For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VSELRLMAVHAHPDDESSKGAATLARYADEGHRVLVVTLTGGERGEILNPAMDLPDVHGRIAEIRRDEMT KAAEILGVEHTWLGFVDSGLPKGDLPPPLPDDCFARVPLEVSTEALVRVVREFRPHVMTTYDENGGYPHP DHIRCHQVSVAAYEAAGDFCRFPDAGEPWTVSKLYYVHGFLRERMQMLQDEFARHGQRGPFEQWLAYWDP DHDFLTSRVTTRVECSKYFSQRDDALRAHATQIDPNAEFFAAPLAWQERLWPTEEFELARSRIPARPPET ELFAGIEP
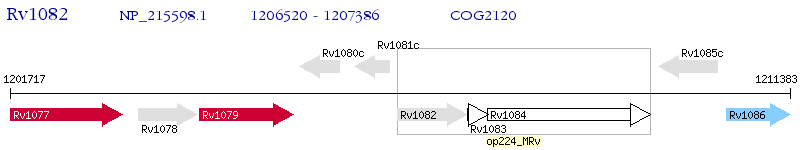
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. tuberculosis H37Rv | Rv1082 | mca | - | 100% (288) | Mycothiol conjugate amidase Mca (Mycothiol S-conjugate amidase) |
| M. tuberculosis H37Rv | Rv1170 | mshB | 4e-28 | 42.22% (180) | N-Acetyl-1-D-myo-Inosityl-2-amino-2-deoxy-alpha- |
| M. tuberculosis H37Rv | Rv0323c | - | 5e-13 | 31.29% (147) | hypothetical protein Rv0323c |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1111 | mca | 1e-174 | 100.00% (288) | Mycothiol conjugate amidase Mca (Mycothiol S-conjugate |
| M. gilvum PYR-GCK | Mflv_2050 | - | 1e-137 | 78.40% (287) | LmbE family protein |
| M. leprae Br4923 | MLBr_02391 | - | 1e-150 | 86.76% (287) | hypothetical protein MLBr_02391 |
| M. abscessus ATCC 19977 | MAB_1203 | - | 1e-116 | 67.25% (287) | LmbE-like protein |
| M. marinum M | MMAR_4385 | mca | 1e-149 | 86.41% (287) | mycothiol conjugate amidase Mca |
| M. avium 104 | MAV_1206 | - | 1e-147 | 84.38% (288) | mycothiol conjugate amidase Mca |
| M. smegmatis MC2 155 | MSMEG_5261 | - | 1e-138 | 77.78% (288) | mycothiol conjugate amidase Mca |
| M. thermoresistible (build 8) | TH_2167 | - | 1e-140 | 79.09% (287) | mycothiol conjugate amidase Mca |
| M. ulcerans Agy99 | MUL_0198 | mca | 1e-149 | 86.06% (287) | mycothiol conjugate amidase Mca |
| M. vanbaalenii PYR-1 | Mvan_4667 | - | 1e-138 | 78.05% (287) | LmbE family protein |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_2050|M.gilvum_PYR-GCK MSELRLMAVHAHPDDESSKGAAAMARYVDEGVRVMVVTLTGGERGDILNP
Mvan_4667|M.vanbaalenii_PYR-1 MSELRLMAVHAHPDDESSKGAATMARYVDEGVRVMVVTLTGGERGDILNP
MSMEG_5261|M.smegmatis_MC2_155 MSELRLMAVHAHPDDESSKGAATTARYAAEGARVMVVTLTGGERGDILNP
TH_2167|M.thermoresistible__bu MTELRLMAVHAHPDDESSKGAATTAKYADEGARVMVVTLTGGERGDILNP
Rv1082|M.tuberculosis_H37Rv MSELRLMAVHAHPDDESSKGAATLARYADEGHRVLVVTLTGGERGEILNP
Mb1111|M.bovis_AF2122/97 MSELRLMAVHAHPDDESSKGAATLARYADEGHRVLVVTLTGGERGEILNP
MMAR_4385|M.marinum_M MSKLRLMAVHAHPDDESSKGAATLARYADEGHRVLVVTLTGGERGEILNP
