For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MAAPDPSMTRIAGPWRHLDVHANGIRFHVVEAVPSGQPEGPDAATPPMQPALARPLVILLHGFGSFWWSW RHQLCGLTGARVVAVDLRGYGGSDKPPRGYDGWTLAGDTAGLIRALGHPSATLVGHADGGLACWTTALLH SRLVRAIALISSPHPAALRRSTLTRRDQRHALLPTLLRYQLPIWPERLLTRNNAAEIERLVRARGCAKWL ASEDFSQAIDHLRQAIQIPAAAHCALEYQRWAVRSQLRSEGRRFIRAMTQQLGMPLLHLRGDADPYVLAD PVERTQRYAPHGRYISIAGAGHFSHEEAPEEVNRHLMRFLEQVHQLS
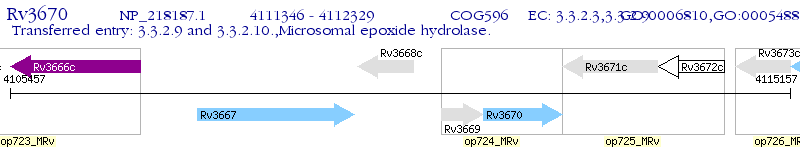
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. tuberculosis H37Rv | Rv3670 | ephE | - | 100% (327) | POSSIBLE EPOXIDE HYDROLASE EPHE (EPOXIDE HYDRATASE) (ARENE-OXIDE HYDRATASE) |
| M. tuberculosis H37Rv | Rv3617 | ephA | 8e-23 | 30.28% (327) | epoxide hydrolase EphA |
| M. tuberculosis H37Rv | Rv1938 | ephB | 3e-18 | 39.72% (141) | epoxide hydrolase EphB |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3694 | ephE | 0.0 | 100.00% (327) | epoxide hydrolase EphE |
| M. gilvum PYR-GCK | Mflv_1377 | - | 1e-145 | 74.14% (321) | alpha/beta hydrolase fold |
| M. leprae Br4923 | MLBr_02297 | - | 1e-154 | 80.25% (324) | putative hydrolase |
| M. abscessus ATCC 19977 | MAB_0422c | - | 1e-130 | 68.99% (316) | epoxide hydrolase EphE |
| M. marinum M | MMAR_5158 | ephE | 1e-159 | 82.30% (322) | epoxide hydrolase EphE |
| M. avium 104 | MAV_0458 | - | 1e-158 | 81.48% (324) | hydrolase, alpha/beta fold family protein |
| M. smegmatis MC2 155 | MSMEG_6184 | - | 1e-135 | 70.59% (323) | hydrolase, alpha/beta fold family protein |
| M. thermoresistible (build 8) | TH_0963 | ephE | 1e-134 | 70.90% (323) | POSSIBLE EPOXIDE HYDROLASE EPHE (EPOXIDE HYDRATASE) |
| M. ulcerans Agy99 | MUL_4244 | ephE | 1e-159 | 82.61% (322) | epoxide hydrolase EphE |
| M. vanbaalenii PYR-1 | Mvan_5429 | - | 1e-145 | 75.31% (324) | alpha/beta hydrolase fold |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_1377|M.gilvum_PYR-GCK ------MTPPDPSVVRIGGPWRHLDVHANGIRFHVVEAQRPSGADDVTR-
MSMEG_6184|M.smegmatis_MC2_155 ------MPPPDPSVTRIDGPWRHLQVHANGIRFHVVEAAPEAVEPGVT--
