For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MHGTDSSSDTPGQPAPSRAELLSAHAEGSISEDTILKTARDRAVDIGVGAVTPAVGALLSMLAKLSGGKA VAEVGTGAGVSGLWLLSGMSDDGVLTTIDIEPEHLRVAKQAFTEAGIGPSRTRLISGRAQEVLTRLADDS YDLVFVDADPIDQPDYVVEGVRLLRSGGVIVVHRAALGDRAGDPAVRDAEVTAVREAARLIAEDERLTPA LVPLGDGVLAAVRD
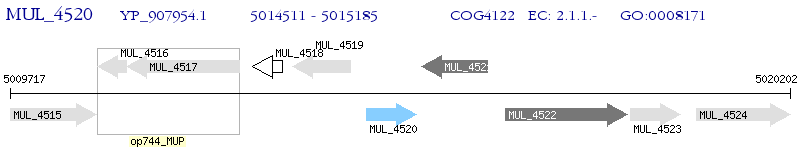
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. ulcerans Agy99 | MUL_4520 | - | - | 100% (224) | methyltransferase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1252c | - | 1e-104 | 89.72% (214) | methyltransferase |
| M. gilvum PYR-GCK | Mflv_2199 | - | 1e-89 | 73.89% (226) | O-methyltransferase family protein |
| M. tuberculosis H37Rv | Rv1220c | - | 1e-104 | 89.72% (214) | methyltransferase |
| M. leprae Br4923 | MLBr_01075 | - | 1e-100 | 82.59% (224) | putative O-methyltransferase |
| M. abscessus ATCC 19977 | MAB_1361c | - | 4e-80 | 69.23% (208) | O-methyltransferase |
| M. marinum M | MMAR_4217 | - | 1e-121 | 99.11% (224) | methyltransferase |
| M. avium 104 | MAV_1364 | - | 1e-105 | 86.61% (224) | O-methyltransferase, family protein 3 |
| M. smegmatis MC2 155 | MSMEG_5073 | - | 5e-93 | 81.25% (208) | O-methyltransferase, family protein 3 |
| M. thermoresistible (build 8) | TH_0546 | - | 2e-90 | 74.55% (224) | PROBABLE METHYLTRANSFERASE |
| M. vanbaalenii PYR-1 | Mvan_4497 | - | 3e-88 | 72.57% (226) | O-methyltransferase family protein |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_2199|M.gilvum_PYR-GCK MLVCQAYEQTELFLRNSATPFVTSGNRCDSRCTRGRVGDAGDSRWLYAAV
Mvan_4497|M.vanbaalenii_PYR-1 -------------MR----------GRFPRRGPGVSVSRQ-SLLWPYAAG
MSMEG_5073|M.smegmatis_MC2_155 --------------------------------------------------
TH_0546|M.thermoresistible__bu --------------------------------------------------
MAB_1361c|M.abscessus_ATCC_199 --------------------------------------------------
MUL_4520|M.ulcerans_Agy99 --------------------------------------------------
MMAR_4217|M.marinum_M --------------------------------------------------
Mb1252c|M.bovis_AF2122/97 --------------------------------------------------
Rv1220c|M.tuberculosis_H37Rv --------------------------------------------------
MLBr_01075|M.leprae_Br4923 --------------------------------------------------
MAV_1364|M.avium_104 --------------------------------------------------
Mflv_2199|M.gilvum_PYR-GCK MASTDEPAAASGDSTARARQAEAIVNHAEHSISEDAIVAAARERAVDIGA
Mvan_4497|M.vanbaalenii_PYR-1 MASTDEPGVGSGDPTARARQAEAIVNHAEHSISEDAIVAAARERAVDIGA
MSMEG_5073|M.smegmatis_MC2_155 MASS----------------AEAIVTHAERSISEDAIVAAARERADDIGA
TH_0546|M.thermoresistible__bu MTSTDDPGSRP-LSHPRSDTAQSLLAHAEQSISEDEIVAAARERAHDIGV
MAB_1361c|M.abscessus_ATCC_199 MTSSADRS-----------PVEAMVSHAEGAISEDDIVAAARERAVDLGA
MUL_4520|M.ulcerans_Agy99 MHGTDSSSDTP--GQPAPSRAELLSAHAEGSISEDTILKTARDRAVDIGV
MMAR_4217|M.marinum_M MHGTDSSSDTP--GQPAPSRAELLSAHAEGSISEDTILKTARDRAVDIGV
Mb1252c|M.bovis_AF2122/97 ---------MP--GQPAPSRGESLWAHAEGSISEDVILAGARERATDIGA
Rv1220c|M.tuberculosis_H37Rv ---------MP--GQPAPSRGESLWAHAEGSISEDVILAGARERATDIGA
MLBr_01075|M.leprae_Br4923 MYGTGNNAVTP--DQAAASRADSLFAHAEGSISEDAILASARERSEEIGA
MAV_1364|M.avium_104 MDGT--DAEAP--GQTAPSRAESLVAHAEASISEDALLAAARERAVDIGA
: : *** :**** :: **:*: ::*.
