For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
AKTFVGARIRQLRSERGFSQAALAQMLEISPSYLNQIEHDVRPLTVAVLLRITEVFGVDATFFASQDDTR LIAELREVLMDRDLDVDVDPAEVADMVGSHPALARAMVNLHRRYQLTTTQLAAATEDRFNVGSGGGSGAI TMPHEEVRDYFYQRQNYLHDLDTAAEALTADMRVHRGDLAGDLSDRLRFVHGVTVIRSVDLGEGVLHRYD PATRRLEVGAHLSSGQTVFRMAAELAYLECGELIDKLVDEGKFTSEESRKLARMGLANYFAAAAVLPYGQ FHEVAEKFRYDIERLSAYYSVSYETTCHRLSTLQRPSMRGVPFSFVRVDRAGNMSKRQSATGFHFSSSGG TCPLWNVYETFANPGKISVQIAEMPDGRAYMWVARTVERRASRYGQPGKTFAIGLGCELRHAQRLVYSEG LVTGDPNVPATPIGAGCRVCERDNCPQRAFPALGRALDIDEHRSTVSPYLVRQQ*
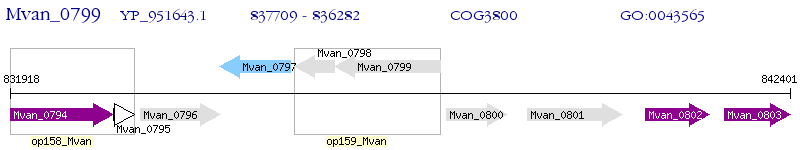
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_0799 | - | - | 100% (475) | hypothetical protein Mvan_0799 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0474c | - | 0.0 | 81.68% (475) | transcriptional regulatory protein |
| M. gilvum PYR-GCK | Mflv_0109 | - | 0.0 | 90.02% (481) | hypothetical protein Mflv_0109 |
| M. tuberculosis H37Rv | Rv0465c | - | 0.0 | 81.89% (475) | transcriptional regulatory protein |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_4118 | - | 0.0 | 81.68% (475) | transcriptional regulatory protein |
| M. marinum M | MMAR_0790 | - | 0.0 | 82.32% (475) | transcriptional regulatory protein |
| M. avium 104 | MAV_4684 | - | 0.0 | 83.16% (475) | DNA-binding protein |
| M. smegmatis MC2 155 | MSMEG_0906 | - | 0.0 | 82.95% (475) | DNA-binding protein |
| M. thermoresistible (build 8) | TH_4348 | - | 0.0 | 80.25% (476) | PUTATIVE protein of unknown function DUF955 |
| M. ulcerans Agy99 | MUL_4534 | - | 0.0 | 82.32% (475) | transcriptional regulatory protein |
CLUSTAL 2.0.9 multiple sequence alignment
MSMEG_0906|M.smegmatis_MC2_155 MPKTFVGSRVRQLRSERGFSQAALAQMLDISPSYLNQIEHDVRPLTVAVL
TH_4348|M.thermoresistible__bu VAKTYVGSRVRQLRNERGYSQAALAEMLGISPSYLNQIEHDVRPLTVPVL
Mb0474c|M.bovis_AF2122/97 MSKTYVGSRVRQLRNERGFSQAALAQMLEISPSYLDQIEHDVRPLTVAVL
Rv0465c|M.tuberculosis_H37Rv MSKTYVGSRVRQLRNERGFSQAALAQMLEISPSYLNQIEHDVRPLTVAVL
