For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VRLVFAGTPEPALPSLQRLIDSDRHEVIAVMTRPDAVAGRRGRPSPSPVAQLAAAHGIPVLKPARPNSGE FVAELSALSPDCCAVVAYGALLGDALLAVPAHGWVNLHFSVLPAWRGAAPVQAALAAGDEVTGATTFQIE RSLDSGPVYGVVTETIRPTDTAGDLLERLSVSGAGLLEATMDGIEDGTLTAVPQPPEGVSVAPKVTVDDA RIRWELPAHVVDRRVRSVTPNPGAWTMIGDQRIKVGPVTVAGDGPKGLEPGEIRVDKKHIHVGTGSDAVL LGTVQPPGKKPMNAADWARGARLDENGWAR
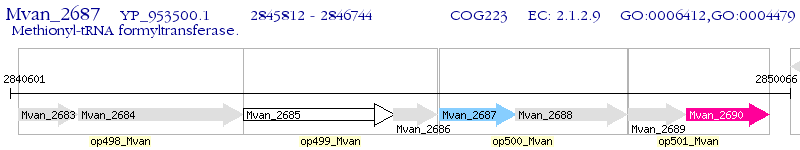
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_2687 | fmt | - | 100% (310) | methionyl-tRNA formyltransferase |
| M. vanbaalenii PYR-1 | Mvan_4857 | purN | 1e-07 | 32.35% (102) | phosphoribosylglycinamide formyltransferase |
| M. vanbaalenii PYR-1 | Mvan_1265 | purU | 6e-05 | 26.37% (91) | formyltetrahydrofolate deformylase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1441 | fmt | 1e-128 | 72.88% (306) | methionyl-tRNA formyltransferase |
| M. gilvum PYR-GCK | Mflv_3730 | fmt | 1e-156 | 87.74% (310) | methionyl-tRNA formyltransferase |
| M. tuberculosis H37Rv | Rv1406 | fmt | 1e-128 | 72.88% (306) | methionyl-tRNA formyltransferase |
| M. leprae Br4923 | MLBr_00552 | fmt | 1e-121 | 70.00% (310) | methionyl-tRNA formyltransferase |
| M. abscessus ATCC 19977 | MAB_2811c | - | 1e-117 | 69.74% (304) | methionyl-tRNA formyltransferase |
| M. marinum M | MMAR_2215 | fmt | 1e-131 | 74.18% (306) | methionyl-tRNA formyltransferase Fmt |
| M. avium 104 | MAV_3372 | fmt | 1e-130 | 72.82% (309) | methionyl-tRNA formyltransferase |
| M. smegmatis MC2 155 | MSMEG_3064 | fmt | 1e-139 | 77.35% (309) | methionyl-tRNA formyltransferase |
| M. thermoresistible (build 8) | TH_1529 | fmt | 1e-133 | 73.33% (315) | PROBABLE METHIONYL-TRNA FORMYLTRANSFERASE FMT |
| M. ulcerans Agy99 | MUL_1802 | fmt | 1e-132 | 74.18% (306) | methionyl-tRNA formyltransferase |
