For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
AFDVMVPLKAALDELTYDNDVRAVVLTGAGHGFSSGADHKSAGSVPHVDGLTRPAFGLRSMEVLDDVILA LRKLHQPVIAAVNGAAIGGGLCLALAADIRVASTSAYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEI MLTGRDVDAEEAERIGLVSRRVADEDLLETCYELAERIAGFSRPGTELTKRTLWSGLDAGSLEAHMQAEG LGQLYIRLLTDNFEEAVAARREKRAPRFSDDR*
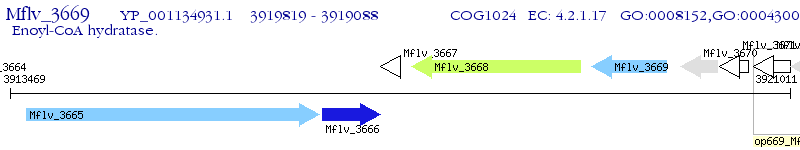
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. gilvum PYR-GCK | Mflv_3669 | - | - | 100% (243) | enoyl-CoA hydratase |
| M. gilvum PYR-GCK | Mflv_4817 | - | 6e-40 | 42.33% (215) | enoyl-CoA hydratase/isomerase |
| M. gilvum PYR-GCK | Mflv_2036 | - | 6e-26 | 33.90% (236) | enoyl-CoA hydratase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1507 | echA12 | 1e-113 | 84.30% (242) | enoyl-CoA hydratase |
| M. tuberculosis H37Rv | Rv1472 | echA12 | 1e-112 | 83.88% (242) | enoyl-CoA hydratase |
| M. leprae Br4923 | MLBr_01241 | - | 3e-92 | 70.37% (243) | enoyl-CoA hydratase |
| M. abscessus ATCC 19977 | MAB_2737c | - | 1e-106 | 76.95% (243) | enoyl-CoA hydratase |
| M. marinum M | MMAR_2278 | echA12 | 1e-109 | 81.48% (243) | enoyl-CoA hydratase, EchA12 |
| M. avium 104 | MAV_3307 | - | 1e-113 | 84.36% (243) | enoyl-CoA hydratase |
| M. smegmatis MC2 155 | MSMEG_3139 | - | 1e-112 | 82.30% (243) | enoyl-CoA hydratase |
| M. thermoresistible (build 8) | TH_2798 | - | 1e-124 | 91.36% (243) | PUTATIVE Enoyl-CoA hydratase |
| M. ulcerans Agy99 | MUL_1480 | echA12 | 1e-108 | 80.66% (243) | enoyl-CoA hydratase |
| M. vanbaalenii PYR-1 | Mvan_2741 | - | 1e-122 | 89.71% (243) | enoyl-CoA hydratase |
CLUSTAL 2.0.9 multiple sequence alignment
Mb1507|M.bovis_AF2122/97 --MPHRCAAQVVAGYRSTVSLVLVEHPRPEIAQITLNRPERMNSMAFDVM
Rv1472|M.tuberculosis_H37Rv --MPHRCAAQVVAGYRSTVSLVLVEHPRPEIAQITLNRPERMNSMAFDVM
MAV_3307|M.avium_104 ------------------MSLVLVDHPRPGVALITLNRPERMNSMAFDVM
MMAR_2278|M.marinum_M ------------------MALVLLEHPRPDIALVTLNRPERMNSMAFDVM
MUL_1480|M.ulcerans_Agy99 ------------------MSLVLLEHPRPDIALVTLNRPERMNSIAFDVM
Mflv_3669|M.gilvum_PYR-GCK --------------------------------------------MAFDVM
TH_2798|M.thermoresistible__bu ---------------------VLVDHPRPHIALVTLNRPERMNSMAFDVM
