For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VEVRSGYATSGDLKLYYEDMGNVDDPPVLLIMGLGAQLVLWRTAFCEKLVAQGLRVVRYDNRDVGLSSRT DSTEQPCPSQPLTARLIRFWLGQRNACAYTLEDMTDDAVALLDHLSIERAHIVGASMGGMIAQIFAARFP TRTRSLAVFFSSNNRPFLPPPAPRALLALLTGPPTGSRRDVVVDNVVRVTKITGSPLYRMPEEQVRTNAA EIYDRSFYPLGVSRQFSAILGSGSLLHYNQRIIAPTVVIHGRADKLVRPSSGRAVARAITGARLVLFDGM GHDLPQQLWDQAVGVLMSNFAKAS
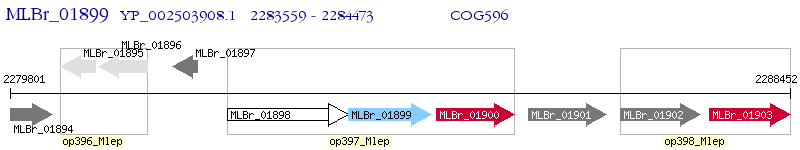
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. leprae Br4923 | MLBr_01899 | lipG | - | 100% (304) | putative hydrolase |
| M. leprae Br4923 | MLBr_02269 | - | 1e-08 | 23.88% (289) | putative hydrolase |
| M. leprae Br4923 | MLBr_00862 | ephD | 1e-06 | 23.18% (289) | short chain dehydrogenase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0665c | lipG | 1e-125 | 71.62% (303) | lipase/esterase LipG |
| M. gilvum PYR-GCK | Mflv_5112 | - | 1e-106 | 63.70% (303) | alpha/beta hydrolase fold |
| M. tuberculosis H37Rv | Rv0646c | lipG | 1e-125 | 71.62% (303) | lipase/esterase LipG |
| M. abscessus ATCC 19977 | MAB_3880 | - | 7e-93 | 54.79% (303) | lipase/esterase LipG |
| M. marinum M | MMAR_0981 | lipG1 | 1e-127 | 71.52% (316) | lipase/esterase LipG1 |
| M. avium 104 | MAV_4515 | - | 1e-126 | 73.36% (304) | hydrolase, alpha/beta fold family protein |
| M. smegmatis MC2 155 | MSMEG_1352 | - | 1e-106 | 59.34% (305) | hydrolase, alpha/beta fold family protein |
| M. thermoresistible (build 8) | TH_1861 | lipG | 1e-105 | 61.06% (303) | PROBABLE LIPASE/ESTERASE LIPG |
| M. ulcerans Agy99 | MUL_0733 | lipG1 | 1e-126 | 71.20% (316) | lipase/esterase LipG1 |
| M. vanbaalenii PYR-1 | Mvan_1240 | - | 1e-103 | 61.51% (304) | alpha/beta hydrolase fold |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_5112|M.gilvum_PYR-GCK -----MQVRTGTARS-------------GELEIHYEDMGDPNDPAVLLIM
Mvan_1240|M.vanbaalenii_PYR-1 -----MQIRSGTAKS-------------GDLDIFYEDMGDPNDPAVLLIM
MMAR_0981|M.marinum_M -----MEIRTGYAQSGVGSGATSERQPAGDLKLYYEDMGDINAPPVLLIM