MMAR_0790|M.marinum_M MSKTFVGSRVRQLRNERGFSQAALAQLLEISPSYLNQIEHDVRPLTVAVL
MUL_4534|M.ulcerans_Agy99 MSKTFVGSRVRQLRNERGFSQAALAQLLEISPSYLNQIEHDVRPLTVAVL
MAV_4684|M.avium_104 MSKTFVGSRVRQLRHERGFSQAALAQMLEISPSYLNQIEHDVRPLTVAVL
MAB_4118|M.abscessus_ATCC_1997 MSKTYVGGRLRQLRSERGFSQAALAQMLEISPSYLNQIEHDVRPLTVAVL
Mvan_0799|M.vanbaalenii_PYR-1 MAKTFVGARIRQLRSERGFSQAALAQMLEISPSYLNQIEHDVRPLTVAVL
Mflv_0109|M.gilvum_PYR-GCK MAKTFVGARVRQLRSERGFSQAALAQMLEISPSYLNQIEHDVRPLTVAVL
:.**:**.*:**** ***:******::* ******:***********.**
MSMEG_0906|M.smegmatis_MC2_155 LRITEVFGVDATFFSPQDDTRLVAELREVTLDRDLGASDVDLTEVADMVA
TH_4348|M.thermoresistible__bu LRITDLFGVDATFFAPEDNTRLIAELREVTLDRDLDIN-VDMTEIADMVS
Mb0474c|M.bovis_AF2122/97 LRITEVFGVDATFFASQDDTRLVAELREVTLDRDLDIA-IDPHEVAEMVS
Rv0465c|M.tuberculosis_H37Rv LRITEVFGVDATFFASQDDTRLVAELREVTLDRDLDIA-IDPHEVAEMVS
MMAR_0790|M.marinum_M LRITEVFGVDATFFSSQDDTRLVAELREVTMDRDLDIS-VDPTEVAEMVS
MUL_4534|M.ulcerans_Agy99 LRITEVFGVDATFFSSQDDTRLVAELREVTMDRDLDIS-VDPTEVAEMVS
MAV_4684|M.avium_104 LRITEVFGVDATFFSSQDDTRLVAELREVTMDRDLDID-VDPTEVAEMVS
MAB_4118|M.abscessus_ATCC_1997 LRITEVFGVDATFFSSQDDSRLIAELREVVQDKDLDID-VDPAEIADVVA
Mvan_0799|M.vanbaalenii_PYR-1 LRITEVFGVDATFFASQDDTRLIAELREVLMDRDLDVD-VDPAEVADMVG
Mflv_0109|M.gilvum_PYR-GCK LRITEVFGVDATFFASQDDTRLIAELREVTMDRDLDVE-VDPAEIAEVVA
****::********:.:*::**:****** *:**. :* *:*::*.
MSMEG_0906|M.smegmatis_MC2_155 SHPKLARAMVNLHRRYRLATTQLAAATEDRYSD--------GSGSGAITM
TH_4348|M.thermoresistible__bu AHPELARAMVNLHQRYRRTTTQLAAATEDRYHDS----SGFGSGSGSITM
Mb0474c|M.bovis_AF2122/97 AHPGLARAVVNLHRRYRITTAQLAAATEERFSD--------GSGRGSITM
Rv0465c|M.tuberculosis_H37Rv AHPGLACAVVNLHRRYRITTAQLAAATEERFSD--------GSGRGSITM
MMAR_0790|M.marinum_M AHPSLARAMVNLHRRYRITTAQLAAATEERYSD--------GSGTGSITM
MUL_4534|M.ulcerans_Agy99 AHPSLARAMVNLHRRYRITTAQLAAATEERYSD--------GSGTGSITM
MAV_4684|M.avium_104 AHPGLARAVVNLHRRYRITTAQLAAATEERFFD--------GSGTGSITM
MAB_4118|M.abscessus_ATCC_1997 GHPALARAMVNLHRRYRITTTQLAAATEDRYTD--------GSGSGSITM