CLUSTAL 2.0.9 multiple sequence alignment
Mvan_2687|M.vanbaalenii_PYR-1 --MRLVFAGTPEPALPSLQRLIDSDRHEVIAVMTRPDAVAGRRGRPSPSP
Mflv_3730|M.gilvum_PYR-GCK --MRLVFAGTPEPALPSLQRLIDSARHDVIAVLTRPDAAAGRRGRPSPSP
Mb1441|M.bovis_AF2122/97 --MRLVFAGTPEPALASLRRLIESPSHDVIAVLTRPDAASGRRGKPQPSP
Rv1406|M.tuberculosis_H37Rv --MRLVFAGTPEPALASLRRLIESPSHDVIAVLTRPDAASGRRGKPQPSP
MLBr_00552|M.leprae_Br4923 MLVRLVFAGTPESALPALCRLIDSPRHDVIAVLTRPDAASGRRGKPEPSP
MAV_3372|M.avium_104 MPVRLVFAGTPEPALPALRRLLDSPRHEVIAVLTRPDAASGRRGKPEPSP
MMAR_2215|M.marinum_M --MRLVFAGTPETALPALHQLIDSPRHDVIAVLTRPDAASGRRGKPEPSP
MUL_1802|M.ulcerans_Agy99 --MRLVFAGTPETALPALHQLIDSPRHDVIAVLTRPDAASGRRGKPEPSP
MSMEG_3064|M.smegmatis_MC2_155 --MRLVFAGTPEPALPSLRRLIESPRHDVVAVLTRPDAAAGRRGKPRPSP
TH_1529|M.thermoresistible__bu --MRLVFAGTPEAALPSLRRLIDXXRHEVVAVLTRPDAAAGRRARPAPSP
MAB_2811c|M.abscessus_ATCC_199 --MRIVFAGTPAPALPSLERLLAS-RHEVVAVLTRPDARAGRGRASASSP
:*:****** .**.:* :*: *:*:**:***** :** . .**
Mvan_2687|M.vanbaalenii_PYR-1 VAQLAAAHGIPVLKPARPNSGEFVAELSALSPDCCAVVAYGALLGDALLA
Mflv_3730|M.gilvum_PYR-GCK VAELAAAHGIPVLKPPRPNSEEFVAELAALAPDCCAVVAYGALLREELLA
Mb1441|M.bovis_AF2122/97 VAREAAERGIPVLRPSRPNSAEFVAELSDLAPECCAVVAYGALLGGPLLA
Rv1406|M.tuberculosis_H37Rv VAREAAERGIPVLRPSRPNSAEFVAELSDLAPECCAVVAYGALLGGPLLA
MLBr_00552|M.leprae_Br4923 VAREALDRGIPLLRPARPNSPVFVSELSEWAPECCVVVAYGALLGSPLLA
MAV_3372|M.avium_104 VAREALDRGIPVLRPARPNSPEFVAELAQLAPDCCAVVAYGALLRDELLA
MMAR_2215|M.marinum_M VARAALERDIPVLRPSRPNSAEFVAELSELAPQCCAVVAYGALLGDALLG
MUL_1802|M.ulcerans_Agy99 VARAALERGIPVLRPSRPNSAEFVAELSELAPQCCAVVAYGALLGDALLG
MSMEG_3064|M.smegmatis_MC2_155 VAQLALEHGIPLLRPDRPNSDEFVAELTELAPDCCAVVAYGALLSQRLLA
TH_1529|M.thermoresistible__bu VAKLASEHGIPVLRPPSPNTDEFIAELTELAPDCCPVVAYGALLSERLLA
