For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
TTAIQYPSASAKARFVRVSPTKARRVIDLVRGKSVEDALDILRWAPQAASEPVAKVIASAAANAQNNEGL DPSTLVVATVFADEGPTAKRIRPRAQGRAFRIRKRTSHITVIVESRPAKKQGASASAARARRAQASKAAS QKEGSE*
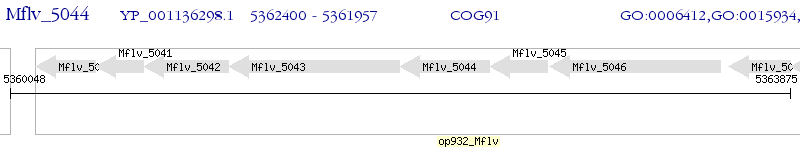
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. gilvum PYR-GCK | Mflv_5044 | rplV | - | 100% (147) | ribosomal protein L22 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0726 | rplV | 3e-60 | 83.69% (141) | 50S ribosomal protein L22 |
| M. tuberculosis H37Rv | Rv0706 | rplV | 3e-60 | 83.69% (141) | 50S ribosomal protein L22 |
| M. leprae Br4923 | MLBr_01858 | rplV | 2e-58 | 80.71% (140) | 50S ribosomal protein L22 |
| M. abscessus ATCC 19977 | MAB_3815c | rplV | 2e-62 | 81.08% (148) | 50S ribosomal protein L22 |
| M. marinum M | MMAR_1036 | rplV | 2e-61 | 83.69% (141) | 50S ribosomal protein L22, RplV |
| M. avium 104 | MAV_4466 | rplV | 1e-60 | 84.29% (140) | 50S ribosomal protein L22 |
| M. smegmatis MC2 155 | MSMEG_1441 | rplV | 4e-66 | 85.03% (147) | 50S ribosomal protein L22 |
| M. thermoresistible (build 8) | TH_0402 | rplV | 6e-62 | 84.40% (141) | PROBABLE 50S RIBOSOMAL PROTEIN L22 RPLV |
| M. ulcerans Agy99 | MUL_0795 | rplV | 2e-61 | 83.69% (141) | 50S ribosomal protein L22 |
| M. vanbaalenii PYR-1 | Mvan_1310 | rplV | 1e-66 | 85.81% (155) | ribosomal protein L22 |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_5044|M.gilvum_PYR-GCK --MTT-AIQYPSASAKARFVRVSPTKARRVIDLVRGKSVEDALDILRWAP
Mvan_1310|M.vanbaalenii_PYR-1 --MTT-AIEYPSAMAKARFVRISATKARRVIDLVRGKSVEEALDILRWAP
MSMEG_1441|M.smegmatis_MC2_155 --MST-VTEFPSATAKARYVRVSATKARRVIDLVRGKSVEEALDILRWAP
TH_0402|M.thermoresistible__bu --MSAPVIEYPSATAKARFVRTSATKARRVIDLVRGKTVAEALDILRWAP
MAB_3815c|M.abscessus_ATCC_199 --MTT-TTEFPSATAKARFVRVSPTKARRVIDLVRGKSVNDAIDILRWAP
Mb0726|M.bovis_AF2122/97 MTAATKATEYPSAVAKARFVRVSPRKARRVIDLVRGRSVSDALDILRWAP
Rv0706|M.tuberculosis_H37Rv MTAATKATEYPSAVAKARFVRVSPRKARRVIDLVRGRSVSDALDILRWAP
MMAR_1036|M.marinum_M ---MTTTTEFPSATAKARFVRVSPRKARRVIDLVRGRSVSDALDILRWAP
MUL_0795|M.ulcerans_Agy99 ---MTTTTEFPSATAKARFVRVSPRKARRVIDLVRGRSVSDALDILRWAP
MAV_4466|M.avium_104 ----MTTTEFPSAVAKARFVRVSPRKARRVIDLVRGRSVTDALDILRWAP
MLBr_01858|M.leprae_Br4923 ----MTTTEFPSAIAKARFVRVSPRKARRVIDLVRGRSVADALDILRWAP
. ::*** ****:** *. ***********::* :*:*******
Mflv_5044|M.gilvum_PYR-GCK QAASEPVAKVIASAAANAQNNEGLDPSTLVVATVFADEGPTAKRIRPRAQ
Mvan_1310|M.vanbaalenii_PYR-1 QSASEPVAKVIASAAANAQNNEGLDPTSLVVATIHADEGPTAKRIRPRAQ
MSMEG_1441|M.smegmatis_MC2_155 QAASEPVAKVIASAAANAQNNEGLDPSTLVVATVYADEGPTAKRIRPRAQ
TH_0402|M.thermoresistible__bu QAASEPVAKVIASAAANAQNNEGLDPDTLVVATIYADEGPTFKRIRPRAQ