MUL_0733|M.ulcerans_Agy99 -----MEIRTGYAQSGVGSGATSERQPAGDLKLYYEDMGDINAPPVLLIM
MLBr_01899|M.leprae_Br4923 -----MEVRSGYATS-------------GDLKLYYEDMGNVDDPPVLLIM
Mb0665c|M.bovis_AF2122/97 -----MDIRSGTAVS-------------GDVKLYYEDMGDLDHPPVLLIM
Rv0646c|M.tuberculosis_H37Rv -----MDIRSGTAVS-------------GDVKLYYEDMGDLDHPPVLLIM
MAV_4515|M.avium_104 ----MVQVRTGHAIS-------------GDLKLYYEDMGDIDDPPVLLIM
MSMEG_1352|M.smegmatis_MC2_155 -----MQTRTGLARAG---------DADHPIDLFYEDMGDPGDPAVLLIM
TH_1861|M.thermoresistible__bu -----MQIRTGFANVD---------GSPEPVDLYYEDLGDPADPPVLLIM
MAB_3880|M.abscessus_ATCC_1997 MSTTAREISEMTASTAT----------VGDIELCYEEFGNPADPAVLLIM
: * :.: **::*: *.*****
Mflv_5112|M.gilvum_PYR-GCK GLGAQLLLWRKGFCEKLINQGLRVIRFDNRDVGLSSKLT---GHHAGAPL
Mvan_1240|M.vanbaalenii_PYR-1 GLGAQLLLWRNGFCEKLINQGLRVIRFDNRDVGLSSKLH---HHRSGAPL
MMAR_0981|M.marinum_M GLGAQLLLWRTGFCQKLVDQGLRVIRYDNRDVGLSTKME---QPSARQPL
MUL_0733|M.ulcerans_Agy99 GLGAQLLLWRTGFCQKPVDQGLRVIRYDNRDVGLSTKME---QPSARQPL
MLBr_01899|M.leprae_Br4923 GLGAQLVLWRTAFCEKLVAQGLRVVRYDNRDVGLSSRTDSTEQPCPSQPL
Mb0665c|M.bovis_AF2122/97 GLGAQMLLWRTDFCARLVAKGLRVIRYDNRDVGLSTKTE---RHRPGQPL
Rv0646c|M.tuberculosis_H37Rv GLGAQMLLWRTDFCARLVAKGLRVIRYDNRDVGLSTKTE---RHRPGQPL
MAV_4515|M.avium_104 GLGAQLLLWRTAFCEKLVGRGLRVIRYDNRDVGLSSKTE---HRSSGQPL
MSMEG_1352|M.smegmatis_MC2_155 GLGAQMLMWRTEFCEKLVDQGLRVIRFDNRDVGLSTKLS---GQRVDSPL
TH_1861|M.thermoresistible__bu GLGAQLILWRDGFCRKLVERGLRVIRFDNRDVGLSSKLD---GRRERTPL
MAB_3880|M.abscessus_ATCC_1997 GIGAQMVFWRPEFCQQLADQGYRVIRFDNRDCGLSTKLDG--VRAGGGSL
*:***:::** ** : :* **:*:**** ***:: .*
Mflv_5112|M.gilvum_PYR-GCK LPRMARSLAGLPSP-AAYTLEDMADDAAALLDHLDIDRAHVVGGSMGGMI
Mvan_1240|M.vanbaalenii_PYR-1 LPRMARSFVGRPSP-AAYTLEDMADDAAALLDHLGIDTAHVVGGSMGGMI
MMAR_0981|M.marinum_M MPQLIRSWLGRRSQ-TPYTLEDMADDAAALLDHLDIERAHIVGASMGGMI
MUL_0733|M.ulcerans_Agy99 MPQLIRSWLGRRSQ-TPYTLEDMADDAAALLDHLDIERAHIVGASMGGMI
MLBr_01899|M.leprae_Br4923 TARLIRFWLGQRNA-CAYTLEDMTDDAVALLDHLSIERAHIVGASMGGMI
