For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MEGFAGKVAVVTGAGSGIGQALAIELGRSGAKLAICDVDTEGLAATEERLKAIGAPVKADRLDVTEREAF LLYADAVKEHYGTVNQIYNNAGIAFTGDVEISEFKDIERVMDVDYWGVVNGTKAFLPHLIASGDGHVINV SSVFGLFSVPGQAAYNSAKFAVRGFTEALRQEMAIAKHPVKVTTVHPGGIKTAIARNATAAEGLDAKELA EAFDKKLANTTPQRAAVIILDGVRKNKARVLVGPDAKILDVIVRITGSGYQRLFSAVAAKAMPR
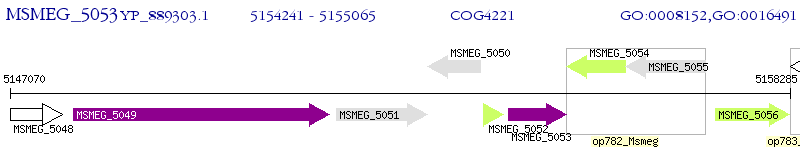
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_5053 | - | - | 100% (274) | short chain alcohol dehydrogenase |
| M. smegmatis MC2 155 | MSMEG_2872 | - | 2e-35 | 36.33% (256) | short-chain dehydrogenase/reductase SDR |
| M. smegmatis MC2 155 | MSMEG_2291 | - | 5e-35 | 33.72% (258) | short chain dehydrogenase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1277c | - | 1e-130 | 83.21% (274) | short-chain type dehydrogenase/reductase |
| M. gilvum PYR-GCK | Mflv_2216 | - | 1e-124 | 79.49% (273) | short-chain dehydrogenase/reductase SDR |
| M. tuberculosis H37Rv | Rv1245c | - | 1e-130 | 83.21% (274) | short-chain type dehydrogenase/reductase |
| M. leprae Br4923 | MLBr_01094 | - | 1e-118 | 79.34% (271) | short chain alcohol dehydrogenase |
| M. abscessus ATCC 19977 | MAB_1388c | - | 1e-116 | 74.45% (274) | short-chain dehydrogenase/reductase |
| M. marinum M | MMAR_4195 | - | 1e-121 | 80.22% (273) | short-chain type dehydrogenase/reductase |
| M. avium 104 | MAV_1384 | - | 1e-130 | 84.98% (273) | short chain alcohol dehydrogenase |
| M. thermoresistible (build 8) | TH_1828 | - | 1e-130 | 83.58% (274) | PROBABLE SHORT-CHAIN TYPE DEHYDROGENASE/REDUCTASE |
| M. ulcerans Agy99 | MUL_4501 | - | 1e-121 | 79.49% (273) | short-chain type dehydrogenase/reductase |
| M. vanbaalenii PYR-1 | Mvan_4479 | - | 1e-131 | 84.25% (273) | short-chain dehydrogenase/reductase SDR |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_2216|M.gilvum_PYR-GCK MQGFAGKVAVVTGAGSGIGQALAIELGRSGALVAISDVDTEGLAVTEERL
Mvan_4479|M.vanbaalenii_PYR-1 MEGFAGKVAVVTGAGSGIGQALAIELGRSGARVAISDVDTEGLAVTEERL