MAB_3815c|M.abscessus_ATCC_199 QAASEPVAKVIASAAANAQNNDGLDPSTLVVSEIFADEGPTAKRIRPRAQ
Mb0726|M.bovis_AF2122/97 QAASGPVAKVIASAAANAQNNGGLDPATLVVATVYADQGPTAKRIRPRAQ
Rv0706|M.tuberculosis_H37Rv QAASGPVAKVIASAAANAQNNGGLDPATLVVATVYADQGPTAKRIRPRAQ
MMAR_1036|M.marinum_M QAASEPVAKVIASAAANAQNNNGLDPATLIVATVYADEGPTAKRIRPRAQ
MUL_0795|M.ulcerans_Agy99 QAASEPVAKVIASAAANAQNNNGLDPATLIVATVYADEGPTAKRIRPRAQ
MAV_4466|M.avium_104 QAASEPVAKVIASAAANAQNNNGLDPATLVVATVYADEGPTAKRIRPRAQ
MLBr_01858|M.leprae_Br4923 QAASEPVARVIASAAANAQNNNGLDPETLVVATVYAGEGPTAKRIRPRAQ
*:** ***:************ **** :*:*: :.*.:*** ********
Mflv_5044|M.gilvum_PYR-GCK GRAFRIRKRTSHITVIVESRPAK----KQGASASAARARRAQAS------
Mvan_1310|M.vanbaalenii_PYR-1 GRAYRIRKRTSHITVIVESRPPKKAG-KQGASASAARARRAQAS------
MSMEG_1441|M.smegmatis_MC2_155 GRAFRIRKRTSHITVIVESRPPKQ----KGASAASARSRRAQGS------
TH_0402|M.thermoresistible__bu GRAFRIRRRTSHITVVVESRPVKKG---RGGSAASARSRRAQAS------
MAB_3815c|M.abscessus_ATCC_199 GRAFRIRKRTSHITVVVESRPKQEKGGKAGASKASSRAARAQGS------
Mb0726|M.bovis_AF2122/97 GRAFRIRRRTSHITVVVESRPAKDQ-----RSAKSSRARRTEASKAASKV
Rv0706|M.tuberculosis_H37Rv GRAFRIRRRTSHITVVVESRPAKDQ-----RSAKSSRARRTEASKAASKV
MMAR_1036|M.marinum_M GRAFRIRKRTSHITVVVESRPSKDQ-----RSSKSSRARRAEGS------
MUL_0795|M.ulcerans_Agy99 GRAFRIRKRTSHITVVVESRPSKDQ-----RSSKSSRARRAEGS------
MAV_4466|M.avium_104 GRAFRIRKRTSHITVVVESRPAKDQ-----RSAKSSRTRRAEAS------
MLBr_01858|M.leprae_Br4923 GRAFRIRRRTSHITVVVESRPVKDQ-----RSAASTKARRAEAS------
***:***:*******:***** : * :::: *::.*
Mflv_5044|M.gilvum_PYR-GCK -----------KAASQ-------------------------------KEG
Mvan_1310|M.vanbaalenii_PYR-1 -----------KAATKKAT--------------------------DSKEG
MSMEG_1441|M.smegmatis_MC2_155 -----------KAAATKKSA-------------ETKE------------G
TH_0402|M.thermoresistible__bu -----------KAASTKKAP-------------ATKATAQKATADEAKGG
MAB_3815c|M.abscessus_ATCC_199 -----------KAAAAKKT--------------------------ESKGG
Mb0726|M.bovis_AF2122/97 GATAPAKKAAAKAPAKKAPASSGVKKTPAKKAPAKKAPAKASETSAAKGG
Rv0706|M.tuberculosis_H37Rv GATAPAKKAAAKAPAKKAPASSGVKKTPAKKAPAKKAPAKASETSAAKGG
MMAR_1036|M.marinum_M -------KAAATAPAKKSSASKAPAKKA-----ATKAESKTSETSEAKGG
MUL_0795|M.ulcerans_Agy99 -------KAAATAPAKKSSASKAPAKKA-----ATKAESKTSEISEAKGG
MAV_4466|M.avium_104 -------KAAAKAPAKKAPAKKAPAKKAPAKTAAKKTPAKTSETSDAKGG
MLBr_01858|M.leprae_Br4923 -------KTAGRAPAKKAGASSGATKMP-----PKKASVKTSEASETKGG
*.: *
Mflv_5044|M.gilvum_PYR-GCK SE
Mvan_1310|M.vanbaalenii_PYR-1 SE
MSMEG_1441|M.smegmatis_MC2_155 SE
TH_0402|M.thermoresistible__bu SQ
MAB_3815c|M.abscessus_ATCC_199 TS
Mb0726|M.bovis_AF2122/97 SD
Rv0706|M.tuberculosis_H37Rv SD
MMAR_1036|M.marinum_M SD
MUL_0795|M.ulcerans_Agy99 SD
MAV_4466|M.avium_104 SD
MLBr_01858|M.leprae_Br4923 SD
:.