Mb0665c|M.bovis_AF2122/97 ATRLVRSWLGLPSQ-AAYTLEDMAADAAALLDHLDVKHAHVVGASMGGMI
Rv0646c|M.tuberculosis_H37Rv ATRLVRSWLGLPSQ-AAYTLEDMAADAAALLDHLDVKHAHVVGASMGGMI
MAV_4515|M.avium_104 VTRLLRSWLGLPSQ-SAYRLEDMADDAAAVLDHLGIDDAHIVGASMGGMI
MSMEG_1352|M.smegmatis_MC2_155 ALRMARSFLGKRSP-AVYTLEDMADDAAALLDHLEIDRAHIVGGSMGGMI
TH_1861|M.thermoresistible__bu PWRMLRSFAGLPSP-AVYTLEDMADDAAGLLDHLQIDAAHIVGASMGGMV
MAB_3880|M.abscessus_ATCC_1997 IPSMAKFLAGVKITGTAYTLVDMAGDAAGLLDHLGIEQAHIVGASMGGMI
: : * * * **: **..:**** :. **:**.*****:
Mflv_5112|M.gilvum_PYR-GCK AQVFAARHHPRTKTLGVIFSSNNQAALPPPGPRQLAAILQRP-KGSSRDE
Mvan_1240|M.vanbaalenii_PYR-1 AQVFAARHRARTGALGVIFSSNNQAALPPPGPRQLAAILQRP-KGSDREA
MMAR_0981|M.marinum_M AQIFAARFAERTNSLAVIFSSNNRPFLPPPAPRALLAVLNGPPPGSPREV
MUL_0733|M.ulcerans_Agy99 AQIFAARFAERTNSLAVIFSSNNRPFLPPPAPRALLAVLNGPPPGSPREV
MLBr_01899|M.leprae_Br4923 AQIFAARFPTRTRSLAVFFSSNNRPFLPPPAPRALLALLTGPPTGSRRDV
Mb0665c|M.bovis_AF2122/97 AQIFAARFAQRTKTLAVIFSSNNHRFLPPPAPRALLALLTGPPPDSPRDV
Rv0646c|M.tuberculosis_H37Rv AQIFAARFAQRTKTLAVIFSSNNHRFLPPPAPRALLALLTGPPPDSPRDV
MAV_4515|M.avium_104 AQIFAARFRQRTKTLAVIFSSNNSALLPPPAPRALLALLKGPPPDSPREV
MSMEG_1352|M.smegmatis_MC2_155 AQVFASQHAARTETLGVIFSSNNQALLPPPGIRQLLSLITGPPADASRDA
TH_1861|M.thermoresistible__bu AQVFAGRHAHRTRSLGVIFSSNNRPFLPPPGPRQLLSIITGPAPDAPRDE
MAB_3880|M.abscessus_ATCC_1997 AQVFAAEHPDRTATVSIIMSSNNQPFLPPPGPRQLMALLKPPPEGASREQ
**:**... ** ::.:::**** ****. * * ::: * .: *:
Mflv_5112|M.gilvum_PYR-GCK IVDNAVRVSRIIGSPGYPAPLDRLRTDAIEGFDRSYYPAGVGRHFAAILG
Mvan_1240|M.vanbaalenii_PYR-1 IIENAVRVSRIIGSPGYPAAEERLRADAIEGYDRSYYPAGVGRHFAAILG
MMAR_0981|M.marinum_M IIDNVVRVTRITGSPGYQPTEEWIRADAAENYDRSYYPWGIPRQFSAVLG
MUL_0733|M.ulcerans_Agy99 IIDNVVRVTRITGSPGYQPTEEWIRADAAENYDRSYYPWGIPRQFSAVLG
MLBr_01899|M.leprae_Br4923 VVDNVVRVTKITGSPLYRMPEEQVRTNAAEIYDRSFYPLGVSRQFSAILG
Mb0665c|M.bovis_AF2122/97 IVDNAVRVSKIIGSPAYPIPEDQVRAEAAESYDRNFHPWGIAQQFSAILG
Rv0646c|M.tuberculosis_H37Rv IVDNAVRVSKIIGSPAYPIPEDQVRAEAAESYDRNFHPWGIAQQFSAILG
