For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MSDDLRVTTAHLRELSAKQGRAAAELATATAVVDGVDTALRFTHGPISWGTAAAVEAVQHARRAAGTGMV KVSQELETKLDTAAGRYHRTDSTMGDALDETIQPR
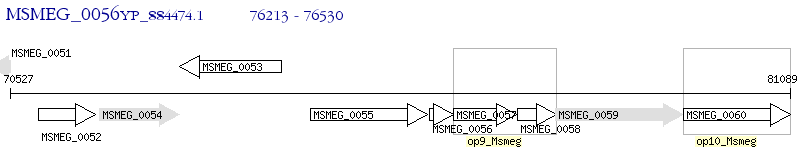
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_0056 | - | - | 100% (105) | hypothetical protein MSMEG_0056 |
| M. smegmatis MC2 155 | MSMEG_0077 | - | 1e-14 | 41.41% (99) | hypothetical protein MSMEG_0077 |
| M. smegmatis MC2 155 | MSMEG_0728 | - | 6e-10 | 36.08% (97) | hypothetical protein MSMEG_0728 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3895 | - | 3e-06 | 25.00% (100) | hypothetical protein Mb3895 |
| M. gilvum PYR-GCK | Mflv_0157 | - | 2e-09 | 33.66% (101) | hypothetical protein Mflv_0157 |
| M. tuberculosis H37Rv | Rv3865 | - | 3e-06 | 25.00% (100) | hypothetical protein Rv3865 |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | - | - | - | - | - |
| M. marinum M | MMAR_1326 | - | 4e-10 | 35.29% (102) | hypothetical protein MMAR_1326 |
| M. avium 104 | - | - | - | - | - |
| M. thermoresistible (build 8) | TH_3927 | - | 2e-07 | 41.94% (62) | PUTATIVE - |
| M. ulcerans Agy99 | MUL_2553 | - | 3e-10 | 35.64% (101) | hypothetical protein MUL_2553 |
| M. vanbaalenii PYR-1 | Mvan_0655 | - | 4e-11 | 33.66% (101) | hypothetical protein Mvan_0655 |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_1326|M.marinum_M --------------------------------------------------
MUL_2553|M.ulcerans_Agy99 --------------------------------------------------
MSMEG_0056|M.smegmatis_MC2_155 --------------------------------------------------
Mflv_0157|M.gilvum_PYR-GCK MALGKLWVDANYFYGVADHANMMSSDPALGSAAIATDAAILMQSLAAVGG
TH_3927|M.thermoresistible__bu --------------------------------------------------
Mvan_0655|M.vanbaalenii_PYR-1 --------------------------------------------------
Mb3895|M.bovis_AF2122/97 --------------------------------------------------
Rv3865|M.tuberculosis_H37Rv --------------------------------------------------
MMAR_1326|M.marinum_M --------------------------------------------------
MUL_2553|M.ulcerans_Agy99 --------------------------------------------------
MSMEG_0056|M.smegmatis_MC2_155 --------------------------------------------------
Mflv_0157|M.gilvum_PYR-GCK STMGATGLATSVATPIINAGLMTLTAASNTLGFGRPEDGERFTDGARQFS
TH_3927|M.thermoresistible__bu --------------------------------------------------
Mvan_0655|M.vanbaalenii_PYR-1 --------------------------------------------------
Mb3895|M.bovis_AF2122/97 --------------------------------------------------
Rv3865|M.tuberculosis_H37Rv --------------------------------------------------
MMAR_1326|M.marinum_M --------------------------------------------------
MUL_2553|M.ulcerans_Agy99 --------------------------------------------------
MSMEG_0056|M.smegmatis_MC2_155 --------------------------------------------------
Mflv_0157|M.gilvum_PYR-GCK EANDELQQAVPPPDWQGDASDAYGLRNTEQQQRAGSMATFDGQIQEVLAE
TH_3927|M.thermoresistible__bu --------------------------------------------------
