For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
SDAPRGEDAVLVAVQAALAGRPGVLTGARALSHFGEHSAGWVALAAAGALAQPAKRRSWLAVGAGAFLAH AAAVVIKRVVRRERPSHPDIAVNVGTPSRLSFPSAHATSTTAAAVLLAQTTGVPAPALLVPPMALSRLVL GVHYPTDVVTGVVVGALVGKAVGKAAGRISNSVGAEGDR*
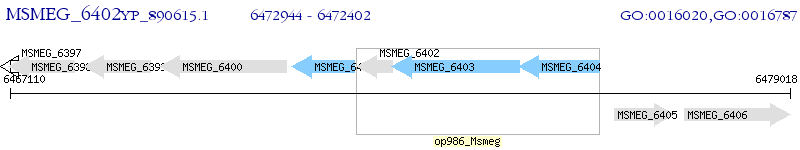
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_6402 | - | - | 100% (180) | PAP2 superfamily protein |
| M. smegmatis MC2 155 | MSMEG_0633 | - | 2e-10 | 36.96% (138) | PAP2 superfamily protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3837c | - | 1e-57 | 66.24% (157) | transmembrane protein |
| M. gilvum PYR-GCK | Mflv_1161 | - | 3e-68 | 77.11% (166) | phosphoesterase, PA-phosphatase related |
| M. tuberculosis H37Rv | Rv3807c | - | 1e-57 | 66.24% (157) | transmembrane protein |
| M. leprae Br4923 | MLBr_00094 | - | 2e-53 | 59.06% (171) | hypothetical protein MLBr_00094 |
| M. abscessus ATCC 19977 | MAB_0172 | - | 5e-62 | 69.01% (171) | phosphoesterase |
| M. marinum M | MMAR_5371 | - | 3e-57 | 67.72% (158) | transmembrane protein |
| M. avium 104 | MAV_0210 | - | 3e-56 | 72.54% (142) | PAP2 superfamily protein |
| M. thermoresistible (build 8) | TH_2991 | - | 7e-64 | 70.52% (173) | PUTATIVE phosphoesterase, PA-phosphatase related |
| M. ulcerans Agy99 | MUL_4991 | - | 3e-56 | 66.46% (158) | transmembrane protein |
| M. vanbaalenii PYR-1 | Mvan_5648 | - | 3e-71 | 75.84% (178) | phosphoesterase, PA-phosphatase related |
CLUSTAL 2.0.9 multiple sequence alignment
MSMEG_6402|M.smegmatis_MC2_155 -------MSDAPRGEDAVLVAVQAALAGRPGVLTGARALSHFGEHSAGWV
TH_2991|M.thermoresistible__bu ---MREQMAEPLGREEAALVTIQAALAGRPGVLAAARALSHFGEHSLGWL
Mflv_1161|M.gilvum_PYR-GCK -------MADSPRGEDAVLVAVQSTLGARPGVLPVARAMSHFGEHSLGWL
Mvan_5648|M.vanbaalenii_PYR-1 -------MADAPRGEDAVLVAVQRALADRPGVLAVARGLSHFGEHSLGWL
MAB_0172|M.abscessus_ATCC_1997 MTGPLPGEVTAPQGETAVLVAVQSALGGRPGVLSTARGLSHFGEHSIGWV
MAV_0210|M.avium_104 -------------------------------MLTVARALSHFGEHSIGWV
Mb3837c|M.bovis_AF2122/97 ------------------MVAVQSALVDRPGMLATARGLSHFGEHCIGWL
Rv3807c|M.tuberculosis_H37Rv ------------------MVAVQSALVDRPGMLATARGLSHFGEHCIGWL
MMAR_5371|M.marinum_M -----MPEPAPPRGEAAAIVAVQSALADRPGVLGTARGLSHFGEHSVGWL
MUL_4991|M.ulcerans_Agy99 -----MPEPAPPRGEAAAIVAVQSALADRPGVLGTARGLSHFGEHSVGWL
MLBr_00094|M.leprae_Br4923 -----MPEDKAPTGELAAIAAVQSVLVDRPGVLPTARGMSHFGEHSIGWL
:* **.:******. **:
MSMEG_6402|M.smegmatis_MC2_155 ALAAAGALAQPA---KRRSWLAVGAGAFLAHAAAVVIKRVVRRERPSHPD
TH_2991|M.thermoresistible__bu GLAGLGALTQPA---RRRSWLAVGVGAFAAHAAAVLIKRVVRRRRPEHEA
Mflv_1161|M.gilvum_PYR-GCK ALAGIGALVSPR---QRSDYLAAGAGAFAAHAAAVVIKRVVRRTRPHHPA
Mvan_5648|M.vanbaalenii_PYR-1 AVAALGAVLSPR---RRRDYVTAGAGALLAHAAAVVIKRVVRRQRPHHPA
MAB_0172|M.abscessus_ATCC_1997 AAGLIGAAINRRDKPRRRSWLAAGAGAFGAHAASVIIKRLVRRRRPSHEA
MAV_0210|M.avium_104 AAAALGALCSPR---RRADWLAAGAGAFVAHAAAVVVKRIVRRERPHHPA
Mb3837c|M.bovis_AF2122/97 ILALLGAIALPR---RRREWLVAGAGAFVAHAIAVLIKRLVRRQRPDHPA
Rv3807c|M.tuberculosis_H37Rv ILALLGAIALPR---RRREWLVAGAGAFVAHAIAVLIKRLVRRQRPDHPA
MMAR_5371|M.marinum_M LVSALGALLFPS---RRREWVVAGAGTFVAHAAAVVIKRVVRRKRPHDPA
MUL_4991|M.ulcerans_Agy99 LVSALGAMLFPS---RRREWVVAGAGTFVAHAAAVVIKRVVRRKRPHDPA
MLBr_00094|M.leprae_Br4923 AISLLGAILVPC---RRRYWLVAGAGVFAAHVAAVLIKRMVRRIRPNHPA
. ** :* ::..*.*.: **. :*::**:*** ** .