MUL_0198|M.ulcerans_Agy99 MSKLRLMAVHAHPDDESSKGAATLARYADEGHRVLVVTLTGGERGEILNP
MLBr_02391|M.leprae_Br4923 MSELRLMAVHAHPDDESSKGAATLARYADEGHRVLVVTLTGGERGEILNP
MAV_1206|M.avium_104 MSELRLMAVHAHPDDESSKGAATLARYADEGHRVLVVTLTGGERGDILNP
MAB_1203|M.abscessus_ATCC_1997 MTGLRLMAVHAHPDDEASKGAATTARYAAEGNEVMVVTLTGGERGDILNP
*: *************:*****: *:*. ** .*:**********:****
Mflv_2050|M.gilvum_PYR-GCK AMDLPEVHGRIHEIRIDEMARAAEILGVEHHWLGFVDSGLPEGDPLPPLP
Mvan_4667|M.vanbaalenii_PYR-1 AMDLPEVHGRMAEIRRDEMARAAEILGVEHHWLGFVDSGLPEGDPPPPLP
MSMEG_5261|M.smegmatis_MC2_155 AMDLPEVHGRIAEVRRDEMAKAAEILGVEHHWLGFVDSGLPEGDPLPPLP
TH_2167|M.thermoresistible__bu AMDLPEVRGRIAEVRRDEMAKAAEILGVEHRWLGFVDSGLPEGDPPPPLP
Rv1082|M.tuberculosis_H37Rv AMDLPDVHGRIAEIRRDEMTKAAEILGVEHTWLGFVDSGLPKGDLPPPLP
Mb1111|M.bovis_AF2122/97 AMDLPDVHGRIAEIRRDEMTKAAEILGVEHTWLGFVDSGLPKGDLPPPLP
MMAR_4385|M.marinum_M AMDLPDVHGHISEIRRDEMAKAAEILGVEHTWLGFVDSGLPKGDPPPPLP
MUL_0198|M.ulcerans_Agy99 AMDLPDVHGHISEIRRDEMAKAAEILGVEHTWLGFVDSGLPKGDPPPPLP
MLBr_02391|M.leprae_Br4923 AMDLPDVHGHIAEIRRDEMAKAAEILGVEHTWLGFIDSGLPKGDPPPPLP
MAV_1206|M.avium_104 AMDLPEVHGHIAEIRRDEMAKAAEILGVEHTWLGFVDSGLPKGDPPPPLP
MAB_1203|M.abscessus_ATCC_1997 AMDRPEVAANIGAIRREEMAKAAAELGVRHRWLGYVDSGLPQGDPLPPLP
*** *:* ..: :* :**::** ***.* ***::*****:** ****
Mflv_2050|M.gilvum_PYR-GCK EGCFALEPLEVPTEALVKVIRDFRPHVMTTYDENGGYPHPDHIKCHQVSV
Mvan_4667|M.vanbaalenii_PYR-1 EGCFALEPLEVPTHALVRVIRDFKPHVITTYDENGGYPHPDHIKCHQVSV
MSMEG_5261|M.smegmatis_MC2_155 DGCFALVPLEEPVKRLVRVIREFRPHVMTTYDENGGYPHPDHIRCHQVSV
TH_2167|M.thermoresistible__bu EGCFATVPLEQATERLVRLIREFRPHVMTTYDENGGYPHPDHIRCHQVSV
Rv1082|M.tuberculosis_H37Rv DDCFARVPLEVSTEALVRVVREFRPHVMTTYDENGGYPHPDHIRCHQVSV
Mb1111|M.bovis_AF2122/97 DDCFARVPLEVSTEALVRVVREFRPHVMTTYDENGGYPHPDHIRCHQVSV
MMAR_4385|M.marinum_M EGCFALVPLEDSIEALVRVVREFRPHVMTTYDENGGYPHPDHIRCHQVSI
MUL_0198|M.ulcerans_Agy99 EGCFALVPLEDSIEALVRVVREFRPHVMTTYDENGGYPHPDHIRCHQVSI
MLBr_02391|M.leprae_Br4923 DDCFALVPLEVCTEALVRVVRKFRPHVLTTYDENGGYPHPDHIRCHQVSV
MAV_1206|M.avium_104 EGCFALVPLEEAIEALVRVVREFRPHVMTTYDETGGYPHPDHIRCHQVSV
MAB_1203|M.abscessus_ATCC_1997 SDSFALVPIEEPIAKLVAVVREFRPHVMTTYDENGGYPHPDHIRCHEVSM
...** *:* ** ::*.*:***:*****.*********:**:**:
Mflv_2050|M.gilvum_PYR-GCK AAYEAAADHLLYPDAGEPWSVSKLYYNHGFLRQRMQVLQDEFAKHGEEGP
Mvan_4667|M.vanbaalenii_PYR-1 AAYEAAADHRLFPEAGDPWTPLKLYYNHGFLRQRMQVLQDEFAKHGKEGP
MSMEG_5261|M.smegmatis_MC2_155 AAYEAAADHLLYPDAGEPWAVQKLYYNHGFLRQRMQLLQEEFAKNGQEGP