TH_1828|M.thermoresistible__bu MQGFAGKVAVITGAGSGIGQALALELGRSGAKVAISDVNTEGLTETEDRL
MSMEG_5053|M.smegmatis_MC2_155 MEGFAGKVAVVTGAGSGIGQALAIELGRSGAKLAICDVDTEGLAATEERL
Mb1277c|M.bovis_AF2122/97 MEGFAGKVAVVTGAGSGIGQALAIELARSGAKVAISDVDTDGLADTEHRL
Rv1245c|M.tuberculosis_H37Rv MEGFAGKVAVVTGAGSGIGQALAIELARSGAKVAISDVDTDGLADTEHRL
MLBr_01094|M.leprae_Br4923 MEGFAGKVAVVTGAGSGIGQALAIELARSGAKLAISDVDGEGLAQTEGQL
MMAR_4195|M.marinum_M MEGFAGKVAVVTGAGSGIGRALAIELARSGAKLAISDVDTEGLAQTEKLV
MUL_4501|M.ulcerans_Agy99 MEGFAGKVAVVTGAGSGIGRALAIELARSGAKLAISDVDTEGLAQTEKLV
MAV_1384|M.avium_104 MQGFAGKVAVVTGAGSGIGQALAVELARSGAKVAISDVDLEGLAHTEEQL
MAB_1388c|M.abscessus_ATCC_199 MEGFSGKVAVVTGAGSGIGRALAVELARSGAKIAISDIDNEGLAETERQV
*:**:*****:********:***:**.**** :**.*:: :**: ** :
Mflv_2216|M.gilvum_PYR-GCK KAIGTHVKSDRLNVTERERFLQYADDVAEHFGKVHQIYNNAGIAFTGDIE
Mvan_4479|M.vanbaalenii_PYR-1 KAIGAQVKSDRLDVTERETFLLYADDVAEHFGSVNQIYNNAGIAFTGDIE
TH_1828|M.thermoresistible__bu KAIGAPVKADRLDVTERAAFEAYADAVVEHFGVVNQIYNNAGIAYTGDIE
MSMEG_5053|M.smegmatis_MC2_155 KAIGAPVKADRLDVTEREAFLLYADAVKEHYGTVNQIYNNAGIAFTGDVE
Mb1277c|M.bovis_AF2122/97 KAISTPVKTDRLDVTEREAFLAYADAVNEHFGTVNQIYNNAGIAFTGDIE
Rv1245c|M.tuberculosis_H37Rv KAISTPVKTDRLDVTEREAFLAYADAVNEHFGTVNQIYNNAGIAFTGDIE
MLBr_01094|M.leprae_Br4923 KAIGASARTDRLDVTEREAFLTYADVVHENFGKVNQIYNNAGIAFTGDVE
MMAR_4195|M.marinum_M TALGAEVKTDRLDVTEREAFLAYADAVNEHFGKVNQIYNNAGIGHTGDVE
MUL_4501|M.ulcerans_Agy99 TALGAEVKTDRLDVTERETFLAYADAVNEHFGKVNQIYNNAGIGHTGDVE
MAV_1384|M.avium_104 KAIGAQYKADRLDVTEREAFLAYADAVKEHFGKVNQIYNNAGIAFTGDVE
MAB_1388c|M.abscessus_ATCC_199 KALGAEVRADRLNVAEREAFLLYADAIKDHFGKVNQIYNNAGIAYQGEVE
.*:.: ::***:*:** * *** : :::* *:********.. *::*
Mflv_2216|M.gilvum_PYR-GCK VSQFKDIERVMDVDFWGVVNGTKAFLPHLIASGDGHVVNVSSVFGLFSVP
Mvan_4479|M.vanbaalenii_PYR-1 ITQFKDIERVMDVDYWGVVNGTKAFLPHLIASGDGHVVNVSSVFGLFSVP
TH_1828|M.thermoresistible__bu VSHFKDIERVMDVDFWGVVNGTKVFLPHLIASGDGHVINISSVFGLFSVP