Mflv_2199|M.gilvum_PYR-GCK GAVTPAVGALLCVLAKLTGAKAVVEVGTGAGVSGLWLLSGMREDGVLTTI
Mvan_4497|M.vanbaalenii_PYR-1 GAVSPAVGALLCVLAKLTGARAVVEVGTGAGVSGLWLLSGMREDGVLTTI
MSMEG_5073|M.smegmatis_MC2_155 GAVTPAVGALLSVLARLTGGRAVVEVGTGAGVSGLWLLSGMRDDGVLTTI
TH_0546|M.thermoresistible__bu GAVTPAVGALLSVLAKVTGAKAVVEVGTGAGISGLWLLSGMRDDGVLTTI
MAB_1361c|M.abscessus_ATCC_199 GPVTPAVGALLSVFARLSGGKAVVEVGTGAGVSGLWLLSGMREDGVLTTI
MUL_4520|M.ulcerans_Agy99 GAVTPAVGALLSMLAKLSGGKAVAEVGTGAGVSGLWLLSGMSDDGVLTTI
MMAR_4217|M.marinum_M GAVTPAVGALLSMLAKLSGGKAVAEVGTGAGVSGLWLLSGMSDDGVLTTI
Mb1252c|M.bovis_AF2122/97 GAVTPAVGALLCLLAKLSGGKAVAEVGTGAGVSGLWLLSGMRDDGVLTTI
Rv1220c|M.tuberculosis_H37Rv GAVTPAVGALLCLLAKLSGGKAVAEVGTGAGVSGLWLLSGMRDDGVLTTI
MLBr_01075|M.leprae_Br4923 RAVTPAVGALLSLLTKLSGGKAVAEVGTGAGVSGLWLLSGMSYDGVLTTI
MAV_1364|M.avium_104 GAVTPAVGALLSLLTKLSGGKAIAEVGTGAGVSGLWLLSGMSDDGVLTTI
.*:*******.::::::*.:*:.*******:********* *******
Mflv_2199|M.gilvum_PYR-GCK DVEPEHQRIAKQAFSEAGVGPGRTRLISGRAQEVLTRLADESYDLVFIDA
Mvan_4497|M.vanbaalenii_PYR-1 DVEPEHQRIAKQAFSEAGVGPGRTRLISGRAQEVLTRLADESYDLVFIDA
MSMEG_5073|M.smegmatis_MC2_155 DVEPEHQRIAKQGFSEAGVGPGRTRLISGRAQEVLTRLADESYDLVFIDA
TH_0546|M.thermoresistible__bu DLEPEHQRIAKQAFTEAGVGPGRTRLISGRAQEVLTRLADESYDLVFIDA
MAB_1361c|M.abscessus_ATCC_199 DVEPEHQRSAKQAFSEASIAPSRTRLIGGRAQEVLPRLADESYDLLVIDA
MUL_4520|M.ulcerans_Agy99 DIEPEHLRVAKQAFTEAGIGPSRTRLISGRAQEVLTRLADDSYDLVFVDA
MMAR_4217|M.marinum_M DIEPEHLRVAKQAFTEAGIGPSRTRLISGRAQEVLTRLADDSYDLVFVDA
Mb1252c|M.bovis_AF2122/97 DIEPEHLRLARQAFAEAGIGPSRTRLISGRAQEVLTRLADASYDLVFIDA
Rv1220c|M.tuberculosis_H37Rv DIEPEHLRLARQAFAEAGIGPSRTRLISGRAQEVLTRLADASYDLVFIDA
MLBr_01075|M.leprae_Br4923 DIEPEYLRLAKQAFSEAGIGPSRTRLISGRGQDVLTRLADESYDLVFIDA
