For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VTARRIAAAVVSLAVTMPAVACSTPEPPDYKSIWTTSATTSTTATTAPPVPLSQYLESIGVSGKQIAPSD LKGLTVSIPTPPGWSPYEGPKINPETVIISKGGHYPTARLVVFLLRGNFDPADVARRGNDDAQLFENFKQ LDASSADFHGFPSSMMQGSYDLEGARMHSWNRIVIPTGPPPDNQRYLVQLTITSLASQAAGSASDIESII NGFVVAVK
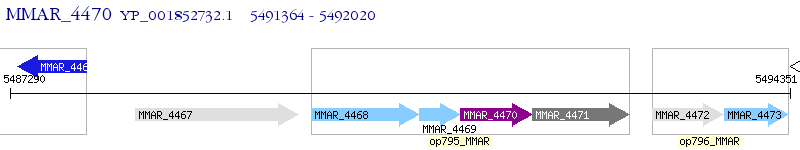
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. marinum M | MMAR_4470 | lpqT | - | 100% (218) | lipoprotein LpqT |
| M. marinum M | MMAR_0056 | mtc28 | 2e-20 | 31.89% (185) | secreted proline rich protein Mtc28 |
| M. marinum M | MMAR_0950 | lpqN | 7e-06 | 21.03% (214) | lipoprotein, LpqN |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1044c | lpqT | 6e-54 | 56.74% (178) | putative lipoprotein LpqT |
| M. gilvum PYR-GCK | Mflv_1941 | - | 8e-59 | 52.13% (211) | putative lipoprotein |
| M. tuberculosis H37Rv | Rv1016c | lpqT | 2e-69 | 59.63% (218) | lipoprotein LpqT |
| M. leprae Br4923 | MLBr_00246 | lpqT | 3e-80 | 67.45% (212) | putative lipoprotein |
| M. abscessus ATCC 19977 | MAB_1145c | - | 2e-25 | 28.90% (218) | putative lipoprotein LpqT precursor |
| M. avium 104 | MAV_1154 | - | 6e-81 | 69.34% (212) | LpqT protein |
| M. smegmatis MC2 155 | MSMEG_5429 | - | 2e-59 | 54.29% (210) | LpqT protein |
| M. thermoresistible (build 8) | TH_0908 | lpqT | 6e-60 | 51.83% (218) | PROBABLE CONSERVED LIPOPROTEIN LPQT |
| M. ulcerans Agy99 | MUL_4640 | lpqT | 1e-124 | 100.00% (218) | lipoprotein LpqT |
| M. vanbaalenii PYR-1 | Mvan_4785 | - | 5e-62 | 51.13% (221) | putative lipoprotein |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_4470|M.marinum_M MTARR--------IAAAVVSLAVTMPAVACSTPEPPDYKSIWTTSATTST
MUL_4640|M.ulcerans_Agy99 MTARR--------IAAAVVSLAVTMPAVACSTPEPPDYKSIWTTSATTST
MLBr_00246|M.leprae_Br4923 MQAIRL------GLHTAAAVVTLSISAVSCGT-KTPDYQLILSKSSTTTT
MAV_1154|M.avium_104 MNRLR-------GCAAVAAAVALAVATAGCAP-KPPDYQSILSKTPTTTT
Mb1044c|M.bovis_AF2122/97 MAGRRCPQDSVRPLAVAVAVATLAMSAVACGP-KSPDFQSILSTSPTTSA
Rv1016c|M.tuberculosis_H37Rv MAGRRCPQDSVRPLAVAVAVATLAMSAVACGP-KSPDFQSILSTSPTTSA
MSMEG_5429|M.smegmatis_MC2_155 MTIV---------AAVMSVLT-----VASCGA-ETPDYQSIWTTTTTTTT