Mvan_5429|M.vanbaalenii_PYR-1 ------MVRPDPSVVRIGGPWRHLDVHANGIRFHVVEAERQAGADDRSK-
TH_0963|M.thermoresistible__bu ------MPPPDPSVTRIDGPWRHHEVHANGIRFHVVEAEPDEADRRIPR-
Rv3670|M.tuberculosis_H37Rv ------MAAPDPSMTRIAGPWRHLDVHANGIRFHVVEAVPSGQPEGPDAA
Mb3694|M.bovis_AF2122/97 ------MAAPDPSMTRIAGPWRHLDVHANGIRFHVVEAVPSGQPEGPDAA
MMAR_5158|M.marinum_M ------MPAPDPSITRIDGPWRHLDIHANGIRFHVVEAIPAGQRDN----
MUL_4244|M.ulcerans_Agy99 ------MPAPDPSITRIDGPWRHLDIHANGIRFHVVEAVPAGQRDN----
MLBr_02297|M.leprae_Br4923 ------MAMSNPSVTRIAGPWHHLDVHANGIRFHVVEAVPT-QPAG----
MAV_0458|M.avium_104 ------MAPPDPSVTRIDGPWRHLDVHANGIRFHVVEALPVGDARG----
MAB_0422c|M.abscessus_ATCC_199 MLFDQTARHPDPSVVRISGPWRHFDVHANGIRFHVVEADAQDAPTA----
.:**:.** ***:* ::************
Mflv_1377|M.gilvum_PYR-GCK -----PLTDRPLVILLHGFGSFWWSWRHQLTGLS---GARLVAVDLRGYG
MSMEG_6184|M.smegmatis_MC2_155 -------SDRPLVILLHGFGSFWWSWRHQLTGLR---DARVVAVDLRGYG
Mvan_5429|M.vanbaalenii_PYR-1 -----PLTDRPLVILLHGFGSFWWSWRHQLAGLT---GARVVAVDLRGYG
TH_0963|M.thermoresistible__bu -------TERPLVILLHGFGSFWYSWRHQLTGLTGLTGARVVAVDLRGYG
Rv3670|M.tuberculosis_H37Rv TPPMQPALARPLVILLHGFGSFWWSWRHQLCGLT---GARVVAVDLRGYG
Mb3694|M.bovis_AF2122/97 TPPMQPALARPLVILLHGFGSFWWSWRHQLCGLT---GARVVAVDLRGYG
MMAR_5158|M.marinum_M ----APSTTQPLVMLLHGFGSFWWSWRHQLRGLT---GARVVAVDLRGYG
MUL_4244|M.ulcerans_Agy99 ----APSTTQPLVMLLHGFGSFWWSWRHQLRGLT---GARVVAVDLRGYG
MLBr_02297|M.leprae_Br4923 QDCAPSATARPLVMLLHGFGSFWWSWRHQLRGLT---EARLVAVDLRGYG
MAV_0458|M.avium_104 ----LPATARPLVMLLHGFGSFWWSWRHQLRGLT---GARVVAVDLRGYG
MAB_0422c|M.abscessus_ATCC_199 -----PVTSRPLVLLLHGFGSFWWSWRHQLGALP---GARVVAVDLRGYG
:***:*********:****** .* **:*********
Mflv_1377|M.gilvum_PYR-GCK GSDKPPRGYDGWTLAGDTAGLVRALGHNSATLVGHADGGLVCWATSVLHP
MSMEG_6184|M.smegmatis_MC2_155 GSDKPPRGYDGWTLAGDTAGLVRALGHTSATLVGHADGGLVCWATSVLHP
Mvan_5429|M.vanbaalenii_PYR-1 GSDKPPRGYDGWTLAGDAAGLVRALGHNTATLVGHADGGLVCWATSVLHP
TH_0963|M.thermoresistible__bu GSDKPPRGYDGWTLAGDTAGLVRALGHPRAALVGHADGGLVCWATAVLHP