Mvan_2741|M.vanbaalenii_PYR-1 -----------MHAVTDGDGFVLVEHPRPHVALVTLNRPERMNSMAFDVM
MSMEG_3139|M.smegmatis_MC2_155 --------------MTDDTRFVLLDRSRPGVALVTLNRPERMNSMAFDVM
MAB_2737c|M.abscessus_ATCC_199 ------------------MSFVLVDRPRPEIALVTLNRPERMNAMAFDVM
MLBr_01241|M.leprae_Br4923 MSQTDASCTIAELPYRSVTDLVVLDFPRPEVALITLNRPGRMNSMALDLM
:*:*:*
Mb1507|M.bovis_AF2122/97 VPLKEALAQVSYDNSVRVVVLTGAGRGFSSGADHKSAGVVPHVENLTRPT
Rv1472|M.tuberculosis_H37Rv VPLKEALAQVSYDNSVRVVVLTGAGRGFSPGADHKSAGVVPHVENLTRPT
MAV_3307|M.avium_104 VPLKEALQKVTYDNAVRVVVLTGAGRGFSSGADHKSAGTVPHVEGLTRPT
MMAR_2278|M.marinum_M VPLKEVLEEVSYDNSVRVVVLTGAGRGFSSGADHKSAGAVPNVGGLTRPT
MUL_1480|M.ulcerans_Agy99 VPLKEVLEEVSYDNSVRVVVLTGAGRGFSSGADHKSAGAVPNVGGLTRPT
Mflv_3669|M.gilvum_PYR-GCK VPLKAALDELTYDNDVRAVVLTGAGHGFSSGADHKSAGSVPHVDGLTRPA
TH_2798|M.thermoresistible__bu VPLKAALDEITYDNDVRVVVLTGAGRGFSSGADHKSAGTVPHVDGLTRPS
Mvan_2741|M.vanbaalenii_PYR-1 VPLKTALDEITHDNDIRVVVLTGAGAGFSSGADHKSAGSVPHVAGLTRPT
MSMEG_3139|M.smegmatis_MC2_155 VPLLELLGDLRHDNSVRVVILTGAGRGFSSGADHKSAGSVPHVDGLTRPT
MAB_2737c|M.abscessus_ATCC_199 LPFKQMLVDISHDNDVRAVVITGAGKGFCSGADQKSAGPIPHIGGLTQPT
MLBr_01241|M.leprae_Br4923 KSLKQVLKRITYDHSVRVVVLTGAGRGFCSGADQKFTAPVPQVEGLTQPV
.: * : :*: :*.*::**** **..***:* :. :*:: .**:*
Mb1507|M.bovis_AF2122/97 YALRSMELLDDVILMLRRLHQPVIAAVNGPAIGGGLCLALAADIRVASSS
Rv1472|M.tuberculosis_H37Rv YALRSMELLDDVILMLRRLHQPVIAAVNGPAIGGGLCLALAADIRVASSS
MAV_3307|M.avium_104 YALRSMEILDEVILALRRLHQPVIAAVNGPAIGGGLCLALAADIRVASTS
MMAR_2278|M.marinum_M YALRSMELLDEVILMLRRLHQPVIAAVNGPAIGGGLCLALAADVRVAAAS
MUL_1480|M.ulcerans_Agy99 YALRSMELLDEVILMLRRLHQPVIAAVNGPAIGGGLCLALAADVRVAAAS
Mflv_3669|M.gilvum_PYR-GCK FGLRSMEVLDDVILALRKLHQPVIAAVNGAAIGGGLCLALAADIRVASTS
TH_2798|M.thermoresistible__bu YGLRSMEVLDDVILTLRKLHQPVIAAVNGAAIGGGLCLALAADIRIASTE
Mvan_2741|M.vanbaalenii_PYR-1 FGLRSMEVLDDVILALRKLHQPVIAAVNGAAIGGGLCLALAADIRVAGSD
MSMEG_3139|M.smegmatis_MC2_155 YALRSMEVLDNVILALRRLHQPVIAAVNGAAIGGGLCLALACDVRIAAHN
MAB_2737c|M.abscessus_ATCC_199 IALRSMELLDEVILTLRRMHQPVIAAINGAAIGGGLCLALACDVRVASQD
MLBr_01241|M.leprae_Br4923 RALRAMELLEEVILALRRLHQPVIAAINGPAIGGGLCLALAADVRVASTR
.**:**:*::*** **::*******:**.***********.*:*:*.