Mvan_0655|M.vanbaalenii_PYR-1 --------------------------------------------------
Mb3895|M.bovis_AF2122/97 --------------------------------------------------
Rv3865|M.tuberculosis_H37Rv --------------------------------------------------
MMAR_1326|M.marinum_M ----------------------------------MANLVVTPQYLQTLAA
MUL_2553|M.ulcerans_Agy99 ----------------------------------MAKLVVTPQYLQTLAA
MSMEG_0056|M.smegmatis_MC2_155 ---------------------------------MSDDLRVTTAHLRELSA
Mflv_0157|M.gilvum_PYR-GCK EAGQVQDTRDFVSTRQVMLAAAIPAAITAKLIPPGGAAAAMAIELAAVAA
TH_3927|M.thermoresistible__bu --------------------------------------------------
Mvan_0655|M.vanbaalenii_PYR-1 ------------------------------MTSPDGELHVEPSHLRQLSA
Mb3895|M.bovis_AF2122/97 ---------------------------------MTGFLGVVPSFLKVLAG
Rv3865|M.tuberculosis_H37Rv ---------------------------------MTGFLGVVPSFLKVLAG
MMAR_1326|M.marinum_M TQTQAAATAASAGAASTN--------------------------------
MUL_2553|M.ulcerans_Agy99 TQTQAAATAASAGAASTN--------------------------------
MSMEG_0056|M.smegmatis_MC2_155 KQGRAAAELATATAVVDG--------------------------------
Mflv_0157|M.gilvum_PYR-GCK TVPFAMQRVTEMVSHSADNADRIRQIGSHYGVVDESAQMPGGLSGAQSGT
TH_3927|M.thermoresistible__bu --------------------------------------------------
Mvan_0655|M.vanbaalenii_PYR-1 RQNAAAAQIADAVPLTAD--------------------------------
Mb3895|M.bovis_AF2122/97 MHNEIVGDIKRATDTVAG--------------------------------
Rv3865|M.tuberculosis_H37Rv MHNEIVGDIKRATDTVAG--------------------------------
MMAR_1326|M.marinum_M ---------------------------------LKNEVWVSHGVISGASN
MUL_2553|M.ulcerans_Agy99 ---------------------------------LKNEVWVSHGVISGASN
MSMEG_0056|M.smegmatis_MC2_155 ---------------------------------VDTALRFTHGPISWGTA
Mflv_0157|M.gilvum_PYR-GCK PAPLRVSESIRGTAGIQQNVAATIDSARSAPDGSAGDVFKTHGVVCTSTA
TH_3927|M.thermoresistible__bu ---------------------------------VTGKVLQTHGIVCAPTI
Mvan_0655|M.vanbaalenii_PYR-1 ---------------------------------ESDHVLSSHGIVCMPTY
Mb3895|M.bovis_AF2122/97 ---------------------------------ISGRVQLTHGSFTSKFN
Rv3865|M.tuberculosis_H37Rv ---------------------------------ISGRVQLTHGSFTSKFN
: :** .
MMAR_1326|M.marinum_M TSFANAEAARRAAANSMNTTSSKLAAKLRTASSAYQGVDEQAGESLDKQI
MUL_2553|M.ulcerans_Agy99 TSFANAEAARRAAANSMNTTSSKLAAKLRTASSAYQGVDEQAGESLDKQI
MSMEG_0056|M.smegmatis_MC2_155 AAVEAVQHARRAAGTGMVKVSQELETKLDTAAGRYHRTDSTMGDALDETI
Mflv_0157|M.gilvum_PYR-GCK AALKSAELARRSAAEKMHLISTDLSAKLGSSATEYSHTEQQNADAVDRQM
TH_3927|M.thermoresistible__bu AALGAAQMSRSNAAKAMERISNDLAEKLDTAAAEYTRTDQQGQQTLDGEM
Mvan_0655|M.vanbaalenii_PYR-1 LAAQAAGVARAKAGVAVQVMSEAFAANLDSAAANYSWTDQDSGSNIDGKM
Mb3895|M.bovis_AF2122/97 DTLQEFETTRSSTGTGLQGVTSGLANNLLAAAGAYLKADDGLAGVIDKIF
Rv3865|M.tuberculosis_H37Rv DTLQEFETTRSSTGTGLQGVTSGLANNLLAAAGAYLKADDGLAGVIDKIF
: :* :. : : : :* ::: * .:. :* :
MMAR_1326|M.marinum_M IER--
MUL_2553|M.ulcerans_Agy99 IER--
MSMEG_0056|M.smegmatis_MC2_155 QPR--
Mflv_0157|M.gilvum_PYR-GCK PPR--
TH_3927|M.thermoresistible__bu RPG--
Mvan_0655|M.vanbaalenii_PYR-1 RPGRR
Mb3895|M.bovis_AF2122/97 G----
Rv3865|M.tuberculosis_H37Rv G----