Rv3670|M.tuberculosis_H37Rv GSDKPPRGYDGWTLAGDTAGLIRALGHPSATLVGHADGGLACWTTALLHS
Mb3694|M.bovis_AF2122/97 GSDKPPRGYDGWTLAGDTAGLIRALGHPSATLVGHADGGLACWTTALLHS
MMAR_5158|M.marinum_M GSDKPPRGYDGWTLAGDTAGLIRALGHSSATLVGHADGGLACWTTALLHP
MUL_4244|M.ulcerans_Agy99 GSDKPPRGYDGWTLAGDTAGLIRALGHSSATLVGHADGGLACWTTALLHP
MLBr_02297|M.leprae_Br4923 GSDKPPRGYDGWTLAGDTAGLIRALGHSSATLVGHAEGGLACWATALLHP
MAV_0458|M.avium_104 GSDKPPRGYDGWTLSGDAAGLIRALGHSSATLVGHADGGLACWATALLHA
MAB_0422c|M.abscessus_ATCC_199 GSDKPPRGYDGWTLAGDTAGLIRALGHTSATLVGHAEGGLVCWATANLHP
**************:**:***:***** *:*****:***.**:*: **.
Mflv_1377|M.gilvum_PYR-GCK RVVKAIAVVSSPHPSALRASALRRRDQGRALLPSMMRYQVPMWPERALTR
MSMEG_6184|M.smegmatis_MC2_155 RAVNAIAVISSPHPVALRASALRRRDQGRALLPWMLGYQVPMWPERKLTR
Mvan_5429|M.vanbaalenii_PYR-1 RVVQAIAVVSSPHPAALRASALTRRDQGQALLPSLLRYQVPIRPERLLTR
TH_0963|M.thermoresistible__bu RVVRSIALVSSPHPAALRASALSNRAQARALLPSLLSYQLPRWPERMLTK
Rv3670|M.tuberculosis_H37Rv RLVRAIALISSPHPAALRRSTLTRRDQRHALLPTLLRYQLPIWPERLLTR
Mb3694|M.bovis_AF2122/97 RLVRAIALISSPHPAALRRSTLTRRDQRHALLPTLLRYQLPIWPERLLTR
MMAR_5158|M.marinum_M RLVRAIALISSPHPAALRRSTLTRRDQGRALLPTLLRYQVPLWPERLLTR
MUL_4244|M.ulcerans_Agy99 RLVRAIALISSPHPAALRRSTLTRRDQGRALLPTLLRYQVPLWPERLLTR
MLBr_02297|M.leprae_Br4923 RLVRAIALINSPHPAALRRSMLTHRNQGYALLPTLLRYQLPIWPERLLTR
MAV_0458|M.avium_104 RLVSAVALISSPHPAALRQSTLARRDQAGALLPTLLRYQVPLWPERLLTR
MAB_0422c|M.abscessus_ATCC_199 RLVNAIAVISSPHPIALRTSTFRGTGQGGALLPSLMRYQVPILPERTLTR
* * ::*::.**** *** * : * **** :: **:* *** **:
Mflv_1377|M.gilvum_PYR-GCK HNGAELERLVRSRASEKWQVSEDFAETVGHLRRAVQIPSAAHCALEYQRW
MSMEG_6184|M.smegmatis_MC2_155 DDAAELERVVRSRAGDAWQQTPDFAETMRHLRRAIQIPGAAHSALEYQRW
Mvan_5429|M.vanbaalenii_PYR-1 DDGAELERLVRSRSSAKWQAGEDFSESIRHMRRAIQIPSAAHCALEYQRW
TH_0963|M.thermoresistible__bu DDGARLERLVRSRAGAQWPATQDFATTVGHLRRAIQIPSAAHSALEYQRW
Rv3670|M.tuberculosis_H37Rv NNAAEIERLVRARGCAKWLASEDFSQAIDHLRQAIQIPAAAHCALEYQRW
Mb3694|M.bovis_AF2122/97 NNAAEIERLVRARGCAKWLASEDFSQAIDHLRQAIQIPAAAHCALEYQRW
MMAR_5158|M.marinum_M DNAEEIERLIRSRTSAKWAASEDFSQSIGYLRRAIQIPAAAHCALEYQRW