TH_0908|M.thermoresistible__bu MSAARY-------AVALTLLAG---ALAGCGD-DIPDYQSIWTTTHTSAA
Mflv_1941|M.gilvum_PYR-GCK MRLWLT-------IALAAA-----LALTPCTT-ETPDYQSIWSTTSATPT
Mvan_4785|M.vanbaalenii_PYR-1 MTVRLR-------PALAAAAILAVPALSGCAA-ETPDYQSVWSTASTTSA
MAB_1145c|M.abscessus_ATCC_199 -----------MRSAALAAAAVAAFAMIGCGS-HNPGQPTGSASSTPESA
* . *. :.: . .:
MMAR_4470|M.marinum_M TATT----APPVPLSQYLESIGVSGKQIAPSDLKGLTVSIPTPPGWSPYE
MUL_4640|M.ulcerans_Agy99 TATT----APPVPLSQYLESIGVSGKQIAPSDLKGLTVSIPTPPGWSPYE
MLBr_00246|M.leprae_Br4923 TTPD-----KPIPLPQYLESIGVTGQQVAPSSLPGLTVSIPTPPGWSPYS
MAV_1154|M.avium_104 TTSAT---AKPVPLSQYLQSIGVTGQQVAPASLPDLTVSIPTPPGWAPFS
Mb1044c|M.bovis_AF2122/97 VSTTT---EVPVPLWKYLESVGVTGEPVAPSSLTDLTVSIPTPPGWAPMK
Rv1016c|M.tuberculosis_H37Rv VSTTT---EVPVPLWKYLESVGVTGEPVAPSSLTDLTVSIPTPPGWAPMK
MSMEG_5429|M.smegmatis_MC2_155 TPS----GEPPVPLWQHLESNGIVGEPVAPEKIPDLTVSMPTPPGWHPYN
TH_0908|M.thermoresistible__bu PTS----QEP-VPIHQYLESVGVVGRPVPPDEVRDFTITLPTPPGWQPYH
Mflv_1941|M.gilvum_PYR-GCK SPSAAADSAEPMSIATFLEQAGVAGQPVAPDKLTDLTVSMPTPKGWEPYR
Mvan_4785|M.vanbaalenii_PYR-1 APTSTDPDAPPMSIATFLEQSGVTGQPVAPDKLTDLTVSMQTPPGWHPYQ
MAB_1145c|M.abscessus_ATCC_199 APP-----AKPASMNEYLHSVGVTVTPVSPESPANVRVDVPLPQGWENLG
.. .: .*.. *: :.* . .. : : * **
MMAR_4470|M.marinum_M GPK---INPETVIISKGGHYPTARLVVFLLRGNFDPADVA-RRGNDDAQL
MUL_4640|M.ulcerans_Agy99 GPK---INPETVIISKGGHYPTARLVVFLLRGNFDPADVA-RRGNDDAQL
MLBr_00246|M.leprae_Br4923 NPN---ITPETLIIAKSGKYPTARLVAFKLRGDFDPTQVI-KHGNDDAQL
MAV_1154|M.avium_104 SPN---ISPQTVMISKGGKFPTARLVVFKLSGDFDPAQVI-KHGNDDAQL
Mb1044c|M.bovis_AF2122/97 NPN---ITPNTEMIAKGESYPTAMLMVFKLHRDFDIAEAL-KHGTADARL
Rv1016c|M.tuberculosis_H37Rv NPN---ITPNTEMIAKGESYPTAMLMVFKLHRDFDIAEAL-KHGTADARL
MSMEG_5429|M.smegmatis_MC2_155 NPN---LAPFTRMIAKGDTYPTAMLLAFKLYGDFDVKAALRDHGYADAQL
TH_0908|M.thermoresistible__bu NPN---FAPGTRVIAKGDTYPIAMLMVFELTGDFDVARAI-EHGHADAER
Mflv_1941|M.gilvum_PYR-GCK NSN---LSPGTRMIAKGETYPTAMLIVFELNGDFDVAQAL-SHANTDAET
Mvan_4785|M.vanbaalenii_PYR-1 NSN---LAPGTRVIAKGDTYPTAMLMVFELDGDFDVAEAL-THANVDAQA
MAB_1145c|M.abscessus_ATCC_199 ALDPAYLIADKPSGADQGRTPSAVVYLIKLGGPLDARKVISEHGFADAQN
. : . . :. * * : : * :* . :. **.