Mvan_0799|M.vanbaalenii_PYR-1 SHPALARAMVNLHRRYQLTTTQLAAATEDRFN------VGSGGGSGAITM
Mflv_0109|M.gilvum_PYR-GCK SHPSLARAMVNLHRRYQLTTTQLAAATEDRFNTAGLSSAGGGSGSGAISM
.** ** *:****:**: :*:*******:*: *.* *:*:*
MSMEG_0906|M.smegmatis_MC2_155 PHEEVRDYFYERQNYLHELDTAAEDLTARMRMHSGDLAAELTNRLTSVHG
TH_4348|M.thermoresistible__bu PHEEVRDYFYQRQNYLHELDTAAEDLTIRMGMTRTELARQLADRLTMVHG
Mb0474c|M.bovis_AF2122/97 PHEEVRDYFYQRQNYLHALDTAAEDLTAQMRMHHGDLARELTRRLTEVHG
Rv0465c|M.tuberculosis_H37Rv PHEEVRDYFYQRQNYLHALDTAAEDLTAQMRMHHGDLARELTRRLTEVHG
MMAR_0790|M.marinum_M PHEEVRDYFYQRQNYLHELDAAAEDLTIQMRLHHGDLRSELTRRLTEVHG
MUL_4534|M.ulcerans_Agy99 PHEEVRDYFYQRQNYLHELDAAAEDLTIQMRLHHGDLRSELTRRLTEVHG
MAV_4684|M.avium_104 PHEEVRDYFYQRQNYLHELDTAAEDLTIKMRLHHGDLARELTRRLTEVHG
MAB_4118|M.abscessus_ATCC_1997 PHEEVRDYFYQRHNYLHELDTAAEDLTVRMRMHRADLAREIADRLTEVHG
Mvan_0799|M.vanbaalenii_PYR-1 PHEEVRDYFYQRQNYLHDLDTAAEALTADMRVHRGDLAGDLSDRLRFVHG
Mflv_0109|M.gilvum_PYR-GCK PHEEVRDYFYQRQNYLHELDLAAENLTADMRVHRGELAGDLADRLRFVHG
**********:*:**** ** *** ** * : :* ::: ** ***
MSMEG_0906|M.smegmatis_MC2_155 VRVVRRIDMGETVLHRYDPATKTLEMSNHISRGQQVFKMAAELAYLEFGD
TH_4348|M.thermoresistible__bu VHIVRRVDMPETVLHRYDPETRTLELNNQLSSGQQVFKMAAELGYLEFGD
Mb0474c|M.bovis_AF2122/97 VRINKRIDLGDTVLHRYDPATNTLEISSHLSPGQQVFKMAAELAYLEFGD
Rv0465c|M.tuberculosis_H37Rv VRINKRIDLGDTVLHRYDPATNTLEISSHLSPGQQVFKMAAELAYLEFGD
MMAR_0790|M.marinum_M VRITRRIDLGDTVLHRFDPATKTLEISNHLASGQQVFKMAAELAYLEFGD
MUL_4534|M.ulcerans_Agy99 VRITRRIDLGDTVLHRLDPATMTLEISNHLASGQQVFKMAAELAYLEFGD
MAV_4684|M.avium_104 VHINRRIDLGDTVLHRYDPETKTLEISNHLSWGQQVFKMAAELAYLEFGD
MAB_4118|M.abscessus_ATCC_1997 VQIARRIDLGDSVLHKYDPASKTLEISNHLSGGQQVFKLAAELAYLEYGD
Mvan_0799|M.vanbaalenii_PYR-1 VTVIRSVDLGEGVLHRYDPATRRLEVGAHLSSGQTVFRMAAELAYLECGE
Mflv_0109|M.gilvum_PYR-GCK VTIIRRIDLGEVVMHRYDPDTRRLEMSAQLSAGQRVFRMAAELAYLECGD
* : : :*: : *:*: ** : **:. ::: ** **::****.*** *:
MSMEG_0906|M.smegmatis_MC2_155 LIDKLVDEGRFTSDESRTLARLGLANYFAAATVLPYRQFHDVAENFRYDI
