For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MGKHHTPEASTFRDRSAVRTTVLALAALTVFVIAMVASYSGAFAKPTLHHMTVAVAAPPEIIDGLRDQHT LSVDEVGDAAAARAQVYQRSADAALAVTPDQRMTIYVAGGGGRSVAAAAELVGRETAERADLSPVVEDVA PTSPGDPSGTVEFYAVIFLSIGASLGAALLGRLMGSVRTPATLALRTVSLAAFSAVLAGAVTAYVGPVLD ALTGHIWQVFGALWLYSMAVAGAVTGVAAAFGTAASVALGLFLVVVGNAAAAGPVGRPLLSGFYSTFTTI VPQGSGVSLLRSISYFGGHGATTPLVTLGIWAAAGCFLAVGATAARVNYRALYERRVGQYRAVLRSRLVP SAD
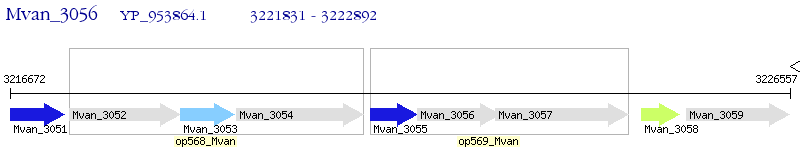
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_3056 | - | - | 100% (353) | integral membrane protein |
| M. vanbaalenii PYR-1 | Mvan_0640 | - | 8e-20 | 27.49% (331) | hypothetical protein Mvan_0640 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | - | - | - | - | - |
| M. gilvum PYR-GCK | Mflv_3330 | - | 1e-154 | 78.93% (356) | integral membrane protein |
| M. tuberculosis H37Rv | - | - | - | - | - |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | - | - | - | - | - |
| M. marinum M | MMAR_0333 | - | 3e-05 | 27.24% (279) | hypothetical protein MMAR_0333 |
| M. avium 104 | - | - | - | - | - |
| M. smegmatis MC2 155 | MSMEG_3573 | - | 1e-122 | 68.28% (331) | integral membrane protein, putative |
| M. thermoresistible (build 8) | TH_2357 | - | 2e-21 | 27.27% (319) | integral membrane protein |
| M. ulcerans Agy99 | - | - | - | - | - |
CLUSTAL 2.0.9 multiple sequence alignment
Mvan_3056|M.vanbaalenii_PYR-1 MGKHHTPE---ASTFR--DRSAVR----------------------TTVL
Mflv_3330|M.gilvum_PYR-GCK MGRHHTPD---IPPIR--DRTAVR----------------------TTLL
MSMEG_3573|M.smegmatis_MC2_155 MGRHHAPEDRRAATTRPVTRTQIR----------------------HIAI
TH_2357|M.thermoresistible__bu MTTQSAPPND-RSTIRRHEQPAAV----------------------RATG
MMAR_0333|M.marinum_M MALAATRKLVRDPQLRRLMAVCATFLTSNSAHGVLLTVFAFSVGGAATVG
* : . *
Mvan_3056|M.vanbaalenii_PYR-1 ALAALTVFVIAMVASYSGAFAKPTLHHMTVAVAAPPEIIDGLRD------
Mflv_3330|M.gilvum_PYR-GCK ALAGLTVFVIAMVASYSGAFANPTLHHMTVAVSAPQQVVDGLRA------
