For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
PSVGGLEGCPPWQGPNPREFRDDVVRVARNRGDGVTIEQIATDFGVHPMTLHKWMRQADIDEGAKPGKST GESAELRELRRRNRLLNRRTRCCAVPRPTCRRPICREKALPARK*
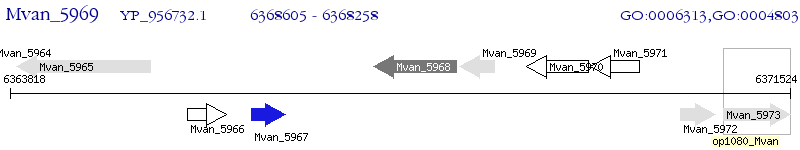
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_5969 | - | - | 100% (115) | transposase IS3/IS911 family protein |
| M. vanbaalenii PYR-1 | Mvan_5960 | - | 2e-36 | 94.59% (74) | transposase IS3/IS911 family protein |
| M. vanbaalenii PYR-1 | Mvan_1095 | - | 2e-36 | 94.59% (74) | transposase IS3/IS911 family protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb2839c | - | 5e-10 | 46.15% (78) | transposase |
| M. gilvum PYR-GCK | Mflv_0616 | - | 7e-36 | 91.89% (74) | transposase IS3/IS911 family protein |
| M. tuberculosis H37Rv | Rv3474 | - | 5e-10 | 46.15% (78) | transposase IS6110 |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_2087 | - | 8e-34 | 87.84% (74) | transposase |
| M. marinum M | MMAR_4592 | - | 9e-32 | 67.31% (104) | transposase |
| M. avium 104 | MAV_5035 | - | 2e-08 | 43.40% (53) | ISAfe7, transposase OrfA |
| M. smegmatis MC2 155 | MSMEG_6889 | - | 2e-34 | 90.54% (74) | transposase |
| M. thermoresistible (build 8) | TH_1992 | - | 2e-08 | 36.84% (76) | POSSIBLE TRANSPOSASE FOR INSERTION ELEMENT IS6110 |
| M. ulcerans Agy99 | - | - | - | - | - |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_0616|M.gilvum_PYR-GCK ---------------------------------------------MPRPY
MAB_2087|M.abscessus_ATCC_1997 ---------------------------MTSELACPKRAGWKDVS-VPRPY
MSMEG_6889|M.smegmatis_MC2_155 ---------------------------------------------MARPY
Mvan_5969|M.vanbaalenii_PYR-1 ---------------------------------MPSVGGLEGCPPWQGPN
MMAR_4592|M.marinum_M ---------------------------------------------MARPY
Mb2839c|M.bovis_AF2122/97 -----------------------------------------MSGGSSRRY
Rv3474|M.tuberculosis_H37Rv -----------------------------------------MSGGSSRRY
TH_1992|M.thermoresistible__bu VLAKLIWQQHWGTWLKFGRADKLFTRLRAARLDHTVEAEIRRLAAVDXKY
MAV_5035|M.avium_104 -----------------------------------MPKEQSPGKPTTRRY
Mflv_0616|M.gilvum_PYR-GCK PREFRDDVVRVARNRD------DGVTIEQIATDFGVH-PMTLHKWMRQAD
MAB_2087|M.abscessus_ATCC_1997 PREFRDDVVRVARNRD------DGVTIEQIATDFGVH-PMTLTKWMRQAD
MSMEG_6889|M.smegmatis_MC2_155 PREFRDDVVRVARNRD------DGVTIEQIATDFGVH-PMTLHKWMRQAD
Mvan_5969|M.vanbaalenii_PYR-1 PREFRDDVVRVARNRG------DGVTIEQIATDFGVH-PMTLHKWMRQAD
MMAR_4592|M.marinum_M PREFRDDVVRVARNRE------DGVTIEQIAADFGVH-PMTLRKWIRQAD
Mb2839c|M.bovis_AF2122/97 PPELRERAVRMVAEIR-GQHDSEWAAISEVARLLGVGCAETVRKWVRQAQ
Rv3474|M.tuberculosis_H37Rv PPELRERAVRMVAEIR-GQHDSEWAAISEVARLLGVGCAETVRKWVRQAQ
TH_1992|M.thermoresistible__bu SPEFRDRALRLLDTTMEDSGVSEFEAIKSVASKLGVS-QESVRRWRRKAE
MAV_5035|M.avium_104 SAEEKAAAVRMVRTLR-AELGTEQGTVQRVARQLGYG-VESVRTWVRQAD
. * : .:*: : ::. :* :* :: * *:*:
Mflv_0616|M.gilvum_PYR-GCK VDEGAKPGKSTGESVELRELRRRNRLLEQENEVLRRASAYLSQANLP---
MAB_2087|M.abscessus_ATCC_1997 VDEGAKPGKSTNDSADLRELRRRNRLLEQENEVLRRAAAYLSQANLP---
MSMEG_6889|M.smegmatis_MC2_155 IDEGAKPGKSTGESAELREARRRIKLLEQENEVLRRATAYLSQANLP---
Mvan_5969|M.vanbaalenii_PYR-1 IDEGAKPGKSTGESAELRELRRRNRLLNRRTRCCAVPRPTCRRPICR---
MMAR_4592|M.marinum_M IDDGAIPGKTTNEPAELRELRRRNRLLEQENEVLRRAAAYACRPVAARAR
Mb2839c|M.bovis_AF2122/97 VDAGARPGTTTEESAELKRLRRDNAELRRANAILKTASAFFAAELDR---
Rv3474|M.tuberculosis_H37Rv VDAGARPGTTTEESAELKRLRRDNAELRRANAILKTASAFFAAELDR---
TH_1992|M.thermoresistible__bu IDAGQRPGVTSTEHAEIRKLKRENAELRRANEILKSASAFFAAELDR---
MAV_5035|M.avium_104 IDEGLAPGVTTAESKRVQELEQENRELKRANEILKRAASFFGAELDR---
:* * ** :: : ::. .: *.: . . .
Mflv_0616|M.gilvum_PYR-GCK -GKGS-TRS-----------
MAB_2087|M.abscessus_ATCC_1997 -GKGS-TRS-----------
MSMEG_6889|M.smegmatis_MC2_155 -GKGS-TRS-----------
Mvan_5969|M.vanbaalenii_PYR-1 -EKALPARK-----------
MMAR_4592|M.marinum_M LRRARPSRGWARQLRRSRPA
Mb2839c|M.bovis_AF2122/97 -----PAR------------
Rv3474|M.tuberculosis_H37Rv -----PAR------------
TH_1992|M.thermoresistible__bu -----PATK-----------
MAV_5035|M.avium_104 -----QHKK-----------