TH_4348|M.thermoresistible__bu LIDKLVDEGHFTSDESRKLARLGLANYFAAATVLPYRQFHGVTENFRYDI
Mb0474c|M.bovis_AF2122/97 LIDAMVTDGKFTSAESRTLARLGLANYFAAATVLPYRQFHDVAENFRYDV
Rv0465c|M.tuberculosis_H37Rv LIDAMVTDGKFTSAESRTLARLGLANYFAAATVLPYRQFHDVAENFRYDV
MMAR_0790|M.marinum_M LIDAMVTDGKFTSDESRTLARLGLANYFAAAAVLPYRQFHDVAENFRYDV
MUL_4534|M.ulcerans_Agy99 LIDAMVTDGKFTSDESRTLARLGLANYFAAAAVLPYRQFHDVAENFRYDV
MAV_4684|M.avium_104 LIDSLVAQGKFTSEESHKLARLGLANYFAAAAVLPYRQFHDVAENFRYDV
MAB_4118|M.abscessus_ATCC_1997 LIDTMVDDGKFTSEESRKLARLGLANYFAAATVLPYRQFHGVAEDFQYDI
Mvan_0799|M.vanbaalenii_PYR-1 LIDKLVDEGKFTSEESRKLARMGLANYFAAAAVLPYGQFHEVAEKFRYDI
Mflv_0109|M.gilvum_PYR-GCK LIEKLVAEGHFTSEESRKLARMGLANYFAAATVLPYGQFHEVAEKFRYDV
**: :* :*:*** **:.***:*********:**** *** *:*.*:**:
MSMEG_0906|M.smegmatis_MC2_155 ERLSAFYQVSYETIGHRLSTLQRPSMRGVPFSFVRVDRAGNMSKRQSATG
TH_4348|M.thermoresistible__bu ERLSAFYRVSYETICHRLSTLQRPSMRGVPFSFVRVDRAGNMSKRQSASG
Mb0474c|M.bovis_AF2122/97 ERLSAFYSVSYETIAHRLSTLQRPSMRGVPFTFVQVDRAGNMSKRQSATG
Rv0465c|M.tuberculosis_H37Rv ERLSAFYSVSYETIAHRLSTLQRPSMRGVPFTFVRVDRAGNMSKRQSATG
MMAR_0790|M.marinum_M ERLSAFYSVSYETIAHRLSTLQRPSMRGVPFSFIRVDRAGNMSKRQSATG
MUL_4534|M.ulcerans_Agy99 ERLSAFYSVSYETIAHRLSTLQRPSMRGVPFSFIRVDRAGNMSKRQSATG
MAV_4684|M.avium_104 ERLSSFYSVSYETIAHRLSTLQRPSMRGVPFSFIRVDRAGNMSKRQSATG
MAB_4118|M.abscessus_ATCC_1997 ERLSAFYSVSYETVCHRLSTLQRPSMRGVPFSFVRVDRAGNMSKRQSATG
Mvan_0799|M.vanbaalenii_PYR-1 ERLSAYYSVSYETTCHRLSTLQRPSMRGVPFSFVRVDRAGNMSKRQSATG
Mflv_0109|M.gilvum_PYR-GCK ERLSAYYSVSYETICHRLSTLQRPSMRGVPFSFVRVDRAGNMSKRQSATG
****::* ***** ****************:*::*************:*
MSMEG_0906|M.smegmatis_MC2_155 FHFSSSGGTCPLWNVYETFAYPGKILVQIAQMPDGRHYMWVARTVERRAH
TH_4348|M.thermoresistible__bu FHFSSSGGTCPLWNVYETFAYPGKIMVQIAEMPDGRKYLWVARTVERRAS
Mb0474c|M.bovis_AF2122/97 FHFSSSGGTCPLWNVYETFANPGKILVQIAQMPDGRNYLWVARTVELRAA
Rv0465c|M.tuberculosis_H37Rv FHFSSSGGTCPLWNVYETFANPGKILVQIAQMPDGRNYLWVARTVELRAA
MMAR_0790|M.marinum_M FHFSSSGGTCPLWNVYETFANPGKILVQIAQMPDGRNYMWVARTVERRAA