MSMEG_3573|M.smegmatis_MC2_155 VMTALAMGVLAMIASYSGAFAKPTLHHMTVAVAAPTQLVDQLRE------
TH_2357|M.thermoresistible__bu IVVVLTVAIAIIAIAFAWPASRSAPGDLPIGVAAPPPVTAQITARLAEQA
MMAR_0333|M.marinum_M VVTLVRMLPSALLAPLAATLATSPRPQVHLAIGVGARGMTTIATVIVVVS
:. : : : . : . : .. .: :.:.. :
Mvan_3056|M.vanbaalenii_PYR-1 QHTLSVDEVGDAAAARAQVYQRSADAALAVTPDQRMTIYVAGGGGRSVAA
Mflv_3330|M.gilvum_PYR-GCK EHALTVHEFGDAASARAQVLERTADTAFAVNPDGHLTVYVAGGGGHSVAA
MSMEG_3573|M.smegmatis_MC2_155 QDSLSVTDVPDEAAAHAQVLQRDADAALVVAAPDRLDIYVAGGGGRSVAT
TH_2357|M.thermoresistible__bu PDAFDVRAFDDDAALRAAIRDREVYGGLSIG-AGPPTLLIATGGSPAVAQ
MMAR_0333|M.marinum_M GAPLGVVLVVLGVDSLMSAAVRSLHAALVVRLATSVAEAATANAATGVLL
.: * . . * .: : : ... .*
Mvan_3056|M.vanbaalenii_PYR-1 AAELVGRETAERADLSPV---------------------------VEDVA
Mflv_3330|M.gilvum_PYR-GCK AAELVGREAAERAGLTPV---------------------------VEDVA
MSMEG_3573|M.smegmatis_MC2_155 AAESVGRAAAEQAGLVPA---------------------------VTDIA
TH_2357|M.thermoresistible__bu LLTQMAGGIAEQSGMPLR---------------------------TEDLA
MMAR_0333|M.marinum_M YASALAGPALAGVMLAFVGVGWALTLPAALFVVGALTALRITVAPIDDMP
:. : *:.
Mvan_3056|M.vanbaalenii_PYR-1 PTSPGDPSG-----------------------------------------
Mflv_3330|M.gilvum_PYR-GCK PTSAGDPSG-----------------------------------------
MSMEG_3573|M.smegmatis_MC2_155 PTSPGNPSG-----------------------------------------
TH_2357|M.thermoresistible__bu APPPDDPRG-----------------------------------------
MMAR_0333|M.marinum_M DTGGARPSGGVARVQLRAVGAGFRAIIESRPAAAAATLFLLTTTLLDGFW
. * *
Mvan_3056|M.vanbaalenii_PYR-1 ----------------TVEFYAVIFLSIGASLGAALLGRLMGSVRTPATL
Mflv_3330|M.gilvum_PYR-GCK ----------------TVEFYAVIFLSIGASLGAALLGRLMGPVRTPATL
MSMEG_3573|M.smegmatis_MC2_155 ----------------TVEFYAIIFISLGASVGAAAFGYLAGPVRRPMTL
TH_2357|M.thermoresistible__bu ----------------VGLAASALPITLAGLLPAIALVLALR-----REV
MMAR_0333|M.marinum_M YVASAPLAADQFGLGPSGVTTIMTIYGVGGLLGAVAMLSIAGRGGLARMF
:.. : * : .
