For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VLRERFGVSERRACTVVGLHRSTMRLTPAPIATEEAELRAWLRRFSTDRPRWGWRRAAKMARKAGWKANN KRIRRLWREEGLRVPQRRRKKRLTGIGVAVGAMSPIRPNVIWAMDFQFDTTADGRILKMLNVIDEFTREA LAIEVDRAINADGVVDVLDRLALTHGAPHYVRFDNGPEFVTHAVSDWCRFNSAGSLFIDPGSPWQNAWIE SFNGRLRDELLNSWRFDSLLEARVIIEDWRCDYNANRPHSAHGELTPAEFALQWTTTHQPQVA
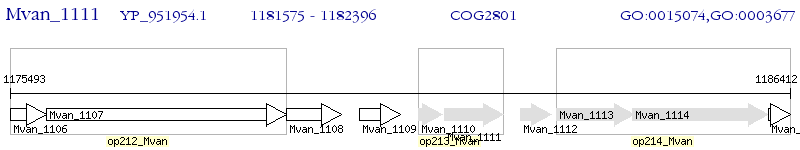
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_1111 | - | - | 100% (273) | integrase catalytic subunit |
| M. vanbaalenii PYR-1 | Mvan_1112 | - | 1e-76 | 93.75% (144) | integrase, catalytic region |
| M. vanbaalenii PYR-1 | Mvan_0580 | - | 1e-39 | 100.00% (75) | integrase, catalytic region |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | - | - | - | - | - |
| M. gilvum PYR-GCK | Mflv_4763 | - | 1e-157 | 96.32% (272) | integrase catalytic subunit |
| M. tuberculosis H37Rv | - | - | - | - | - |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_2088 | - | 1e-07 | 23.14% (229) | transposase |
| M. marinum M | MMAR_3784 | - | 1e-147 | 90.48% (273) | transposase, ISMyma01_aa2-like protein |
| M. avium 104 | - | - | - | - | - |
| M. smegmatis MC2 155 | MSMEG_1866 | - | 8e-57 | 44.77% (277) | transposase B |
| M. thermoresistible (build 8) | TH_3317 | - | 2e-07 | 22.65% (234) | PUTATIVE Integrase, catalytic region |
| M. ulcerans Agy99 | MUL_2691 | - | 1e-05 | 43.10% (58) | transposase ISMmr01_aa2-like |
CLUSTAL 2.0.9 multiple sequence alignment
Mvan_1111|M.vanbaalenii_PYR-1 --------------------------------------------------
Mflv_4763|M.gilvum_PYR-GCK MPASGDHRVDVAPLDCPVRRHESQRGQTAQGTRSRERPAQEAGRQPGPRH
MMAR_3784|M.marinum_M --------------------------------------------------
MSMEG_1866|M.smegmatis_MC2_155 --------------------------------------------------
MUL_2691|M.ulcerans_Agy99 --------------------------------------------------
MAB_2088|M.abscessus_ATCC_1997 --------------------------------------------------
TH_3317|M.thermoresistible__bu --------------------------------------------------
Mvan_1111|M.vanbaalenii_PYR-1 ----------------------MLRERFGVSERRACTVVGLHRSTMRLTP
Mflv_4763|M.gilvum_PYR-GCK RHAQGDFGGKLLTPNRKRSAVIALRERFGVSERRACTVVGLHRSTMRLTP
MMAR_3784|M.marinum_M ----------------------MLRERFGVSERRACTVVGLHRSTMRLQS
MSMEG_1866|M.smegmatis_MC2_155 ----------------------MLTTTMGMSERLACKAVGLARSTCRRLP
MUL_2691|M.ulcerans_Agy99 ---------------MRTGTVRHLQRVLAVSERFACRVTGQQRATQRHES
MAB_2088|M.abscessus_ATCC_1997 ------------------------------MTRRVLKLARQPYYRWRANP
TH_3317|M.thermoresistible__bu ---------MIAFIDAHRDQFGVELICRVLRAAIAGFLTSRGYRAAKARP
. . : .