MSMEG_5053|M.smegmatis_MC2_155 ISEFKDIERVMDVDYWGVVNGTKAFLPHLIASGDGHVINVSSVFGLFSVP
Mb1277c|M.bovis_AF2122/97 VSQFKDIERVMDVDFWGVVNGTKAFLPHLIASGDGHVINISSVFGLFSAP
Rv1245c|M.tuberculosis_H37Rv VSQFKDIERVMDVDFWGVVNGTKAFLPHLIASGDGHVINISSVFGLFSAP
MLBr_01094|M.leprae_Br4923 VSHFKDIERVMDVDYWGVVNGTKAFLPYLISSGDGHVINISSVFGLFSVP
MMAR_4195|M.marinum_M VCAFKDIDRVMDVDFGGVLNGTKAFLPYLIASGDGHVINVSSVFGLFSVS
MUL_4501|M.ulcerans_Agy99 VCAFKDIDRVMDVDFGGVLNGTKTFLPYLIASGDGHVINVSSVFGLFSVS
MAV_1384|M.avium_104 VSQFKDIERVMDVDFWGVVNGTKAFLPHLIASGDGHVVNVSSVFGLFSVP
MAB_1388c|M.abscessus_ATCC_199 ESQFKDIERIIDVDFWGVVNGTKAFLPHLIASGDGHVVNVSSLFGVLSMP
****:*::***: **:****.***:**:******:*:**:**::* .
Mflv_2216|M.gilvum_PYR-GCK GQGAYNAAKFAVRGFTEALRQEMALAGHPVKVSCVHPGGIKTAIARNAGA
Mvan_4479|M.vanbaalenii_PYR-1 GQAAYNSAKFAVRGFTEALRQEMALAGHPVKVSCVHPGGIKTAIARNASA
TH_1828|M.thermoresistible__bu GQAAYNAAKFAVRGFTEALRQEMILAGHPVKVTTVHPGGIKTGIARNMTA
MSMEG_5053|M.smegmatis_MC2_155 GQAAYNSAKFAVRGFTEALRQEMAIAKHPVKVTTVHPGGIKTAIARNATA
Mb1277c|M.bovis_AF2122/97 GQAAYNSAKFAVRGFTEALRQEMALAGHPVKVTTVHPGGVKTAIARNATA
Rv1245c|M.tuberculosis_H37Rv GQAAYNSAKFAVRGFTEALRQEMALAGHPVKVTTVHPGGVKTAIARNATA
MLBr_01094|M.leprae_Br4923 GQAAYNSAKFAVRGFTEALREEMALAGRPVNVTTVYPGGIKTAIARNATA
MMAR_4195|M.marinum_M GQAAYNAAKFAVRGFTEALRQEMILAGHPVGVTTVHPGGIKTAIARNATA
MUL_4501|M.ulcerans_Agy99 GQAAYNAAKFAVRGFTEALRQEMILAGHPVGVTTVHPGGIKTAIARNATA
MAV_1384|M.avium_104 GQAAYNSAKFAVRGFTEALRQEMAAAGHPVAVTTVHPGGIKTAIARNATA
MAB_1388c|M.abscessus_ATCC_199 GQSAYNSAKFAVRGFTESLRQEMIIGKKPVAVTCVHPGGIKTAIARNSTT
**.***:**********:**:** . :** *: *:***:**.**** :
Mflv_2216|M.gilvum_PYR-GCK AEGLDAEELAKTFDKKLASTTPEKAAQVILDGVRKNKARILIGNDAKMFE
Mvan_4479|M.vanbaalenii_PYR-1 AEGLDAEELAKAFDKKLASTTPEKAAKIILDGVRKNRARILVGNDAKVFD
TH_1828|M.thermoresistible__bu AEGLDAQELARAFDTKLARTSPEKAAQIILDGVRKNKARVLVGTDAKILD
MSMEG_5053|M.smegmatis_MC2_155 AEGLDAKELAEAFDKKLANTTPQRAAVIILDGVRKNKARVLVGPDAKILD
Mb1277c|M.bovis_AF2122/97 AEGLDQAELAETFDKRVAHLSPQRAAQIILTGVAKNKARVLVGVDAKVLD
Rv1245c|M.tuberculosis_H37Rv AEGLDQAELAETFDKRVAHLSPQRAAQIILTGVAKNKARVLVGVDAKVLD
MLBr_01094|M.leprae_Br4923 AEGLDVSKIASRFDTWVAHTSPQHAARIILKAVRKKKARVLVGPDAKVAN
MMAR_4195|M.marinum_M AEGLDSDELAKMFDKRVARTSPERAAKIILGAVRKNKARVLVGPDAKALD
MUL_4501|M.ulcerans_Agy99 AEGLDSDELAKMFDKRVARTSPERAAKIILGAVRKNKARVLVGPDAKALD
MAV_1384|M.avium_104 AEGLDQAELAKLFDKRLAKTTPQRAAQIILDAVRKKKARVLVGSDAKALD
MAB_1388c|M.abscessus_ATCC_199 AEGYDQQAMAAMFDKYLANTSPEAAARIILTAVRKKKPRVLVGPDAKILD
*** * :* **. :* :*: ** :** .* *::.*:*:* *** :
Mflv_2216|M.gilvum_PYR-GCK IAIRALG-AGYQSVFPKVVARLTPPAK--
Mvan_4479|M.vanbaalenii_PYR-1 VVVRVLG-AGYQSLFPKAVARLTPPAK--
TH_1828|M.thermoresistible__bu AIVRLTG-SGYQRLFPKFVARTMPR----
MSMEG_5053|M.smegmatis_MC2_155 VIVRITG-SGYQRLFSAVAAKAMPR----
Mb1277c|M.bovis_AF2122/97 LVVRLTG-SGYQRIFPIITGRLIPRPR--
Rv1245c|M.tuberculosis_H37Rv LVVRLTG-SGYQRIFPIITGRLIPRPR--
MLBr_01094|M.leprae_Br4923 VVVRFSGGAGYQRLFAQVASRLILNQR--
MMAR_4195|M.marinum_M IVVRLTG-SGYQRLFMPVLGRLVPASHR-
MUL_4501|M.ulcerans_Agy99 IVVRLTG-SGYQRLFMPVLGRLVPASHR-
MAV_1384|M.avium_104 ILVRLTG-SGYQRLFGPVMSRLLPN----
MAB_1388c|M.abscessus_ATCC_199 VIVRLTG-ARYQDIFSVVTRFILPRPGKK
:* * : ** :*