Mvan_3056|M.vanbaalenii_PYR-1 ALRTVSLAAFSAVLAGAVTAYVGPVLDALTG----HIWQVFGALWLYSMA
Mflv_3330|M.gilvum_PYR-GCK ALRTASLSVFSAVLAGVVTLYTGPVLDALTG----HSWQVFGALWLYSMA
MSMEG_3573|M.smegmatis_MC2_155 ALRTVTLAAYSALLAGLVTVYVDPVLDALVG----HPWQVFGVLWLYCLA
TH_2357|M.thermoresistible__bu WTRFAATVVFAALAAWTIALLLRFVFGSITD----NFWGVTAGLTLGILA
MMAR_0333|M.marinum_M TLARLSEALLLAAIGAASLPALGLVLAAGVGGGSAASWAIAPTLVQRSVN
: * . *: : .. * : * :
Mvan_3056|M.vanbaalenii_PYR-1 VAGAVTGVAAAFGTAASVALG---LFLVVVG-------------------
Mflv_3330|M.gilvum_PYR-GCK VGGAVTGVAAAFGSAASVVLG---LFLVVVG-------------------
MSMEG_3573|M.smegmatis_MC2_155 VGGAVTGVSAAFGPVASLMLT---AFLVIVG-------------------
TH_2357|M.thermoresistible__bu AGLTVLGLGSVFGRAGLAIGA---LLAILLG-------------------
MMAR_0333|M.marinum_M RASMVTAAASLQSVSQLAIAAGAGIAAVLVGSLGLTGALAIVSGAAALIT
. * . .: . :::*
Mvan_3056|M.vanbaalenii_PYR-1 -----------------NAAAAGPVGRPLLSGFYSTFTTIVPQGSGVSLL
Mflv_3330|M.gilvum_PYR-GCK -----------------NAAAAGPVGRPLLSGFYSTFTAIVPQGSGVSLL
MSMEG_3573|M.smegmatis_MC2_155 -----------------NAAAAGPVGRPLLSSFYSTFNLIVPQGSGVSLL
TH_2357|M.thermoresistible__bu -----------------NPLSGLNAAPELLPRGWGQLGQLLPQGANATLL
MMAR_0333|M.marinum_M VLAWPQVRRADKLSDDDSAKLAVIRATPVLAPLPALALEQLARTATRMAV
.. . . :*. . :.: : :
Mvan_3056|M.vanbaalenii_PYR-1 RSISYFGGHGATTPLVTLGIWAAAGCFLAVGATAAR---VNYRAL-----
Mflv_3330|M.gilvum_PYR-GCK RSISYFGGHGAAMSWVTLVVWAVTGAVLATLATAVR---VNYRAL-----
MSMEG_3573|M.smegmatis_MC2_155 RSVEYFDGNGAAQPLLTLAVWAAAGCVLAVLGTAVRSARVRNSALRVARR
TH_2357|M.thermoresistible__bu RSTAYFDGAGATTAIWVLVCWVLVGTALIAVAALRQ--------------
MMAR_0333|M.marinum_M PAGSDVIRQGDRGDRFYMIASGLADATVNGRRVATLGPGGSFGEIALLHD
: . * : .. : .
Mvan_3056|M.vanbaalenii_PYR-1 --------YER---RVGQYRAVLRSRLVPSAD------------------
Mflv_3330|M.gilvum_PYR-GCK --------YERRVERAGGSRTLLRPGLLSSTD------------------
MSMEG_3573|M.smegmatis_MC2_155 CAPGRGPIYRGRHDRNRSERALLPAGVLSPAD------------------
TH_2357|M.thermoresistible__bu ----------------RRTAEPHPAEMH----------------------
MMAR_0333|M.marinum_M VARSATVTARKDLDLVAVDRAEFLAALSSNTAAGLRLGGLARTRLGTVPV
. :
Mvan_3056|M.vanbaalenii_PYR-1 --------------------------------------------------
Mflv_3330|M.gilvum_PYR-GCK --------------------------------------------------
MSMEG_3573|M.smegmatis_MC2_155 --------------------------------------------------
TH_2357|M.thermoresistible__bu --------------------------------------------------
MMAR_0333|M.marinum_M EERLIELGPDPALNSQSLSELLAAQPALAGIGDGALRELADTAKVLAAPD
Mvan_3056|M.vanbaalenii_PYR-1 --------------------------------------------------
Mflv_3330|M.gilvum_PYR-GCK --------------------------------------------------
MSMEG_3573|M.smegmatis_MC2_155 --------------------------------------------------
TH_2357|M.thermoresistible__bu --------------------------------------------------
MMAR_0333|M.marinum_M GALITCEGDHGDTYYVILEGAAQVLDDGTAVHHLGPGEGFGEQAILRDVP
Mvan_3056|M.vanbaalenii_PYR-1 --------------------------
Mflv_3330|M.gilvum_PYR-GCK --------------------------
MSMEG_3573|M.smegmatis_MC2_155 --------------------------
TH_2357|M.thermoresistible__bu --------------------------
MMAR_0333|M.marinum_M RTATVRAVGDTTLVAVDREAFQRARR