Mvan_1111|M.vanbaalenii_PYR-1 APIA--TEEAELRAWLRRFSTDRPRWGWRRAAKMARKAG-WKANNKRIRR
Mflv_4763|M.gilvum_PYR-GCK APVT--TEEAELRAWLRRFSTDRPRWGWRRAAKMARRAG-WKANNKRIRR
MMAR_3784|M.marinum_M APIT--TEEAELRAWLRRFSTDWPRWGWRRAAKMARRAG-WAVNNKRIRR
MSMEG_1866|M.smegmatis_MC2_155 LAETPADPDAEMRGWLRSYATKHPCHGFRRAWAALRYDERREVNKKKIHR
MUL_2691|M.ulcerans_Agy99 CAAPSEGPDGALRDWLRQYAKDHPRRGFRPAYYDVRAET----------A
MAB_2088|M.abscessus_ATCC_1997 ITDAELVEAYRANALFDAHGED-PEFGYRYLVEEAREAG-EPMAERTAWR
TH_3317|M.thermoresistible__bu PSERAMRDELLIAELRTVHEQNFSVYGVKKMHHAMKRRG-WRVGREQTRR
. . . . * : :
Mvan_1111|M.vanbaalenii_PYR-1 LWREEGLRVPQRRR--KKRLTGIGVAVG------AMSPIRPNVIWAMDFQ
Mflv_4763|M.gilvum_PYR-GCK LWREEGLRVPQRRR--KKRLTGIGVAVG------AMSPIRPNVIWAMDFQ
MMAR_3784|M.marinum_M LWREEGLRVPQRRK--KKRLTGIGVEVG------AMSPICPNVIWAMDFQ
MSMEG_1866|M.smegmatis_MC2_155 LWREEGLQVRVHSP--RKRAG--VSSIP------PIEADAPNVVWAIDFQ
MUL_2691|M.ulcerans_Agy99 LLRK----------------------------------------------
MAB_2088|M.abscessus_ATCC_1997 ICSQQQLWSVFGKK--RGKNGKPGPPVHDDLVQRDFTAETPNMLWLSDIT
TH_3317|M.thermoresistible__bu LMRKAGLRGVQRGKPVFTTISDPAHCRPADLVNRQFSAEAPDRLWVADIT
: :
Mvan_1111|M.vanbaalenii_PYR-1 FDTTADGRILKMLNVIDEFTREALAIEVDRAIN-ADGVVDVLDRLALTHG
Mflv_4763|M.gilvum_PYR-GCK FDTTADGRTLKMLNVIDEFTREALAIEVDRAIN-ADGVVDVLDRLALTYG
MMAR_3784|M.marinum_M FDTTADGRTLKMLNVIDEFTREALAIEVDRAIN-ADGVVAVLDRLTAQRA
MSMEG_1866|M.smegmatis_MC2_155 FDSTIDGKAIKIASMIDEHTRVSLLNIVERSIT-ADRLVEELKKVFVAAG
MUL_2691|M.ulcerans_Agy99 --------------------------------------------------
MAB_2088|M.abscessus_ATCC_1997 EHYTGDGK-LYLCAIKDAFSNRIVGYSIDSRMK-SRLATAALRNAVGRRG
TH_3317|M.thermoresistible__bu FVRTWQGFCYTAF-VTDACTKAIKGWAVAASMRTEDLPLEAFNHAVWHSD
Mvan_1111|M.vanbaalenii_PYR-1 --APHYVRFDN-GPEFVTHAVSDWCRFNSAGSLFIDPGSPWQNAWIESFN
Mflv_4763|M.gilvum_PYR-GCK --APHYVRFDN-GPEFVANAVADWCRFNSAGSLFIDPGSPWQNAWIESFN
MMAR_3784|M.marinum_M --TPHYVRFDN-GPEFVAHAVSDWCRFNSAGSLFIDPGSPWQNAWIESFN
MSMEG_1866|M.smegmatis_MC2_155 G-PPRVLRMDN-GPEFISQALQQFCDG-RIGMSYIPPGTPWNNGHIESFN
MUL_2691|M.ulcerans_Agy99 --------------------------------------------------
MAB_2088|M.abscessus_ATCC_1997 V-VAGCVVHTDRGSQFRSRKFVQALHHHGMVGSMGRVGAAGDNAAMESFF
TH_3317|M.thermoresistible__bu SDLSELIHHSDRGSQYLSLTYTDRLAELGIAPSVGSRGDSYDNALAEAVN
Mvan_1111|M.vanbaalenii_PYR-1 GRLRDELLNSWR-FDSLLEARVIIEDWRC-DYNANRPHSAHGELTPAEFA
Mflv_4763|M.gilvum_PYR-GCK GRLRDELLNLWR-FDSLLEARVIIEDWRR-DYNANRPHSAHGELTPAEFA
MMAR_3784|M.marinum_M GRLRDELLNSWR-FDSLLEAQVLIEDWRR-DYNANRPHTAHGELTPTEFA
MSMEG_1866|M.smegmatis_MC2_155 NRLRKECLNRNH-WNTLLEARVVIGDFKD-EHNHRHRHSALGYRTPAEYA
MUL_2691|M.ulcerans_Agy99 --------------------------------------------------
MAB_2088|M.abscessus_ATCC_1997 SLLQKNVLDRRR-WATREELRIAIVTWIERTYHRRRRQGALGRLTPVEYE
TH_3317|M.thermoresistible__bu AAYKTELINRGKPWRSVDDVELATAGWVA-WYNQERLHEALGYVPPAEYE
Mvan_1111|M.vanbaalenii_PYR-1 LQWTTTHQPQVA--------
Mflv_4763|M.gilvum_PYR-GCK LQWTTTHQPQVA--------
MMAR_3784|M.marinum_M LQWTTTHQPQAA--------
MSMEG_1866|M.smegmatis_MC2_155 AVCSCTHTPVACSIN-----
MUL_2691|M.ulcerans_Agy99 --------------------
MAB_2088|M.abscessus_ATCC_1997 AIMTTTASQAA---------
TH_3317|M.thermoresistible__bu AALTGTSHPASQPTPALATN