MAB_2811c|M.abscessus_ATCC_199 VAALAREHGVAVLTPARPNEPEFVRELAQLDVDCCAVVAYGALLKPELLA
** * :.:.:* * ** *: **: :** ******** **.
Mvan_2687|M.vanbaalenii_PYR-1 VPAHGWVNLHFSVLPAWRGAAPVQAALAAGDEVTGATTFQIERSLDSGPV
Mflv_3730|M.gilvum_PYR-GCK VPALGWVNLHFSVLPAWRGAAPVQAALAAGDEVTGATTFQIELSLDSGPV
Mb1441|M.bovis_AF2122/97 VPPHGWVNLHFSLLPAWRGAAPVQAAIAAGDTITGATTFQIEPSLDSGPI
Rv1406|M.tuberculosis_H37Rv VPPHGWVNLHFSLLPAWRGAAPVQAAIAAGDTITGATTFQIEPSLDSGPI
MLBr_00552|M.leprae_Br4923 VPPRGWVNLHFSLLPAWRGAAPVQAAIAAGDTITGATTFQIEPSLDSGPV
MAV_3372|M.avium_104 VPPHGWINLHFSLLPAWRGAAPVQAAIAAGDTITGATTFRIEPALDSGPI
MMAR_2215|M.marinum_M VPPQGWVNLHFSLLPAWRGAAPVQAAIAAGDAVTGATTFQIEPSLDSGPV
MUL_1802|M.ulcerans_Agy99 VPPQGWVNLHFSLLPAWRGAAPVQAAIAAGDAVTGATTFQIEPSLDSGPV
MSMEG_3064|M.smegmatis_MC2_155 VPRHGWINLHFSLLPAWRGAAPVQAAIAAGDTVTGATTFQIEPALDSGPV
TH_1529|M.thermoresistible__bu VPRHGWVNLHFSLLPAWRGAAPVQAAIAAGDTVTGASTFRIEPALDSGPI
MAB_2811c|M.abscessus_ATCC_199 VPRLGWVNLHFSLLPAWRGAAPVQASIAAGDEITGATTFLIEPALDSGPV
** **:*****:************::**** :***:** ** :*****:
Mvan_2687|M.vanbaalenii_PYR-1 YGVVTETIRPTDTAGDLLERLSVSGAGLLEATMDGIEDGTLTAVPQPPEG
Mflv_3730|M.gilvum_PYR-GCK YGVVTETIRPTDTAGDLLGRLAESGAGLLEATMDGIEDGTLTAVPQPAEG
Mb1441|M.bovis_AF2122/97 YGVVTEVIQPTDTAGDLLKRLAVSGAALLSTTLDGIADQRLTPRPQPADG
Rv1406|M.tuberculosis_H37Rv YGVVTEVIQPTDTAGDLLKRLAVSGAALLSTTLDGIADQRLTPRPQPADG
MLBr_00552|M.leprae_Br4923 YGVVTETIQPTDTAGDLLERLAVSGATLLSSTLDGIADAILTPRQQPVDG
MAV_3372|M.avium_104 YGVVTEAIRPTDTAGELLARLAVSGAELLSATLDGIADSTLTPRPQPAEG
MMAR_2215|M.marinum_M YGVVTETIRPTDTAGDLLGRLAVSGAELLSATLDGIAEGALTARPQPADG
MUL_1802|M.ulcerans_Agy99 YGVVTETIRPTDTAGDLLGRLAVSGAELLSATLDGIAEGALTARPQPADG
MSMEG_3064|M.smegmatis_MC2_155 YGVVTETVRDTDTAGDLLERLSDSGAELLERTIDGIADGSLTAVPQPSEG
TH_1529|M.thermoresistible__bu YGVVTETIRPTDTAGDLLDRLADSGAELLERTLDGIEDGTLTAVPQPSEG
MAB_2811c|M.abscessus_ATCC_199 YGVVTERISPNDTAGALLGRLAESGAGLLESTMDGIEDGALVAVPQAVDG
****** : .**** ** **: *** **. *:*** : *.. *. :*
Mvan_2687|M.vanbaalenii_PYR-1 VSVAPKVTVDDARIRWELPAHVVDRRVRSVTPNPGAWTMIGDQRIKVGPV
Mflv_3730|M.gilvum_PYR-GCK VSIAPKVSVDDARIRWELPAHVVDRRIRSVTPNPGAWTMAGELRIKVGPV
Mb1441|M.bovis_AF2122/97 VSVAPKITVANARVRWDLPAAVVERRIRAVTPNPGAWTLIGDLRVKLGPV
