
Mflv_3727|M.gilvum_PYR-GCK TGTVFADDPALTARLPDGSLADHQPLRVVVGMREISPDATVLNDDSRTMV
Mvan_2690|M.vanbaalenii_PYR-1 TGTVFVDDPALTARLPDGSLADHQPLRVVVGKREISPDATVLNDDSRTMV
Rv1409|M.tuberculosis_H37Rv TGTVLADDPALTARLADGSLAPQQPLRVVVGKRDIPPEARVLNDEARTMM
Mb1444|M.bovis_AF2122/97 TGTVLADDPALTARLADGSLAPQQPLRVVVGKRDIPPEARVLNDEARTMM
MMAR_2218|M.marinum_M TGTVLVDDPTLTARLSDGSLAAEQPLRVVVGKRDIPPEAKVLNDDARTMM
MUL_1805|M.ulcerans_Agy99 TGTVLVDDPTLTARLSDGSLAAEQPLRVVVGKRDIPPEAKVLNDDARTMM
MLBr_00555|M.leprae_Br4923 TGTVLADNPTLTARLANGALADRQPLRVVVGMREIPLEANVLNDDSPTMV
MAV_3368|M.avium_104 TGTVLADDPALTARLPDGTLADRQPLRVVVGMREIPSEAKVLNDDSRTMV
TH_1007|M.thermoresistible__bu TGTVLADDPTLTARRPDGSLAERQPLRVVVGTRDISADANVLNDDSRTMV
MSMEG_3067|M.smegmatis_MC2_155 TGTVFVDDPSLTARLPDGTLADRQPLRVVVGRREVSSDANVLNDDSRTMV
MAB_2808c|M.abscessus_ATCC_199 TGTVFADDPQLTARTPDNELVPHQPLRVVVGMREVSSDAKVLNDDSHTMV
****:.*:* **** .:. *. .******** *::. :* ****:: **:
Mflv_3727|M.gilvum_PYR-GCK IRTRDPHEVIQALTDRTDVLVEGGPTLAGAFLRAGAIDRILAYVAPILLG
Mvan_2690|M.vanbaalenii_PYR-1 IRTHNPHEVIKALADRTDVFVEGGPTLAGAFLRAGVIDRILAYVAPILLG
Rv1409|M.tuberculosis_H37Rv IRTHEPMEVLRALSDRTDVLLEGGPTLAGAFLRAGAINRILAYVAPILLG
Mb1444|M.bovis_AF2122/97 IRTHEPMEVLRALSDRTDVLLEGGPTLAGAFLRAGAINRILAYVAPILLG
MMAR_2218|M.marinum_M IKTHEPVEVLRALSDRTDVLLEGGPTLAGAFLRAGAINRILAYVAPILLG
MUL_1805|M.ulcerans_Agy99 IKTHEPVEVLRALSDRTDVLLEGGPTLAGAFLRAGAINRILAYVAPILLG
MLBr_00555|M.leprae_Br4923 IRTHDPMEVLKALADRTDVLLEGGPTLAGAFLRAGTINRILAYVAPILLG
MAV_3368|M.avium_104 IRTHDPAEVLKAVSDRTDVLLEGGPTLAGAFVRAGLVNRILVYLAPTLLG
TH_1007|M.thermoresistible__bu IRTHDPHEVVRALADRTDVLLEGGPTLAGAFVRAGLVDRILAYVAPVLLG
MSMEG_3067|M.smegmatis_MC2_155 IRTHDPHEVLRALSDRTDIMLEGGPTLAGAFLRAGVVNRILAYVAPILLG
MAB_2808c|M.abscessus_ATCC_199 IRTHDPAEVLAALGDRTDIILEGGPTLAGAFLRAGLVDRILAYLAPMLLG
*:*::* **: *: ****:::**********:*** ::***.*:** ***
Mflv_3727|M.gilvum_PYR-GCK GPVTAVDDVGVSSITQAHRWRFDGMQAIGPDVLLSLVPN---
Mvan_2690|M.vanbaalenii_PYR-1 GPITAVDDVGVLSIAQAQRWRFDGMTAVGPDVLLSLTPV---
Rv1409|M.tuberculosis_H37Rv GPVTAVDDVGVSNITNALRWQFDSVEKVGPDLLLSLVAR---
Mb1444|M.bovis_AF2122/97 GPVTAVDDVGVSNITNALRWQFDSVEKVGPDLLLSLVAR---
MMAR_2218|M.marinum_M GPVTAVDDVGVSSIAHALRWQFDGVERCGPDLLLSLVSR---
MUL_1805|M.ulcerans_Agy99 GPVTAVDDVGVSSIAHALRWQFDGVERCGPDLLLSLVSR---
MLBr_00555|M.leprae_Br4923 GPVTAVDDVGVPNIARALRWQFDGIDRAGSDLVISLVTR---
MAV_3368|M.avium_104 GPVTAVDDVGVPTIAKALRWQFDGVDRVGPDLLLSLVPRSG-
TH_1007|M.thermoresistible__bu GPVTVVDDVGVTTIGNAHRWRFDGMTSVGPDVLLSLVPR---
MSMEG_3067|M.smegmatis_MC2_155 GPITAVDDVGVPSIAHAQRWRFDCVQTIGPDVLMSLVPN---
MAB_2808c|M.abscessus_ATCC_199 GNVPAVEDVGVTTLSRALRWEYDQIERMGPDVLLSLVPRLPV
* :..*:**** .: .* **.:* : *.*:::**..