MUL_4534|M.ulcerans_Agy99 FHFSSSGGTCPLWNVYETFANPGKILVQIAQMPDGRNYMWVARTVERRAA
MAV_4684|M.avium_104 FHFSSSGGTCPLWNVYETFANPGKILVQIAQMPDGRNYMWVARTVERRAA
MAB_4118|M.abscessus_ATCC_1997 FHFSSSGGTCPLWNVYETFGNPGKILVQVAQMPDGRNYLWVARTVERRAS
Mvan_0799|M.vanbaalenii_PYR-1 FHFSSSGGTCPLWNVYETFANPGKISVQIAEMPDGRAYMWVARTVERRAS
Mflv_0109|M.gilvum_PYR-GCK FHFSSSGGTCPLWNVYETFANPGKIGVQIAEMPDGRAYMWVARTVERRAA
*******************. **** **:*:***** *:******* **
MSMEG_0906|M.smegmatis_MC2_155 RYGQPGKTFAIGLGCELRHAHRLVYSEGLDLSGNV---STPIGAGCRVCE
TH_4348|M.thermoresistible__bu RYGQPGKTFAIGLGCELRHAHRLVYSEGLDLSGEN---ASPIGVGCRVCE
Mb0474c|M.bovis_AF2122/97 RYGQPGKTFAIGLGCELRHAHRLVYSEGLDLSGDPNTAATPIGAGCRVCE
Rv0465c|M.tuberculosis_H37Rv RYGQPGKTFAIGLGCELRHAHRLVYSEGLDLSGDPNTAATPIGAGCRVCE
MMAR_0790|M.marinum_M RYGQPGKTFAIGLGCELRHAHRLVYSEGLDLSGDPNIAATPIGAGCRVCE
MUL_4534|M.ulcerans_Agy99 RYGQPGKTFAIGLGCELRHAHRLVYSEGLDLSGDPNIAATPIGAGCRVCE
MAV_4684|M.avium_104 RYGQPGKTFAIGLGCELRHAHRLVYSEGLDVSGGPGTVATPIGAGCRVCE
MAB_4118|M.abscessus_ATCC_1997 RYGQPGKTFAIGLGCELRHAHRLVYSQGLDLSSEG--ATTPIGVGCRVCE
Mvan_0799|M.vanbaalenii_PYR-1 RYGQPGKTFAIGLGCELRHAQRLVYSEGL-VTGDPNVPATPIGAGCRVCE
Mflv_0109|M.gilvum_PYR-GCK RYGQPGKTFAIGLGCELRHAQRLVYSEGLDLSGDPSVRATPIGAGCRVCE
********************:*****:** ::. ::***.******
MSMEG_0906|M.smegmatis_MC2_155 RDNCPQRAFPALGKALDLDEHRSTVSPYLVRPSEVKPLG
TH_4348|M.thermoresistible__bu RDNCPQRAFPALGRALDLDEHRSTVSPYLVR----RP--
Mb0474c|M.bovis_AF2122/97 RDNCPQRAFPALGRALDLDEHRSTVSPYLVKQL------
Rv0465c|M.tuberculosis_H37Rv RDNCPQRAFPALGRALDLDEHRSTVSPYLVKQL------
MMAR_0790|M.marinum_M RDNCPQRAFPALGRALDLDEHRSTVSPYLVKQP------
MUL_4534|M.ulcerans_Agy99 RDNCPQRAFPALGRALDLDEHRSTVSPYLVKQP------
MAV_4684|M.avium_104 RDNCPQRAFPALGRALDLDEHRSTVSPYLVKQP------
MAB_4118|M.abscessus_ATCC_1997 RDNCPQRAFPALGRALDLDEHRSTVSPYLVLQEGAHP--
Mvan_0799|M.vanbaalenii_PYR-1 RDNCPQRAFPALGRALDIDEHRSTVSPYLVRQQ------
Mflv_0109|M.gilvum_PYR-GCK RDNCPQRAFPALGRALDIDEHRSTVSPYLVRQP------
*************:***:************