For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
SAKDHPNNAPGVPMVFPPWFERFQIKYINPALKPIARYLPGTATITHQGRKSGKPYKTIVTVYRKGNVLA IALGHGKTDWVKNVLAAGEADVFFARRQVHIVNPRILPAGSQSDELPMMARVQLRRIGVFVADIA*
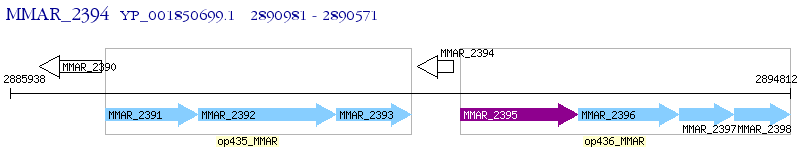
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. marinum M | MMAR_2394 | - | - | 100% (136) | hypothetical protein MMAR_2394 |
| M. marinum M | MMAR_3396 | - | 3e-14 | 33.91% (115) | hypothetical protein MMAR_3396 |
| M. marinum M | MMAR_3392 | - | 8e-10 | 36.78% (87) | hypothetical protein MMAR_3392 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1624c | - | 6e-62 | 81.62% (136) | hypothetical protein Mb1624c |
| M. gilvum PYR-GCK | Mflv_3614 | - | 5e-40 | 62.10% (124) | hypothetical protein Mflv_3614 |
| M. tuberculosis H37Rv | Rv1598c | - | 6e-62 | 81.62% (136) | hypothetical protein Rv1598c |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_2671 | - | 9e-36 | 56.91% (123) | hypothetical protein MAB_2671 |
| M. avium 104 | MAV_3188 | - | 5e-59 | 77.37% (137) | hypothetical protein MAV_3188 |
| M. smegmatis MC2 155 | MSMEG_3204 | - | 2e-38 | 57.02% (121) | hypothetical protein MSMEG_3204 |
| M. thermoresistible (build 8) | TH_1163 | - | 1e-41 | 59.68% (124) | CONSERVED HYPOTHETICAL PROTEIN |
| M. ulcerans Agy99 | MUL_1571 | - | 4e-75 | 98.53% (136) | hypothetical protein MUL_1571 |
| M. vanbaalenii PYR-1 | Mvan_2803 | - | 1e-41 | 59.50% (121) | hypothetical protein Mvan_2803 |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_2394|M.marinum_M MSAKDHPNNAPGVPMVFPPWFERFQIKYINPALKPIARYLPGTATITHQG
MUL_1571|M.ulcerans_Agy99 MSAKDHPNNAPGVPMVFPPWFERFQIKYINPALKPIARCLPGTATITHQG
Mb1624c|M.bovis_AF2122/97 MSAKDHPNNAPGVPMVFPLWLERLQVKYINRALKPIARYLPGTATIEHRG
Rv1598c|M.tuberculosis_H37Rv MSAKDHPNNAPGVPMVFPLWLERLQVKYINRALKPIARYLPGTATIEHRG
MAV_3188|M.avium_104 MSAKDHPNNAPGVPMVFPVWFENLQVKYLNSAIKPLARFLPGTATIKHRG
Mflv_3614|M.gilvum_PYR-GCK ---------MSDLPRMFPEWLDRIQIKYMNPVIKPLAGKVPGISVIKHRG
TH_1163|M.thermoresistible__bu ---------MTDLPRMFPPWLDRIQIRYFNPVIKPFAGRLPGLSLLRHRG
Mvan_2803|M.vanbaalenii_PYR-1 --------------MIFPPWFEKLQIKYINPVIRPLSKRMPGLGVIKHRG
MSMEG_3204|M.smegmatis_MC2_155 --------------MLFPPWLERLQLKYVNPVMTPVAKRLPGFTVIKHRG
MAB_2671|M.abscessus_ATCC_1997 --------------MLLPEWLERAQIKYINPLMAPIARFLPSFATVTHYG
::* *::. *::*.* : *.: :*. : * *
MMAR_2394|M.marinum_M RKSGKPYKTIVTVYRKGNVLAIALGHGKTDWVKNVLAAGEADVFFAR-RQ
MUL_1571|M.ulcerans_Agy99 RKSGKPYKTIVTVYRKGNVLAIAPGHGKTDWVKNVLAAGEADVFFAR-RQ
Mb1624c|M.bovis_AF2122/97 RKSGKPYQTIVTAYRKDGVLAIALAHGKTDWVKNVLAAGEADVHFAR-GV
Rv1598c|M.tuberculosis_H37Rv RKSGKPYQTIVTAYRKDGVLAIALAHGKTDWVKNVLAAGEADVHFAR-GV
MAV_3188|M.avium_104 RTSGKQYETIITPYRKGNTLAIALGHGKTNWVKNVLAAGEADVVFGRDRS
Mflv_3614|M.gilvum_PYR-GCK RTSGRDYQTIVTAYRGDGRLAIVLGHGKTDWVKNVLAAGEADVQLFR-DQ
TH_1163|M.thermoresistible__bu RRSGRDYETVVTTYRRGNELAIVLGHGKTDWVKNVLAAGEADVVVRG-RE
Mvan_2803|M.vanbaalenii_PYR-1 RTSGTEYETIVTPYRKGTVLAIGLAHGKTNWVKNVLAAGEADIQIGK-KT
MSMEG_3204|M.smegmatis_MC2_155 RKSGRAYETVVNSYRKDNVFAVILAHGKTNWVKNVLAAGGAEVKVFG-RE
MAB_2671|M.abscessus_ATCC_1997 RTSGTKYETTVNAFRKGNVVSIALIHGKTNWTKNVIAAGGADITLFGGRQ
* ** *:* :. :* . .:: ****:*.***:*** *:: .
MMAR_2394|M.marinum_M VHIVNPRILPAGSQSDELPMMARVQLRRIGVFVADIA-
MUL_1571|M.ulcerans_Agy99 VHIVNPRILPAGSQSDELPMMARVQLRRIGVFVADIA-
Mb1624c|M.bovis_AF2122/97 VHVINPRIVPAGSDGQGLPRMARLQLRRIGVFVGDIA-
Rv1598c|M.tuberculosis_H37Rv VHVINPRIVPAGSDGQGLPRMARLQLRRIGVFVGDIA-
MAV_3188|M.avium_104 VHLINPRILPAGSDGAGLPFMARVQLRWMGVFVADIA-
Mflv_3614|M.gilvum_PYR-GCK VHIVNPRIEPNGDGVTALPAIARRQAKKIAVFVADIA-
TH_1163|M.thermoresistible__bu WHVVNPRIVPPGGEVAALPGFARLQAKNSAVFVADIA-
Mvan_2803|M.vanbaalenii_PYR-1 LHLVRPRVLPAGTEDPSLPRTAQMLAKRSGVFVADIV-
MSMEG_3204|M.smegmatis_MC2_155 FEITNPRILPAGSDDPSLPRIARLSARKAGILVAEIV-
MAB_2671|M.abscessus_ATCC_1997 IHVVNPRIVEQGQGDPALTRATRQVNKRAGVFVAEIAQ
.: .**: * *. :: : .::*.:*.