MAV_4515|M.avium_104 IIDNAVRVGRIIGSTRYRVPEEQARAEAAEGYDRNYYPQGVARHFSAVLG
MSMEG_1352|M.smegmatis_MC2_155 VLDNAVRLSRLNGSPAYPRSEEEIRAEAAELYDRAYYPAGIARHFAAILG
TH_1861|M.thermoresistible__bu VIENAVKVGRLNGSPAYPVPDEQLRADAATAFDRSYHPVGVARQFDAILG
MAB_3880|M.abscessus_ATCC_1997 IIANSVQVGRIIGSPKYRQSQSKSYLHAAEYYDRSYYPKGFSRQFAAIMG
:: * *:: :: **. * . . .* :** ::* *. ::* *::*
Mflv_5112|M.gilvum_PYR-GCK SGSLRRYDRQITAPTVVIHGRADKLMRPSGGRAIARAIPGARLVLFDGMG
Mvan_1240|M.vanbaalenii_PYR-1 SGSLRRYDRQITAPTVVIHGTADKLMRPSGGRAIARAIPRARLVLFDGMG
MMAR_0981|M.marinum_M SGSLLGYNRRTVAPTVVIHGRADKLMRPFGGRAVAKAINGARMVEFDGMG
MUL_0733|M.ulcerans_Agy99 SGSLLGYNRRTVAPTVVIHGRADKLMRPFGGRAVAKAINGARMVEFDGMG
MLBr_01899|M.leprae_Br4923 SGSLLHYNQRIIAPTVVIHGRADKLVRPSSGRAVARAITGARLVLFDGMG
Mb0665c|M.bovis_AF2122/97 SGSLLRYDRRIVAPTVVIHGRADKLMRPFGGRAVARAINGARLVLIDGMG
Rv0646c|M.tuberculosis_H37Rv SGSLLRYDRRIVAPTVVIHGRADKLMRPFGGRAVARAINGARLVLIDGMG
MAV_4515|M.avium_104 SGSLRRYNRRTTAPTVVIHGRADKLMRPFGGRAVAGAIDGARLVLFDGMG
MSMEG_1352|M.smegmatis_MC2_155 SGSLRHHNERVTAPTVVIHGRADRLMPPSGGRAIANAIRGARMVLIDGMG
TH_1861|M.thermoresistible__bu SGSLRRYNRRTTSPTVVIHGRADKLMRPTGGRAVARAIPGARLTLFDGMG
MAB_3880|M.abscessus_ATCC_1997 TGSLAPFDNRITAPALVLHGKADKLMRPSGARAIARAIPGSRLALIDGMG
:*** .:.: :*::*:** **:*: * ..**:* ** :*:. :****
Mflv_5112|M.gilvum_PYR-GCK HDLPEPLWDDIVGELKTTFAEAP-------------
Mvan_1240|M.vanbaalenii_PYR-1 HELPEPLWDDIVGELKTTFAEIS-------------
MMAR_0981|M.marinum_M HDLPRQLWDQVIGVLTSNFEQAK-------------
MUL_0733|M.ulcerans_Agy99 HDLPRQLWDQVIGVLTSNFEQAE-------------
MLBr_01899|M.leprae_Br4923 HDLPQQLWDQAVGVLMSNFAKAS-------------
Mb0665c|M.bovis_AF2122/97 HDLPRQLWDRVIGELTRNFSEAG-------------
Rv0646c|M.tuberculosis_H37Rv HDLPRQLWDRVIGELTRNFSEAG-------------
MAV_4515|M.avium_104 HDLPQQLWDQVIGVLANNFAKAS-------------
MSMEG_1352|M.smegmatis_MC2_155 HELPEPVWDEVIGELKTNFAVVG-------------
TH_1861|M.thermoresistible__bu HDLPEPLWDEIVAELDKNITAGR-------------
MAB_3880|M.abscessus_ATCC_1997 HDLPEPLWNHIITKLTENFARSEPAAETPESEAAGR
*:**. :*: : * .: