For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
SSAALSYPLFDADNHLYETEESLTKYLPKQYRGVIQYVQVNGRTKIAIRGQISDYIPNPTFEVVARPGAM EEYFRIGNPDGKSRREIFGEPMRSIPAFREPGPRLELMNELGVDRSLMFPTLASLIEERMRDDPLLIHIV VHALNEWLHEEWGFRYQDRIFTVPVVSLPIVEKAIEELDWVVERGARAVLVRPAPVPGYRGPRSFALPEF DPFWHRCVEHDVLVAMHSSDSGYARYTAEWDGGDKEMLPFQTNAFSMLNEWRPVQDAVASWVIHGALYRV PRLKVAVIEAGSKWLFPLLDQLADVYKKTPESFPGGDPVEEIKNRIHVSPFYEDGIADLIKLIGTDRVLY GSDYPHPEGLAQPRHYADALQDLSVDDQAKIMGGNLARLMSV*
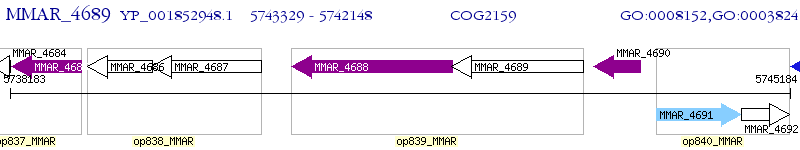
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. marinum M | MMAR_4689 | - | - | 100% (393) | hypothetical protein MMAR_4689 |
| M. marinum M | MMAR_3988 | - | e-139 | 58.88% (394) | metal-dependent hydrolase |
| M. marinum M | MMAR_0138 | - | 2e-79 | 39.28% (387) | hypothetical protein MMAR_0138 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | - | - | - | - | - |
| M. gilvum PYR-GCK | Mflv_2524 | - | 0.0 | 73.90% (387) | amidohydrolase 2 |
| M. tuberculosis H37Rv | - | - | - | - | - |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_0299 | - | 1e-07 | 22.67% (300) | putative amidohydrolase (aminocarboxymuconate-semialdehyde |
| M. avium 104 | MAV_0970 | - | 0.0 | 75.91% (386) | amidohydrolase family protein |
| M. smegmatis MC2 155 | MSMEG_4773 | - | 0.0 | 75.97% (387) | amidohydrolase family protein |
| M. thermoresistible (build 8) | TH_0534 | - | 0.0 | 76.55% (388) | amidohydrolase family protein |
| M. ulcerans Agy99 | MUL_0308 | - | 0.0 | 99.49% (393) | hypothetical protein MUL_0308 |
| M. vanbaalenii PYR-1 | Mvan_4134 | - | 0.0 | 74.68% (387) | amidohydrolase 2 |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_4689|M.marinum_M ----MSSAALSYPLFDADNHLYETEESLTKYLPKQYRGVIQYVQVNGRTK
MUL_0308|M.ulcerans_Agy99 ----MSSAALSYPLFDADNHLYETEESLTKYLPKQYRGVIQYVQVNGRTK
Mflv_2524|M.gilvum_PYR-GCK MGQLSHRVEIPFPLFDADNHLYEPPEAMTKYLPKEYKDIVQYVEINGRTK
Mvan_4134|M.vanbaalenii_PYR-1 MGQLSHRVDIPFPLFDADNHLYEPPEAMTKYLPKEYKDVVQYVEVNGRTK
MSMEG_4773|M.smegmatis_MC2_155 MGQLSHRVDIPFPLFDADNHLYEPPEAMTKYLPKDYKDVVQYVEVNGRTK
TH_0534|M.thermoresistible__bu MGQLSHRVELPFPLFDADNHLYEPPEAMTKYLPKDYKDVVQYVQVNGRTK
MAV_0970|M.avium_104 MGQLSHREDVPFPIFDADNHLYEPPEALTKFLPKEYKDFVQYVQINGRTK
MAB_0299|M.abscessus_ATCC_1997 ---------MTSVRIDIHAHLWS-------------DEYLSLLDGYGRPA
:. :* . **:. :. :: **.
