For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
LNSAPMVRRAVAGVLGAGAVSGAVLFGVLGAAPASAQPVPQNDPPNCTAADFAGVASGVSAATSAYLFTH PEVNTFFSNLHGAPRAQIDAEVAEYLASHPQVEAELKGVRQPLADIKHRCGFTPTDPDGPDF*
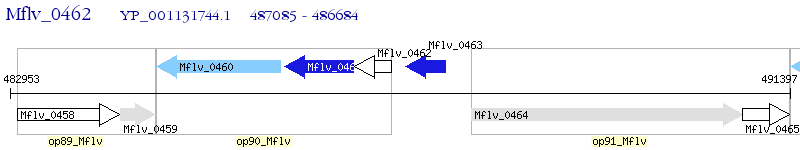
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. gilvum PYR-GCK | Mflv_0462 | - | - | 100% (133) | hypothetical protein Mflv_0462 |
| M. gilvum PYR-GCK | Mflv_4803 | - | 5e-30 | 56.90% (116) | hypothetical protein Mflv_4803 |
| M. gilvum PYR-GCK | Mflv_4804 | - | 7e-28 | 55.24% (105) | hypothetical protein Mflv_4804 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0209 | - | 6e-06 | 27.87% (122) | hypothetical protein Mb0209 |
| M. tuberculosis H37Rv | Rv0203 | - | 6e-06 | 27.87% (122) | hypothetical protein Rv0203 |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_1608c | - | 4e-25 | 51.82% (110) | hypothetical protein MAB_1608c |
| M. marinum M | MMAR_3300 | - | 2e-28 | 53.91% (115) | hypothetical protein MMAR_3300 |
| M. avium 104 | MAV_2237 | - | 4e-26 | 49.17% (120) | hypothetical protein MAV_2237 |
| M. smegmatis MC2 155 | MSMEG_0243 | - | 3e-36 | 62.28% (114) | hypothetical protein MSMEG_0243 |
| M. thermoresistible (build 8) | TH_3768 | - | 7e-37 | 59.85% (132) | PUTATIVE conserved hypothetical protein |
| M. ulcerans Agy99 | MUL_1329 | - | 2e-27 | 52.83% (106) | hypothetical protein MUL_1329 |
| M. vanbaalenii PYR-1 | Mvan_0190 | - | 1e-49 | 72.18% (133) | hypothetical protein Mvan_0190 |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_0462|M.gilvum_PYR-GCK ------------------------MLNSAPMVRRAV--------------
Mvan_0190|M.vanbaalenii_PYR-1 ------------------------MLFSATAVRRLV--------------
MSMEG_0243|M.smegmatis_MC2_155 ----------------------------------MV--------------
TH_3768|M.thermoresistible__bu ------------------------MVLSARTVRRAV--------------
MAB_1608c|M.abscessus_ATCC_199 --------------------------MTARVAHKLT--------------
MMAR_3300|M.marinum_M MTREPQGSVADATHHSVPAPARKHLSCAMRAVPAAFGYSMKESFMVLFGA
MUL_1329|M.ulcerans_Agy99 --------------------------------------------------
MAV_2237|M.avium_104 ------------------------MAPFARSARRVIGERIAA------GA
Mb0209|M.bovis_AF2122/97 ------------------------MKTGTATTRRRL--------------
Rv0203|M.tuberculosis_H37Rv ------------------------MKTGTATTRRRL--------------
Mflv_0462|M.gilvum_PYR-GCK --AGVLGAGAVSGAVLFGVLG-------AAPASAQPVPQNDPPNCTAADF
Mvan_0190|M.vanbaalenii_PYR-1 --VGALGAGAVSGGMLVGAL---------PSASAQPAPP--PPNCTAADF
MSMEG_0243|M.smegmatis_MC2_155 --AGAAGAGAVAGAMLFGAI---------PSALADPAPN--PPNCTAADL
TH_3768|M.thermoresistible__bu --AGTIGAGAIAGATLFSSI---------PFAFADPEPE--PPGCTAADL
MAB_1608c|M.abscessus_ATCC_199 --AGLC---AVA-ASLAGPL----------MAAADP-----PPNCTAADL
MMAR_3300|M.marinum_M ARSRLVAGGIVAGAMFLGAAGFASAVAGADPDDPAPVPAPAPPNCTAADL
MUL_1329|M.ulcerans_Agy99 --------------MFLGAAGFASAVAGADPDDPAPVPAPAPPNCTAADL
MAV_2237|M.avium_104 AGVAMTVAGLVA-TLVLTAG---PAAADWPMDDPEPAPASLP-NCTAADL
Mb0209|M.bovis_AF2122/97 -------LAVLIALALPGAA---------VALLAEPSATGASDPCAASEV
Rv0203|M.tuberculosis_H37Rv -------LAVLIALALPGAA---------VALLAEPSATGASDPCAASEV
. . * . *:*::.