Mb1507|M.bovis_AF2122/97 AYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEIMLTGRDVSAEEAERI
Rv1472|M.tuberculosis_H37Rv AYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEIMLTGRDVSAEEAERI
MAV_3307|M.avium_104 AYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEIMLTGRDVTAEEAERI
MMAR_2278|M.marinum_M AYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEIMLTGRDVTAEEAERI
MUL_1480|M.ulcerans_Agy99 AYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEIMLTGRDVTAEEAERI
Mflv_3669|M.gilvum_PYR-GCK AYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEIMLTGRDVDAEEAERI
TH_2798|M.thermoresistible__bu AYFRAAGINNGLTASELGLSYLLPRAIGSSRAFEIMLTGRDVDAEEAERI
Mvan_2741|M.vanbaalenii_PYR-1 AYFRAAGINNGLTASELGLSYLLPRAIGASRAFEIMLTGRDVDADEAERI
MSMEG_3139|M.smegmatis_MC2_155 AYFRAAGINNGLTASELGLSYLLPRAIGTSRAFEIMLTGRDVDAAEAERI
MAB_2737c|M.abscessus_ATCC_199 AYFRAAGINNGLTASELGLSYLLPRAIGTSRASDIMLTGRDVDADEAERI
MLBr_01241|M.leprae_Br4923 AYFRAAGINNGLSASELGLSYLLPRAVGSSRAFEIMLSGRDVGAEEAEQI
************:*************:*:*** :***:**** * ***:*
Mb1507|M.bovis_AF2122/97 GLVSRQVPDEQLLDACYAIAARMAGFSRPGIELTKRTLWSGLDAASLEAH
Rv1472|M.tuberculosis_H37Rv GLVSRQVPDEQLLDACYAIAARMAGFSRPGIELTKRTLWSGLDAASLEAH
MAV_3307|M.avium_104 GLVSCQVPEEQLLDTCYAIAARIAAFSRPGIELTKRTLWSGLDAGSLEGH
MMAR_2278|M.marinum_M GLVSCRVPDGQLLDTCYAIAARMAAFSRPGIELTKRTLWSGLDAASLEGH
MUL_1480|M.ulcerans_Agy99 GLVSCRVPDGQLLDTCYAITARMAAFSRPGIELTKRTLWSGLDAASLEGH
Mflv_3669|M.gilvum_PYR-GCK GLVSRRVADEDLLETCYELAERIAGFSRPGTELTKRTLWSGLDAGSLEAH
TH_2798|M.thermoresistible__bu GLVSRRVPPAELLDTCYELADRIAGFSRPGTELTKRTLWSGLDAGSLESH
Mvan_2741|M.vanbaalenii_PYR-1 GLVSRRVPAEDLLPTCYDIAERIAGFSRPGTELTKRTLWTGLDAGSLEAH
MSMEG_3139|M.smegmatis_MC2_155 GLVSRTVPDDDLLETCFEMAQRMRGFSRPGIELTKRTLWSGLDAASLEGH
MAB_2737c|M.abscessus_ATCC_199 GLVSRKVASESLLEECYAIGERIAGFSRPGIELTKRTIWSGLDAASLESH
MLBr_01241|M.leprae_Br4923 GLVSYRVSDDRLLDTCYSIAARMATFSRSGTELTKRALWGGLDAASLDKH
**** *. ** *: : *: ***.* *****::* ****.**: *
Mb1507|M.bovis_AF2122/97 MQAEGLGQLFVRLLTANFEEAVAARAEQRAPVFTDDT-------
Rv1472|M.tuberculosis_H37Rv MQAEGLGQLFVRLLTANFEEAVAARAEQRAPVFTDDT-------
MAV_3307|M.avium_104 MQAEGLGQLFVRLLTANFEEAVAARAERRPPVFTDDK-------
MMAR_2278|M.marinum_M MQAEGLGQLFVRLLTANFEEAVAARAEKRPAVFTDDK-------
MUL_1480|M.ulcerans_Agy99 MQAEGLGQLFVRLLTANFEEAVAARAEKRPAVFTDDK-------
Mflv_3669|M.gilvum_PYR-GCK MQAEGLGQLYIRLLTDNFEEAVAARREKRAPRFSDDR-------
TH_2798|M.thermoresistible__bu MHAEGLGQLYIRLLTHNFEEAVAARKEKRAPRFPDDKI------
Mvan_2741|M.vanbaalenii_PYR-1 MQAEGLGQLYIRLLTSNFEEAVAARKEKRPPSFTDDK-------
MSMEG_3139|M.smegmatis_MC2_155 MQAEGLGQLFVRLLTANFEEAVAARAEKRPPAFTDDK-------
MAB_2737c|M.abscessus_ATCC_199 MHQEGLGQLYVRLLTDNFEEATAARKEKRPAEFRDKR-------
MLBr_01241|M.leprae_Br4923 MQSESLAQLFIALHTSNFEEAAAPCTEKRPTVLVDARGCATSPG
*: *.*.**:: * * *****.*. *:*.. : *