For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MTVVGAVLPELKLYGDPTFIVSTALATRDFQDVHHDRDKAVAQGSKDIFVNILTDTGLVQRYVTDWAGPS ALIKSIGLRLGVPWYAYDTVTFSGEVTAVNDGLITVKVVGRNTLGDHVTATVELSMRDS
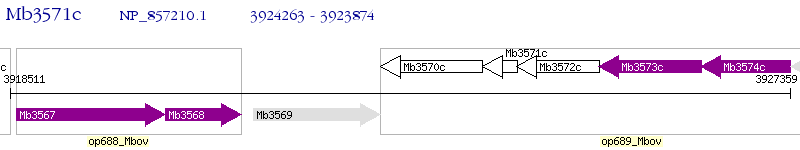
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3571c | - | - | 100% (129) | hypothetical protein Mb3571c |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. gilvum PYR-GCK | Mflv_1512 | - | 2e-55 | 78.40% (125) | dehydratase |
| M. tuberculosis H37Rv | Rv3541c | - | 9e-71 | 100.00% (129) | hypothetical protein Rv3541c |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_0617 | - | 3e-51 | 73.81% (126) | hypothetical protein MAB_0617 |
| M. marinum M | MMAR_5028 | - | 2e-58 | 86.99% (123) | hypothetical protein MMAR_5028 |
| M. avium 104 | MAV_0620 | - | 9e-62 | 88.10% (126) | hypothetical protein MAV_0620 |
| M. smegmatis MC2 155 | MSMEG_5991 | - | 9e-56 | 78.46% (130) | MaoC like domain-containing protein |
| M. thermoresistible (build 8) | TH_0365 | - | 5e-56 | 80.95% (126) | CONSERVED HYPOTHETICAL PROTEIN |
| M. ulcerans Agy99 | MUL_4102 | - | 2e-59 | 87.80% (123) | hypothetical protein MUL_4102 |
| M. vanbaalenii PYR-1 | Mvan_5254 | - | 2e-57 | 81.60% (125) | MaoC-like dehydratase |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_1512|M.gilvum_PYR-GCK ---MSAPSIEVGTTLPELTIYGDPTFIVSTAIATRDYQDVHHDRDKAQAK
Mvan_5254|M.vanbaalenii_PYR-1 ---MSAPALEVGTKLPELKIYGDPTFIVSTAIATRDYQDVHHDRDKAQAK
MSMEG_5991|M.smegmatis_MC2_155 MGAQSMTVIEVGTALPELAIYGDPTFIVSTAIATRDYQDVHHDRDKAQAK
TH_0365|M.thermoresistible__bu ---MSTPVVEVGTKLPELKIHADPTFIVSTALATRDFQDVHHDRDKAQAK
Mb3571c|M.bovis_AF2122/97 -------MTVVGAVLPELKLYGDPTFIVSTALATRDFQDVHHDRDKAVAQ
Rv3541c|M.tuberculosis_H37Rv -------MTVVGAVLPELKLYGDPTFIVSTALATRDFQDVHHDRDKAVAQ
MAV_0620|M.avium_104 ---MSAPVIEVGTTLPELKLYGDPTFIISTALATRDFQDVHHDRDKAQAK
MMAR_5028|M.marinum_M ---MSAPTMDVGTKLPELKLHGDPTFIVSTALATRDFQDVHHDRDKAQAK
MUL_4102|M.ulcerans_Agy99 --------MDVGTKLPELKLHGDPTFIVSTALATRDFQDVHHDRDKAQAK
MAB_0617|M.abscessus_ATCC_1997 -------MTTVGEKLPELKLEGTPTFIVSTALATRDFQDVHHDRDLAQAK
** **** : . ****:***:****:******** * *:
Mflv_1512|M.gilvum_PYR-GCK GSKDIFVNILTDTGLVQRYLTDWAGPAARIRSIGLRLGVPWYAYDTITFT
Mvan_5254|M.vanbaalenii_PYR-1 GSKDIFVNILTDTGLVQRYLTDWAGPTARIKSIGLRLGVPWYAYDTITFT
MSMEG_5991|M.smegmatis_MC2_155 GSKDIFVNILTDTGLVQRYLTDWAGPNARIKSIGLRLGVPWYAYDTVTFR
TH_0365|M.thermoresistible__bu GSKDIFVNILTDTGLVQRYLTDWAGPAARIKSIKLRLGVPWYAYDTVTFT
Mb3571c|M.bovis_AF2122/97 GSKDIFVNILTDTGLVQRYVTDWAGPSALIKSIGLRLGVPWYAYDTVTFS
Rv3541c|M.tuberculosis_H37Rv GSKDIFVNILTDTGLVQRYVTDWAGPSALIKSIGLRLGVPWYAYDTVTFS
MAV_0620|M.avium_104 GSKDIFVNILTDTGLVQRYVTDWAGPSALIKSIGLRLGVPWYAYDTVTFS
MMAR_5028|M.marinum_M GSKDIFVNILTDTGLVQRYVTDWAGPSALIKSIGLRLGVPWYAYDTVIFS
MUL_4102|M.ulcerans_Agy99 GSKDIFVNILTDTGLVQRYVTDWAGPSALIKSIGLRLGLPWYAYDTVTFS
MAB_0617|M.abscessus_ATCC_1997 GSKDIFINILSDTGLVERFVTDWAGPAARVKSISLRLGVPWYAYDTTVFT
******:***:*****:*::****** * ::** ****:******* *
Mflv_1512|M.gilvum_PYR-GCK GEVTAVDDDIATVKVVGANSLGNHVIATATLALGDH-
Mvan_5254|M.vanbaalenii_PYR-1 GEVTAVDDGVATVKVLGSNSLGDHVIATATLSLGDR-
MSMEG_5991|M.smegmatis_MC2_155 GEVTAIEDDLVTVKVTGSNSLGNHVIATATLNLGDKA
TH_0365|M.thermoresistible__bu GEVTDVVDGLVTVKVVGSNSLGDHIIATTTLTMGES-
Mb3571c|M.bovis_AF2122/97 GEVTAVNDGLITVKVVGRNTLGDHVTATVELSMRDS-
Rv3541c|M.tuberculosis_H37Rv GEVTAVNDGLITVKVVGRNTLGDHVTATVELSMRDS-
MAV_0620|M.avium_104 GEVTAVQDGLVTLKVFGRNSLGDHVIATVTLTMGDA-
MMAR_5028|M.marinum_M GEVTTIEDGLITVKVVGSNSLGNHVIATVTLTVGGA-
MUL_4102|M.ulcerans_Agy99 GEVTAIEDGLITVKVVGSNSLGNHVIATVTLTVGGA-
MAB_0617|M.abscessus_ATCC_1997 GEVAAVEDGVVEVDVVGNNSLGAHVTAKVKLTIGAEQ
***: : *.: :.* * *:** *: *.. * :