MMAR_4689|M.marinum_M IAIRGQISDYIPNPTFEVVARPGAMEEYFRIGNPDGKSRREIFGEPMRSI
MUL_0308|M.ulcerans_Agy99 IAIRGQISDYIPNPTFEVVARPGAMEEYFRIGNPDGKSRREIFGEPMRSI
Mflv_2524|M.gilvum_PYR-GCK IALKGQISNYIPNPTFSVVAKPGAWEEYFKFGNPDGKSKRELFGEPMKAI
Mvan_4134|M.vanbaalenii_PYR-1 IALKGQISNYIPNPTFSVVAKPGAWEEYFKFGNPDGKSKRELFGEPMKAI
MSMEG_4773|M.smegmatis_MC2_155 IAINGQISNYIPNPTFSVVAKPGAWEEYFKFGNPDGKSKRELFGEPMRSI
TH_0534|M.thermoresistible__bu IAIRGQISNYIPNPTFEVVARPGAWEEYFKFGNPEGKSKRELFGEPMRSI
MAV_0970|M.avium_104 IALRGVISNYIPNPTFEVVARPGAWEEYFKYGNPEGKSKRELFGEPMRAI
MAB_0299|M.abscessus_ATCC_1997 TDVHRGLG--------------------------AGATRDDLDG------
:. :. * :: :: *
MMAR_4689|M.marinum_M PAFREPGPRLELMNELGVDRSLMFPTLASLIEERMRDDPLLIHIVVHALN
MUL_0308|M.ulcerans_Agy99 PAFREPGPRLELMNELGVDRSLMFPTLASLIEERMRDDPLLIHIVVHALN
Mflv_2524|M.gilvum_PYR-GCK PAFFEPEPRIKVMDELGIQRSLMFPTLASLIEERLSDDPVAIHVIIHALN
Mvan_4134|M.vanbaalenii_PYR-1 PAFFEPEPRIKVMDELGIQRSLMFPTLASLIEERLSDDPVAIHVIIHALN
MSMEG_4773|M.smegmatis_MC2_155 PAFFEPEPRLELMDQLGVDRSLMFPTLASLIEERLRDDPVAIHVIIHSLN
TH_0534|M.thermoresistible__bu PAFFEPGPRLELMDELGVDRTLMFPTLASLIEERLRDDPVAIHVLIHALN
MAV_0970|M.avium_104 PAFFEPGPRLEKMNELGLDRTLMFPTLASLIEERLRDDPVAIHVLIHALN
MAB_0299|M.abscessus_ATCC_1997 --------RFELMDEAGVDLQILTATPAS---PHFEDQAAAVASARFIND
*:: *:: *:: :: .* ** :: *:. : . :
MMAR_4689|M.marinum_M EWLHEEWGFRYQDRIFTVPVVSLPIVEKAIEELDWVVERGARAVLVRPAP
MUL_0308|M.ulcerans_Agy99 EWLHEEWGFRYQDRIFTVPVVSLPIVEKAIEELDWVVERGARAVLVRPAP
Mflv_2524|M.gilvum_PYR-GCK EWLHEVWGFNYQGRIFTTPVITLPIVEKAIEELEWAVKRGARAILIRPAP
Mvan_4134|M.vanbaalenii_PYR-1 EWLHEVWGFNYQGRIFTTPVITLPIVEKAIEELEWAVKRGARAILIRPAP
MSMEG_4773|M.smegmatis_MC2_155 LWLDEVWGFNYKNRIFTTPVITLPIVEKAIEELEWAVSRGARAILIRPAP
TH_0534|M.thermoresistible__bu QWLDEVWGFNYKNRIFTTPVITLPIVEKAIEELEWCVKRGARAILIRPAP
MAV_0970|M.avium_104 QWLDEVWGFNYQNRIFTTPVITLPIVEKAIEELEWVVKRGARAILVRPAP
MAB_0299|M.abscessus_ATCC_1997 EYAKLSS--EYPGRFGALASLPLPHVPAALEELARAIDELG----MYGAS
: . .* .*: : . :.** * *:*** :.. . : *.
MMAR_4689|M.marinum_M VPGYRGPRSFALPEFDPFWHRCVEHDVLVAMHSSDSGYARYTAEWDGGDK
MUL_0308|M.ulcerans_Agy99 VPGYRGPRSFALPEFDPFWHRCVEHDVLVAMHSSASGYARYTAEWDGGDK
Mflv_2524|M.gilvum_PYR-GCK VPGFRGPRSFALPEFDPFWERVVHHDIFVGMHSSDSGYSRYTSEWDGAAQ
Mvan_4134|M.vanbaalenii_PYR-1 VPGFRGPRSFALPEFDPFWERVVHHDIFVGMHSSDSGYSRYTSEWDGAAQ
MSMEG_4773|M.smegmatis_MC2_155 VPGFRGPRSFALPEFDPFWERVVHHDVFVGMHSSDSGYSRYTSEWDGAAQ
TH_0534|M.thermoresistible__bu VPGFRGPRSFALPEFDPFWERVVEYDVFVGMHSSDSGYSRYTSEWDGSQQ
MAV_0970|M.avium_104 VPGFRGPRSFALPEFDPFWERVVEYDVLVGMHSSDSGYSRYTSEWDGADQ
MAB_0299|M.abscessus_ATCC_1997 ITTSVLGRSIADPIFDPLYAELNRRRAVLFVHPAGCG----------AES
:. **:* * ***:: . . .: :*.: .* . .
