For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VSCLPVSVLVVAKAPEPGRVKTRLAAAIGDKVAADIAAAALLDTLDAVAAAPVTARAVALTGDLDSAADS AEIRRRLKSFTVFRQRGDAFADRLANAHVDAADGYPVLQIGMDTPQVTAELLADCARLLLQIPAVLGLAF DGGWWVLGIRTPTAAECLRAVPMSQPDTGELTLKALRDNGIDVTLVQRLGDFDIVDDIALVRDCCAPGSR FAQATRAAGL
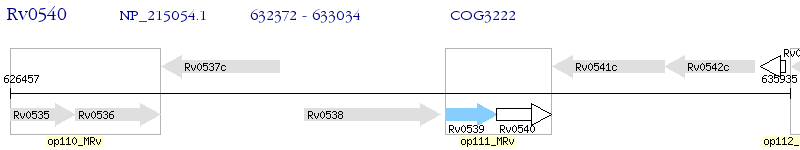
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. tuberculosis H37Rv | Rv0540 | - | - | 100% (220) | hypothetical protein Rv0540 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0554 | - | 1e-121 | 100.00% (220) | hypothetical protein Mb0554 |
| M. gilvum PYR-GCK | - | - | - | - | - |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | - | - | - | - | - |
| M. marinum M | MMAR_0886 | - | 3e-88 | 72.73% (220) | hypothetical protein MMAR_0886 |
| M. avium 104 | MAV_4604 | - | 2e-88 | 75.12% (217) | hypothetical protein MAV_4604 |
| M. smegmatis MC2 155 | MSMEG_0996 | - | 6e-73 | 67.61% (213) | hypothetical protein MSMEG_0996 |
| M. thermoresistible (build 8) | - | - | - | - | - |
| M. ulcerans Agy99 | MUL_0639 | - | 2e-88 | 72.73% (220) | hypothetical protein MUL_0639 |
| M. vanbaalenii PYR-1 | Mvan_0887 | - | 2e-67 | 62.56% (219) | hypothetical protein Mvan_0887 |
CLUSTAL 2.0.9 multiple sequence alignment
Rv0540|M.tuberculosis_H37Rv MSCLPVSVLVVAKAPEPGRVKTRLAAAIGDKVAADIAAAALLDTLDAVAA
Mb0554|M.bovis_AF2122/97 MSCLPVSVLVVAKAPEPGRVKTRLAAAIGDKVAADIAAAALLDTLDAVAA
MMAR_0886|M.marinum_M MSHVPVTLLVVAKAPEPGLAKTRLAATVGDQTAADIAAAALLDTLDAVAA
MUL_0639|M.ulcerans_Agy99 MSHVPVTLLVVAKAPEPGLAKTRLAATVGDQTAADIAAAALLDTLDAVAA
MAV_4604|M.avium_104 --MIPVTLLVVAKAPEPGRAKTRLAAAVGERAAAEIAAAALLDTLDAVAA
MSMEG_0996|M.smegmatis_MC2_155 --MLAVTVLVVAKAPVPGLAKTRLAAGVGADVAADIAAAALLDTLDAVAG
Mvan_0887|M.vanbaalenii_PYR-1 --MIPVGVLVVAKAPVPGLAKTRLAAALGADVAADIAAAALLDTLDAVAD
:.* :******* ** .****** :* .**:**************
Rv0540|M.tuberculosis_H37Rv APVTARAVALTGDLDSAADSAEIRRRLKSFTVFRQRGDAFADRLANAHVD
Mb0554|M.bovis_AF2122/97 APVTARAVALTGDLDSAADSAEIRRRLKSFTVFRQRGDAFADRLANAHVD
MMAR_0886|M.marinum_M TPAASRVVALTGDLANAARGAEIRQRLESFTVIGQRGDDFADRLANAHAD
MUL_0639|M.ulcerans_Agy99 TPAASRVVALTGDLANAARGAEIRRRLESFTVIGQRGDDFADRLANAHAD
MAV_4604|M.avium_104 APVAATVVALTGDLDGAARGAEIRARLASFTVIAQRGNDFGARLANAHAD
MSMEG_0996|M.smegmatis_MC2_155 APVAHRVVAMTGDLAEACRGREIADRLADFTVVAQRGDGFAERLVHAHSD
Mvan_0887|M.vanbaalenii_PYR-1 CPAAAKVVALTGNLDDARCADEIRSRLDNFIVIGQRGADLAERLVHAHRD
*.: .**:**:* * . ** ** .* *. *** :. **.:** *
Rv0540|M.tuberculosis_H37Rv AADG---YPVLQIGMDTPQVTAELLADCARLLLQIPAVLGLAFDGGWWVL
Mb0554|M.bovis_AF2122/97 AADG---YPVLQIGMDTPQVTAELLADCARLLLQIPAVLGLAFDGGWWVL
MMAR_0886|M.marinum_M AADG---YPVLQVGMDTPQISAALLAECARRLLETSAVLGFAVDGGWWVL
MUL_0639|M.ulcerans_Agy99 AADG---YPVLQVGMDTPQISAALLAECARRLLETSAVLGFAVDGGWWVL
MAV_4604|M.avium_104 AADG---LPVLQIGMDTPQVTAELLAGCARRLLDAPAVLGLARDGGWWVF
MSMEG_0996|M.smegmatis_MC2_155 AARP--GFPVLQIGMDTPQVSAGLLSDCGQALLGADAVLGMACDGGWWVL
Mvan_0887|M.vanbaalenii_PYR-1 AAAATGGLPIVQIGMDTPQVSPELLERCAQSLLGSDAVLGLAGDGGWWVL
** *::*:******::. ** *.: ** ****:* ******:
Rv0540|M.tuberculosis_H37Rv GIRTPTAAECLRAVPMSQPDTGELTLKALRDNGIDVTLVQRLGDFDIVDD
Mb0554|M.bovis_AF2122/97 GIRTPTAAECLRAVPMSQPDTGELTLKALRDNGIDVTLVQRLGDFDIVDD
MMAR_0886|M.marinum_M GVHTPAAAECLRGVPMSQPNTGELTLKALREGGIDVATVQPLADVDVVAD
MUL_0639|M.ulcerans_Agy99 GVHTPAAAECLRGVPMSQPNTGELTLKALREGGFDVATVQPLADVDVVAD
MAV_4604|M.avium_104 GVAAPALADCLRAVPMSAADTGDLTLKALRDNGIEVATVQTLADVDVVGD
MSMEG_0996|M.smegmatis_MC2_155 GVHDAAAAACLRTVPMSRPDTGELTLAALRDNGITVQLLAELADVDTIDD
Mvan_0887|M.vanbaalenii_PYR-1 GVRDAGFAECLRNVPMSTPDTGAVTLAALQDTDIHVELVCELVDVDTIDD
*: . * *** **** .:** :** **:: .: * : * *.* : *
Rv0540|M.tuberculosis_H37Rv IALVRDCCAPGSRFAQATRAAGL
Mb0554|M.bovis_AF2122/97 IALVRDCCAPGSRFAQATRAAGL
MMAR_0886|M.marinum_M VAVVRGACGPDSRFARVTRLAGI
MUL_0639|M.ulcerans_Agy99 VAVVRGACGPDSRFARVTRLAGI
MAV_4604|M.avium_104 VAVVRDACPPASRFARATRAAGL
MSMEG_0996|M.smegmatis_MC2_155 IDPVRRACRPGSRFACATEGV--
Mvan_0887|M.vanbaalenii_PYR-1 VGAVRAGCGPGSRFSRATAGVGP
: ** * * ***: .* .