Mflv_0462|M.gilvum_PYR-GCK AGVASGVSAATSAYLFTHPEVNTFFSNLHGAPRAQ-IDAEVAEYLASHPQ
Mvan_0190|M.vanbaalenii_PYR-1 SGVASGVSAASSAYLFTHPDVNTFLTGLHGMPHEQ-IRTEVANYLEMHPQ
MSMEG_0243|M.smegmatis_MC2_155 AGVASGVSAGTSAYLFTHPQVNDFFTSLEGLPREE-VKPKVADYLAANPQ
TH_3768|M.thermoresistible__bu AGVAAGVSAATSAYLFTHPEVNDFFTGLEGQHRAD-VRDTVAEYLDANPQ
MAB_1608c|M.abscessus_ATCC_199 AGVATGVSAATSSYLFTHPDVNDFFTGLKGLPKDQ-VRQRAQQYLEENPQ
MMAR_3300|M.marinum_M AGVSAGVAAATSVYLFSHPEVNDYFTGLRGQPRDD-IRDQLQQYMDANPG
MUL_1329|M.ulcerans_Agy99 AGVSAGVAAATSVYLFSHPEVNDYFTGLRGQPRDD-IRDQLQQYMDANPG
MAV_2237|M.avium_104 AQVSAGVAAATSAYLFSHPDVNAYFTSLKGQPRDE-MRAQLKQYMDANPQ
Mb0209|M.bovis_AF2122/97 ARTVGSVAKSMGDYLDSHPETNQVMTAVLQQQVGPGSVASLKAHFEANPK
Rv0203|M.tuberculosis_H37Rv ARTVGSVAKSMGDYLDSHPETNQVMTAVLQQQVGPGSVASLKAHFEANPK
: . .*: . . ** :**:.* :: : :: :*
Mflv_0462|M.gilvum_PYR-GCK VEAELKGVRQPLADIKHRCG-FTPTDPDGPDF--------
Mvan_0190|M.vanbaalenii_PYR-1 VNAELRGIRQPLADIKHRCG-FTPEDPDGPNFP-------
MSMEG_0243|M.smegmatis_MC2_155 VKDELTAVRQPLVDLKNRCG--TAPAPELP----------
TH_3768|M.thermoresistible__bu VKAELTHIRQPLVDIKNRCGGPDLATDDIP----------
MAB_1608c|M.abscessus_ATCC_199 VKADLHEIRRPLGDFRERCQ-------ER-----------
MMAR_3300|M.marinum_M VHAELQGIRQPLTDFRDRCQQ-------------------
MUL_1329|M.ulcerans_Agy99 VHAELQGIRQPLTDFRDRCQQ-------------------
MAV_2237|M.avium_104 THADLEGIRQPLTDFQDRCR--------------------
Mb0209|M.bovis_AF2122/97 VASDLHALSQPLTDLSTRCSLPISGLQAIGLMQAVQGARR
Rv0203|M.tuberculosis_H37Rv VASDLHALSQPLTDLSTRCSLPISGLQAIGLMQAVQGARR
. :* : :** *: **