TH_2167|M.thermoresistible__bu AAYEAAADYRLFPDAGPPWAVLKLYYNHGFIRQRMKLLQDEFEKNGMEGP
Rv1082|M.tuberculosis_H37Rv AAYEAAGDFCRFPDAGEPWTVSKLYYVHGFLRERMQMLQDEFARHGQRGP
Mb1111|M.bovis_AF2122/97 AAYEAAGDFCRFPDAGEPWTVSKLYYVHGFLRERMQMLQDEFARHGQRGP
MMAR_4385|M.marinum_M GAYEAAADYLRFPDAGEPWTVSKLYYIHGFLRERMRILQDEFIKRGQEGP
MUL_0198|M.ulcerans_Agy99 GAYEAAADYLRFPDAGEPWTVSKLYYIHGFLRERMRILQDEFIKRGQEDP
MLBr_02391|M.leprae_Br4923 DAYEAACDYRRFPDAGKPWTVSKLYYNHGFLRARMQLLHDEFAKHGQAGP
MAV_1206|M.avium_104 GAYEAAGDYRRFPDAGEPWTVSKLYYNHGFLRARMQLLHDEAVKHGHEPP
MAB_1203|M.abscessus_ATCC_1997 AAFEAAADPEQFPEAGEPWTVSKIYYNHGFMRARMQLLNDECKKHGYKGP
*:*** * :*:** **: *:** ***:* **::*::* :.* *
Mflv_2050|M.gilvum_PYR-GCK FAKWLEKWDPDDDVLDKRVTTRIECAKYFGKRDDALLAHATQIDPNSFFF
Mvan_4667|M.vanbaalenii_PYR-1 FATWLEKWDPDEDLLAKRVTTRIECAKYFAQRDEALLAHATQIDPNSFFF
MSMEG_5261|M.smegmatis_MC2_155 FAKWLEHWDPDNDVFANRVTTRVHCAEYFHQRDDALRAHATQIDPKGDFF
TH_2167|M.thermoresistible__bu FAKWLERWDPENDVFEKRVTTRVECAKYFGRRDDALRAHATQIDPNSFFF
Rv1082|M.tuberculosis_H37Rv FEQWLAYWDPDHDFLTSRVTTRVECSKYFSQRDDALRAHATQIDPNAEFF
Mb1111|M.bovis_AF2122/97 FEQWLAYWDPDHDFLTSRVTTRVECSKYFSQRDDALRAHATQIDPNAEFF
MMAR_4385|M.marinum_M FGKWLEHWHPDHDPFANRVTTRVECSGYFAQRDDALRAHATQIDPNAEFF
MUL_0198|M.ulcerans_Agy99 FGKWLEHWHPDHDPFANRVTTRVECSGYFGQRDDALRAHATQIDPNAEFF
MLBr_02391|M.leprae_Br4923 FDKWLAQSNPAHDPFESRVTTRVECSAYFSQRDDALRAHATQIDPKAEFF
MAV_1206|M.avium_104 FKKWLEHWDPAHDPFESRVTTRVECSAYFSQRDDALRAHTTQIDPDHDFF
MAB_1203|M.abscessus_ATCC_1997 FDEWLERWPADDDTFDRRVTTRVPCADFFDARDAALRAHASQIPPDSSFF
* ** . .* : *****: *: :* ** ** **::** *. **
Mflv_2050|M.gilvum_PYR-GCK TTPMEWQQRLWPTEEFELARSRVPVVLPETDLFAGIEDLGD---
Mvan_4667|M.vanbaalenii_PYR-1 STPMEWQQRLWPTEEFELARSRVPVTLPETHLFAGIEGYE----
MSMEG_5261|M.smegmatis_MC2_155 HAPIEWQQRLWPTEEFELARARVPVTLPEDDLFKGVEP------
TH_2167|M.thermoresistible__bu AAPIEWQERLWPTEEFELARSRVPVRLPETDLFAGIEDHVDKLD
Rv1082|M.tuberculosis_H37Rv AAPLAWQERLWPTEEFELARSRIPARPPETELFAGIEP------
Mb1111|M.bovis_AF2122/97 AAPLAWQERLWPTEEFELARSRIPARPPETELFAGIEP------
MMAR_4385|M.marinum_M AAPLSWQRELWPTEEFELARSRIPIGLPETELFAGIESSQ----
MUL_0198|M.ulcerans_Agy99 AAPLSWQRELWPTEEFELARSRIPIGLPETELFAGIESSQ----
MLBr_02391|M.leprae_Br4923 AAPISWQQRLWPTEEFELARSRVPTRLPEHDLFAGIEAAG----
MAV_1206|M.avium_104 AAPIAWQQRLWPTEEFELARSRVPVRLPEDDLFAGIEPDA----
MAB_1203|M.abscessus_ATCC_1997 AVPREFERKLWPTEDFELAKSRVPTSTPEDDLFAGVETK-----
.* ::..*****:****::*:* ** .** *:*