MMAR_4689|M.marinum_M EMLPFQTNAFSMLNEWRPVQD--AVASWVIHGALYRVPRLKVAVIEAGSK
MUL_0308|M.ulcerans_Agy99 EMLPFQTNAFSMLNEWRPVQD--AVASWVIHGALYRVPRLKVAVIEAGSK
Mflv_2524|M.gilvum_PYR-GCK EMLPFQTNAMSILNEWRPIQD--AVASWVIHGALFRFPKLKVGIVEAGSK
Mvan_4134|M.vanbaalenii_PYR-1 EMLPFQTNAMSILNEWRPIQD--AVASWVIHGALFRFPKLKVGIVEAGSK
MSMEG_4773|M.smegmatis_MC2_155 EMLPFQTNAMSILNEWRPIQD--AVASWVIHGALFRHPKLKVGIVEAGSK
TH_0534|M.thermoresistible__bu EMLPFQTNAMAILNEWRPIQD--AVASWVIHGALFRHPKLKVGIVEAGSK
MAV_0970|M.avium_104 EMLPFQTNAMGILNEWRPIQD--SVASWVIHGALYRHPKLKVAIVEAGSK
MAB_0299|M.abscessus_ATCC_1997 SLITDHSLTWSIG---APIEDTVAIMHLIVAGVPSRYPDMKIVTCHLGGA
.::. :: : .: *::* :: :: *. * * :*: . *.
MMAR_4689|M.marinum_M WLFPLLDQLADVYKKTPESFPGGDPVEEIKNRIHVSPFYEDG--IADLIK
MUL_0308|M.ulcerans_Agy99 WLFPLLDQLADVYKKTPESFPGGDPVEEIKNRIHISPFYEDG--IADLIK
Mflv_2524|M.gilvum_PYR-GCK WMFPLLDSMAEVYKKAPEAFLG-NPMEEIKNRIYVSPFYEEG--IDDLIN
Mvan_4134|M.vanbaalenii_PYR-1 WMFPLLDSMAEVYKKAPEAFLG-NPIEEIKNRIYVSPFYEEG--IDDLIN
MSMEG_4773|M.smegmatis_MC2_155 WMFPLLDSMAEVYKKAPEAFLG-NPIEEIKNRIYVSPFYEEG--IDDLIN
TH_0534|M.thermoresistible__bu WMFPLLDSMAEVYKKAPEAFLG-NPIEEIKNRIYVSPFYEEG--IDELIN
MAV_0970|M.avium_104 WMTPLLDGLAEVFRKAPEAFPS-DPVEMVKNRIHVSPFFEDG--IDDLVN
MAB_0299|M.abscessus_ATCC_1997 LPMVLERAHRQVTEWEATQCPE-APRDAARRLWYDTVGHDHTPALRAAVA
* :* . . * : :. : : .:. : :
MMAR_4689|M.marinum_M LIGTDRVLYGSDYPHPEGLAQPRHYADALQDLSVDDQAKIMGGNLARLMS
MUL_0308|M.ulcerans_Agy99 LIGTDRVLYGSDYPHPEGLAQPRHYADALQDLSVDDQAKIMGGNLARLMS
Mflv_2524|M.gilvum_PYR-GCK LIGVDQVLYGSDWPHPEGLAEPTHYVTALEHLAVEDQAKIMGGNLGRLVT
Mvan_4134|M.vanbaalenii_PYR-1 LIGVDQVLYGSDWPHPEGLAEPTHYVTALEHLSVEDQAKIMGGNLGRLVT
MSMEG_4773|M.smegmatis_MC2_155 LIGVDQVLYGSDWPHPEGLAEPTHYVTALEHLSVEDQAKIMGGNLGRLVT
TH_0534|M.thermoresistible__bu LVGVDQVLYGSDWPHPEGLAEPTHYVTALEHLSLEDQAKIMGGNLSRLVT
MAV_0970|M.avium_104 LVGVDQVLYGSDWPHPEGLAEPTYYVNALSHLPVEDQAKIMGGNLGRLVT
MAB_0299|M.abscessus_ATCC_1997 SLGVDRLIFGSDFPYQAGPRYVSSATYIEEGLSAEDASAILDHNGAELLG
:*.*::::***:*: * . . *. :* : *:. * ..*:
MMAR_4689|M.marinum_M V---
MUL_0308|M.ulcerans_Agy99 V---
Mflv_2524|M.gilvum_PYR-GCK T---
Mvan_4134|M.vanbaalenii_PYR-1 T---
MSMEG_4773|M.smegmatis_MC2_155 T---
TH_0534|M.thermoresistible__bu V---
MAV_0970|M.avium_104 V---
MAB_0299|M.abscessus_ATCC_1997 LTGV