MUL_4244|M.ulcerans_Agy99 DNAEEIERLIRSRTSAKWAASEDFSQSIGYLRRAIQIPAAAHCALEYQRW
MLBr_02297|M.leprae_Br4923 NNAAEIERLVRSRSSAKWQACEDFAVTIDHLRIAIQIPAAAHCALEYQRW
MAV_0458|M.avium_104 DGGAEIERLVRSRSCAKWLASEDFSETIGHLRTAIQIPAAAHCALEYQRW
MAB_0422c|M.abscessus_ATCC_199 HRAAELERLVRSRAGTRWTSTDDFAQSMKHLRTAICIPGVAHCALEYQRW
. . .:**::*:* * **: :: ::* *: **..**.*******
Mflv_1377|M.gilvum_PYR-GCK AVRSQLRAEGRRFMRSMRAPLSVPMLHLRGDADPYVLPDPVYRTQRYAPH
MSMEG_6184|M.smegmatis_MC2_155 AVRSQLRGEGRRFMASMRRPVNVPVLHLRGDEDPYVLPDPVYRTQRYAPH
Mvan_5429|M.vanbaalenii_PYR-1 AVRSQLRAEGRRFMSSMKRPLGVPVLHLRGEQDPYVLADPVYRTQRYAPH
TH_0963|M.thermoresistible__bu AVRSQLRSEGHRFMRLMTRPVRVPVLHLHGESDPYVLADPVHRTQRHAPH
Rv3670|M.tuberculosis_H37Rv AVRSQLRSEGRRFIRAMTQQLGMPLLHLRGDADPYVLADPVERTQRYAPH
Mb3694|M.bovis_AF2122/97 AVRSQLRSEGRRFIRAMTQQLGMPLLHLRGDADPYVLADPVERTQRYAPH
MMAR_5158|M.marinum_M AVRSQLRGEGHRFMRSMNQRPGVPILHLRGDADPYVLAAPVERTQRYAPH
MUL_4244|M.ulcerans_Agy99 AVRSQLRREGHRFMRSMNQRPGVPILHLRGDADPYVLAAPVERTQRYAPH
MLBr_02297|M.leprae_Br4923 AVRSQLRNEGRKFMKSMSRPFSVPLLHVRGDADPYLLADSVDRTRRYAPH
MAV_0458|M.avium_104 AVRSQLRDEGRRFLRSMRRQLGVPLLHLRGEEDPYVLADPVDRTRRYAPQ
MAB_0422c|M.abscessus_ATCC_199 AVRSQLRSEGWRFMRSMNRELSVPVLHMRGESDPYVLADPVHRSLPYAPN
******* ** :*: * :*:**::*: ***:*. .* *: :**:
Mflv_1377|M.gilvum_PYR-GCK GRYVSIAGAGHFAHEEQPAAVNAQLSRFLSQVYT----
MSMEG_6184|M.smegmatis_MC2_155 GRYVSITGAGHFAHEEQPDAVNEQLGRFLAQIRS----
Mvan_5429|M.vanbaalenii_PYR-1 GRYVSIEGAGHFSHEEAPAAVNEQLRRFLNQVYG----
TH_0963|M.thermoresistible__bu GRYVSVPGAGHFAHEEAPAVVNEELGRFLTGRTGR---
Rv3670|M.tuberculosis_H37Rv GRYISIAGAGHFSHEEAPEEVNRHLMRFLEQVHQLS--
Mb3694|M.bovis_AF2122/97 GRYISIAGAGHFSHEEAPEEVNRHLMRFLEQVHQLS--
MMAR_5158|M.marinum_M GRYVSVAGVGHYSHEEAPEEINRHLMRFFEQTRDARLS
MUL_4244|M.ulcerans_Agy99 GRYVSVAGVGHYSHEEAPEEINRHLMRFFEQTRDARLS
MLBr_02297|M.leprae_Br4923 GRYVSVSDAGHFSHEEAPEEINRYLTRFLKQVHGPRLS
MAV_0458|M.avium_104 GRYVTVAGAGHFSHEEAPEEINRHLTSFLEDVHGPAVN
MAB_0422c|M.abscessus_ATCC_199 GQFVALRGVGHYVHEEDPAAVNEHLRRFTGHR------
*::::: ..**: *** * :* * *