MSMEG_6402|M.smegmatis_MC2_155 IAVNVGTPSRLSFPSAHATSTTAAAVLLAQTTGVP---APALLVPPMALS
TH_2991|M.thermoresistible__bu IRVHVATPSRLSFPSAHATSTTAAALLLARTTGSV---LPLLLIPPMAVS
Mflv_1161|M.gilvum_PYR-GCK IAVNVATPSRLSFPSAHATSTTAAALLLARPTGLP---LPALLIPPMALS
Mvan_5648|M.vanbaalenii_PYR-1 IAVNVGTPSRLSFPSAHATSTTAAAVLLARPTGLP---LPALLVPPMALS
MAB_0172|M.abscessus_ATCC_1997 VRVNVSTPSRLSFPSSHATSTAAAAVLFAPLTGLP---LPALLIPPMALS
MAV_0210|M.avium_104 VVVNVGTPSRLSFPSAHATSTTAAAILLGRITGLP---LPAVLVPPMALS
Mb3837c|M.bovis_AF2122/97 IAVNVDTPSQLSFPSAHATSTTAAALLMGRATGLP---LPVVLVPPMALS
Rv3807c|M.tuberculosis_H37Rv IAVNVDTPSQLSFPSAHATSTTAAALLMGRATGLP---LPVVLVPPMALS
MMAR_5371|M.marinum_M VTVNVGTPSKLSFPSAHATSTAAAAILLSRAAGVPEAAGAAVLVPPMALS
MUL_4991|M.ulcerans_Agy99 VTVNVGTPSKLSFPSAHATSTAAAAILLSRAAGVSEAAGAAVLVPPMALS
MLBr_00094|M.leprae_Br4923 VTVNVGTPSPLSFPSAHATSTAAAAILIGRASRLPKGIVAAVLVAPMALS
: *:* *** *****:*****:***:*:. : . :*:.***:*
MSMEG_6402|M.smegmatis_MC2_155 RLVLGVHYPTDVVTGVVVGALVGKAVGKAAGRISNS---VGAEGDR----
TH_2991|M.thermoresistible__bu RLVLGVHYPTDVLTGVAVGAATATAVDLVTRKEADG---L-SEADR----
Mflv_1161|M.gilvum_PYR-GCK RLVLGVHYPTDVLAGVVVGAAVAKAVERA----------LEGATT-----
Mvan_5648|M.vanbaalenii_PYR-1 RLVLGVHYPSDVLAGVVVGAVVGKAVGRLGDTLADR---AEGVSSS----
MAB_0172|M.abscessus_ATCC_1997 RLVLGVHYPTDVAAGAALGALIGAAVRRADSRLALK---EEQR-------
MAV_0210|M.avium_104 RMVLGVHYPSDVACGIALGAGVAAVARRAP---------QLARGDQ----
Mb3837c|M.bovis_AF2122/97 RILLGVHYPSDVAVGVALGATVGAIVDSVGGGRQ------RARKR-----
Rv3807c|M.tuberculosis_H37Rv RILLGVHYPSDVAVGVALGATVGAIVDSVGGGRQ------RARKR-----
MMAR_5371|M.marinum_M RILLGVHYPSDVAAGIAVGASVAAIAARLD-KVS-----TKGMSL-----
MUL_4991|M.ulcerans_Agy99 RILLGVHYPSDVAAGIAVGASVAAIAARLD-KVS-----TKGMSL-----
MLBr_00094|M.leprae_Br4923 RIVLGVHYPSDVAFGVVLGAAVAGTTARFDSRLSRRWTVQHGLSSGSAVK
*::******:** * .:** . .