Rv1406|M.tuberculosis_H37Rv VSVAPKITVANARVRWDLPAAVVERRIRAVTPNPGAWTLIGDLRVKLGPV
MLBr_00552|M.leprae_Br4923 VSFAPKITVEQARVCWDLPAPVVERRIRAVTPNPGAWTLVGKLRVKLGPV
MAV_3372|M.avium_104 VSIAPKITVEQARVRWDLPAPVVERRIRAVTPNPGAWTVIGDLRIKLGPV
MMAR_2215|M.marinum_M VTLAPKISVEQARVRWELPAPIIERRIRAVTPNPGAWTLIGDLRVKLGPV
MUL_1802|M.ulcerans_Agy99 VTLAPKISVEQARVRWELPVPIIERRIRAVTPNPGAWTLIGDLRVKLGPV
MSMEG_3064|M.smegmatis_MC2_155 ITVAPKITVESARVRWDLPAHVVDRRIRAVTPNPGAWTMIGELRVKVGPV
TH_1529|M.thermoresistible__bu VSYAPKITVEAARIRWDLPAHVVERRIRAMTPNPGAWTMIGDLRVKLGPV
MAB_2811c|M.abscessus_ATCC_199 VSIAPKITVEQARIRWELPAHAIDRHIRAMTPEPGAWTNIGGTRVKVGPV
:: ***::* **: *:**. ::*::*::**:***** * *:*:***
Mvan_2687|M.vanbaalenii_PYR-1 TVAG------DGPKGLEPGEIRVDKK-----HIHVGTGSDAVLLGTVQPP
Mflv_3730|M.gilvum_PYR-GCK TVPD------DGPKDLEPGEIRVGKK-----HVHVGTATDAVLLGTVQPP
Mb1441|M.bovis_AF2122/97 HLDA----AHRPSKPLPPGGIHVERT-----SVWIGTGSEPVRLGQIQPP
Rv1406|M.tuberculosis_H37Rv HLDA----AHRPSKPLPPGGIHVERT-----SVWIGTGSEPVRLGQIQPP
MLBr_00552|M.leprae_Br4923 RFDSGAVEVPRLLKPLLPGGIHVDHK-----SVWIGTGSDPVRLSKVQPQ
MAV_3372|M.avium_104 RLGAAS-DLPAPPEPLPPGAIHVDRK-----SVWVGTASDPVRLDQIQPP
MMAR_2215|M.marinum_M YLDA----AVKPPGPLPPGAIQVDRK-----HVWVGTGSEPLRLGQVQPP
MUL_1802|M.ulcerans_Agy99 YLDA----AVEPPGPLPPGAIQVDRK-----HVWVGTGSEPLRLGQVQPP
MSMEG_3064|M.smegmatis_MC2_155 TVDQ----AAEADGPLAPGEIRVGRN-----SVHVGTGSHPVRLGQIQPP
TH_1529|M.thermoresistible__bu TVDD----DAE---PLAPGEIRAGRDGSGKQAVWFGTGTHPVRLGWIQPP
MAB_2811c|M.abscessus_ATCC_199 GVAT--------DEQLPPGAVAVRKR-----DVLVGTGTTAIALDEVQPQ
. * ** : . : : .**.: .: *. :**
Mvan_2687|M.vanbaalenii_PYR-1 GKKPMNAADWARGARLDENGWAR
Mflv_3730|M.gilvum_PYR-GCK GKKSMNAADWARGARAEDIRRAR
Mb1441|M.bovis_AF2122/97 GKKLMNAADWARGARLDLAARAT
Rv1406|M.tuberculosis_H37Rv GKKLMNAADWARGARLDLAARAT
MLBr_00552|M.leprae_Br4923 GKKFMNAVDWAHGARLDPAARAS
MAV_3372|M.avium_104 GKKFMNAVDWARGARLDPAARAT
MMAR_2215|M.marinum_M GKKLMNAADWARGARLDPSVRAS
MUL_1802|M.ulcerans_Agy99 GKKLMNAADWARGARLDPSVRAS
MSMEG_3064|M.smegmatis_MC2_155 GKKLMNAADWARGARLEEPVSAS
TH_1529|M.thermoresistible__bu GKKPMNAPDWARGARLDDTARAR
MAB_2811c|M.abscessus_ATCC_199 GKKVMKAADWARGARLDAEVHAL
*** *:* ***:*** : *