MMAR_4470|M.marinum_M FENFKQLDASSADFHGFPSSMMQGSYDLEGARMHSWNRIVIPTGPPPDNQ
MUL_4640|M.ulcerans_Agy99 FENFKQLDASSADFHGFPSSMMQGSYDLEGARMHSWNRIVIPTGPPPDNQ
MLBr_00246|M.leprae_Br4923 FENFRQLDVSTANYNGFPSAMIQGSYDLEGRRLHAWNRIVIPTGPPPSKQ
MAV_1154|M.avium_104 FENFKQLDASGAPYNGFPSSMIQGSYDLDGMRLHSWNRIVIATGSPPAHQ
Mb1044c|M.bovis_AF2122/97 STNFTELDSSTADFNGFPSSMIQGSYDLHGRRLHTWNRIVFPTGAGQAAL
Rv1016c|M.tuberculosis_H37Rv STNFTELDSSTADFNGFPSSMIQGSYDLHGRRLHTWNRIVFPTGAPPAKQ
MSMEG_5429|M.smegmatis_MC2_155 SENFKQLNASLDDWRGFPSAMIEGSYDLNGRRMQSYNRVVIPTTEQAPPQ
TH_0908|M.thermoresistible__bu SQNFKRLNASNENWRGFPSSMIEGSYDLNGRRMQSYNRIVIPTGPPPERQ
Mflv_1941|M.gilvum_PYR-GCK SENFTELNTSTADFGGFPSSMIEGSYDLSGTRMHSYNRIVIATGAPPTSQ
Mvan_4785|M.vanbaalenii_PYR-1 SENFRELNASTADFNGFPSSMIEGSYDLNGTRMHSYNRIVIATGAPPDRQ
MAB_1145c|M.abscessus_ATCC_199 TQNFQKIASSLDDYQGYPSAAIEGTYENQGVRVHAWNRDVIVPDGP---L
** .: * : *:**: ::*:*: * *::::** *: .
MMAR_4470|M.marinum_M RYLVQLTITSLAS--QAAGSASDIESIING-----FVVAVK---------
MUL_4640|M.ulcerans_Agy99 RYLVQLTITSLAS--QAAGSASDIESIING-----FVVAVK---------
MLBr_00246|M.leprae_Br4923 QYLVQLTITSLAN--EAVAQSNDIEAIIRG-----FVVAPK---------
MAV_1154|M.avium_104 RYLVQLTITSLAD--QAAAESTDIEAIIHG-----FVVAAK---------
Mb1044c|M.bovis_AF2122/97 PGAAHHHESGQRGRQARFRHRGDHRRIRRRGKVIVRTVGAQLSEQLTRIE
Rv1016c|M.tuberculosis_H37Rv RYLVQLTITSLAN--EAVKHASDIEAIIAG-----FVVAAK---------
MSMEG_5429|M.smegmatis_MC2_155 RYLIQLTVTTYAE--QAEAQAEDIEAIIAG-----FTVAKK---------
TH_0908|M.thermoresistible__bu RYLIQLTVTSFAE--DAAEHGPDIETIIRD-----FSVAPK---------
Mflv_1941|M.gilvum_PYR-GCK RYLVQFTVTGYAD--KAAEEAPDIEEIIRG-----FTVKVP---------
Mvan_4785|M.vanbaalenii_PYR-1 RYLVQFTVTGFAD--KAAEDGPDVESVIRG-----LTVALPR--------
MAB_1145c|M.abscessus_ATCC_199 HYLVQFTVTTTEA--QSAALSQDVAALTGG-----LKITKR---------
: : * : :
MMAR_4470|M.marinum_M ----------
MUL_4640|M.ulcerans_Agy99 ----------
MLBr_00246|M.leprae_Br4923 ----------
MAV_1154|M.avium_104 ----------
Mb1044c|M.bovis_AF2122/97 FPQCSSHGLA
Rv1016c|M.tuberculosis_H37Rv ----------
MSMEG_5429|M.smegmatis_MC2_155 ----------
TH_0908|M.thermoresistible__bu ----------
Mflv_1941|M.gilvum_PYR-GCK ----------
Mvan_4785|M.vanbaalenii_PYR-1 ----------
MAB_1145c|M.abscessus_ATCC_199 ----------