MAV_1364|M.avium_104 DIEPEYLRLAKQAFAEAGIGPSRTRLIGGRAQEVLTRLADESYDLVFIDA
*:***: * *:*.*:**.:.*.*****.**.*:**.**** ****:.:**
Mflv_2199|M.gilvum_PYR-GCK APADQPQFVVEGVRLLRPGGAIVVHRAALGGRAGDASAKDSEVSAVREAA
Mvan_4497|M.vanbaalenii_PYR-1 APADQPHFVTEGVRLLRPGGAIVVHRAALGGRAGDATAKDSEVAAVREAA
MSMEG_5073|M.smegmatis_MC2_155 DPVDQPQFVVEGVRLLRSGGAIVVHRAALGGRAGDADARDAEVTAVREAA
TH_0546|M.thermoresistible__bu DPIDLPQFVTEGVRLLRSGGAIVLHRAGLGGRVGDAGAADAEVSAVREAT
MAB_1361c|M.abscessus_ATCC_199 APADQPDFLAGGIRLLRPGGAVVIHGASAGGRAGDPSANDPDVVGVREAA
MUL_4520|M.ulcerans_Agy99 DPIDQPDYVVEGVRLLRSGGVIVVHRAALGDRAGDPAVRDAEVTAVREAA
MMAR_4217|M.marinum_M DPIDQPDYVVEGVRLLRSGGVIVVHRAALGGRAGDPAARDAEVTAVREAA
Mb1252c|M.bovis_AF2122/97 DPIDQPDYVAEGVRLLRSGGVIVVHRAALGGRAGDPGARDAEVIAVREAA
Rv1220c|M.tuberculosis_H37Rv DPIDQPDYVAEGVRLLRSGGVIVVHRAALGGRAGDPGARDAEVIAVREAA
MLBr_01075|M.leprae_Br4923 DPIDQPAYVVEGVRLLRSCGIIVVHRAALGGRAGDPAARDAEVTAVREAA
MAV_1364|M.avium_104 DPIDQPDYVVEGVRLLRPGGVIVVHRAALGGRAGDPAARDAEVVAVREAA
* * * ::. *:****. * :*:* *. *.*.**. . *.:* .****:
Mflv_2199|M.gilvum_PYR-GCK RLIAEDERLTPVLIPLGDGLLAAARD
Mvan_4497|M.vanbaalenii_PYR-1 RLIAEDDRLTPVLIPLGDGLLAAARD
MSMEG_5073|M.smegmatis_MC2_155 RLIAEDERLTPVLIPLGDGLLAAVRD
TH_0546|M.thermoresistible__bu RLIAEDERLTPVVIPLGDGLLAAARD
MAB_1361c|M.abscessus_ATCC_199 RIIADDGRFTAVLIPLGDGLLAATRD
MUL_4520|M.ulcerans_Agy99 RLIAEDERLTPALVPLGDGVLAAVRD
MMAR_4217|M.marinum_M RLIAEDERLTPALVPLGDGVLAAVRD
Mb1252c|M.bovis_AF2122/97 RLIAEDERLTPALVPLGDGVLAAVRD
Rv1220c|M.tuberculosis_H37Rv RLIAEDERLTPALVPLGDGVLAAVRD
MLBr_01075|M.leprae_Br4923 RLIAENERLTPALVPLGDGLLAAVRE
MAV_1364|M.avium_104 RLIAEDERLTPALVPLGDGILAAVRD
*:**:: *:*..::*****:***.*: