For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VGMIAGRTAGDMPGLDIAEERSWQNYLDSALRLYATLNRSLVDQHHLTLNDVRLLDYLDKSPTGSARMGD LADRLMSLPSRVTRQIRRLEVQNFVVRGASPEDGRGVVAKITDEGRAAVREAMVTYGQGVRTHFLGRLSR PQIAAMGENCRRISAALKNGAPPAKIGRV*
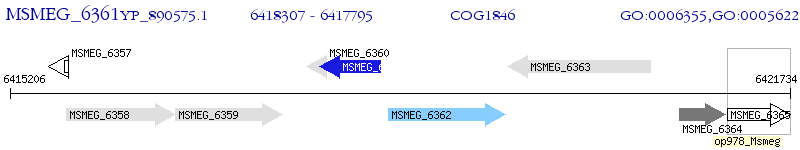
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_6361 | - | - | 100% (170) | transcriptional regulator, MarR family protein |
| M. smegmatis MC2 155 | MSMEG_5579 | - | 1e-30 | 46.26% (147) | MarR-family protein transcriptional regulator |
| M. smegmatis MC2 155 | MSMEG_5595 | - | 4e-13 | 32.65% (147) | MarR-family protein transcriptional regulator |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0758 | - | 2e-07 | 36.05% (86) | transcriptional regulatory protein |
| M. gilvum PYR-GCK | Mflv_1190 | - | 5e-70 | 76.47% (170) | MarR family transcriptional regulator |
| M. tuberculosis H37Rv | Rv0737 | - | 2e-07 | 36.05% (86) | transcriptional regulatory protein |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_0208 | - | 2e-34 | 47.62% (147) | MarR family transcriptional regulator |
| M. marinum M | MMAR_1075 | - | 2e-08 | 38.37% (86) | transcriptional regulator |
| M. avium 104 | MAV_4420 | - | 2e-08 | 33.94% (109) | transcriptional regulator, MarR family protein |
| M. thermoresistible (build 8) | TH_1664 | - | 9e-60 | 79.02% (143) | transcriptional regulator, MarR family protein |
| M. ulcerans Agy99 | MUL_0833 | - | 1e-08 | 38.37% (86) | transcriptional regulator |
| M. vanbaalenii PYR-1 | Mvan_5614 | - | 2e-69 | 75.88% (170) | MarR family transcriptional regulator |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_1075|M.marinum_M MAPD--NKRDPIAAARANWEAGGWGDVAQGMVAVTSVMRAHQILLARVEA
MUL_0833|M.ulcerans_Agy99 MAPD--NKRDPIAAARANWEAGGWGDVAQGMVAVTSVMRAHQILLARVEA
MAV_4420|M.avium_104 MADSRPHERDPIAAARANWERAGWGDVAPGMVAVTSVMRAHQILLARVET
Mb0758|M.bovis_AF2122/97 MASD---NRDPIAAARANWERSGWGDVSLGMVAVTSVMRAHQILLARVET
Rv0737|M.tuberculosis_H37Rv MASD---NRDPIAAARANWERSGWGDVSLGMVAVTSVMRAHQILLARVET
Mflv_1190|M.gilvum_PYR-GCK ------MPMEGIIGGRTASDTPGL-DIAEQRAWQNFLDAALRLYATMNRS
Mvan_5614|M.vanbaalenii_PYR-1 ------MPMEGIIGGRTASDTPGL-DIAEQRAWQNFLDAALRLYATMNRS
MSMEG_6361|M.smegmatis_MC2_155 --------MVGMIAGRTAGDMPGL-DIAEERSWQNYLDSALRLYATLNRS
TH_1664|M.thermoresistible__bu ------------------------------------LDAALRLYATLNKT
MAB_0208|M.abscessus_ATCC_1997 ------------MSTRSESEVPGL-DLAEFRSWQNFWVATNRLNFLLNRK
: :: .
MMAR_1075|M.marinum_M ALRPYDLSFSRYELLRLLAFSRTGALPITKASDRLQVHVTSVTHAIRRLE
MUL_0833|M.ulcerans_Agy99 ALRPYDLSFSRYELLRLLAFSRTGALPITKASDRLQVHVTSVTHAIRRLE
MAV_4420|M.avium_104 ALRPYDLSFSRYELLRLLAFSRTGALPITKASDRLQVHVTSVTHAIRRLE
Mb0758|M.bovis_AF2122/97 ALRPYDLSFSRFELLRLLAFSRIGALPITKASDRLQVHVTSVTHAIRRLE
Rv0737|M.tuberculosis_H37Rv ALRPYDLSFSRFELLRLLAFSRIGALPITKASDRLQVHVTSVTHAIRRLE
Mflv_1190|M.gilvum_PYR-GCK LSDEHGLTLNDVRLLDLLAKSPTGSARMGDVAEALMSLPSRVTRQIHRLE
Mvan_5614|M.vanbaalenii_PYR-1 LSDEHGLTLNDVRLLDLLAKSPTGSARMGDVAESLMSLPSRVTRQIHRLE
MSMEG_6361|M.smegmatis_MC2_155 LVDQHHLTLNDVRLLDYLDKSPTGSARMGDLADRLMSLPSRVTRQIRRLE
TH_1664|M.thermoresistible__bu LVDEHHLTLNDVRLLDMLDKSPSGSARMGDLADALMSLPSRVTRQIRRLE
MAB_0208|M.abscessus_ATCC_1997 MVASHSISLTDFHVLQQLLRSANGSCRMGDIANSLLASRSRITHQVRRLE
: :::. .:* * * *: : . :: * : :*: ::***
MMAR_1075|M.marinum_M ADGLVRRVPHPTDGRTTLVQITDLGRSTVEDATVTLNEHVFANIGMSTGE
MUL_0833|M.ulcerans_Agy99 ADGLVRRVPHPTDGRTTLVQITDLGRSTVEDATVTLNEHVFANIGMSTGE
MAV_4420|M.avium_104 ADGLVQRVPHPTDGRTTLVQITELGRSTVEDATVTLNKQVFADIGMSDTE
Mb0758|M.bovis_AF2122/97 ADGLVRRVPHPTDGRTTLVQITELGRSTVEDATVTLNEQVFANVGMGAEE
Rv0737|M.tuberculosis_H37Rv ADGLVRRVPHPTDGRTTLVQITELGRSTVEDATVTLNEQVFANVGMGAEE
Mflv_1190|M.gilvum_PYR-GCK VQGLVGRVASPDDGRGVLASITAEGRTALATAMQTYGSAVRTHFLDRLSR
Mvan_5614|M.vanbaalenii_PYR-1 VQGLVSRIASPDDGRGVLASITPEGRSALATAMQTYGVAVRSHFLDRLSR
MSMEG_6361|M.smegmatis_MC2_155 VQNFVVRGASPEDGRGVVAKITDEGRAAVREAMVTYGQGVRTHFLGRLSR
TH_1664|M.thermoresistible__bu AHNLVRRCASPDDGRGVLAIITDAGREEVRKAMVTYGNEVRVHFLSRLSR
MAB_0208|M.abscessus_ATCC_1997 EQGLVKRGSMPQDRRAVLAALTEQGRTLGEAAMRTYSECVREHYLMRLTR
..:* * . * * * .:. :* ** * * . * . .
MMAR_1075|M.marinum_M SEALVSSIETLRRSAGDF--------------
MUL_0833|M.ulcerans_Agy99 SEALVSSIETLRRSAGDF--------------
MAV_4420|M.avium_104 SQQLASSIETLRRNAGDF--------------
Mb0758|M.bovis_AF2122/97 SQALVSAVETLRRNAGDF--------------
Rv0737|M.tuberculosis_H37Rv SQALVSAVETLRRNAGDF--------------
Mflv_1190|M.gilvum_PYR-GCK PQ-VTAMGENCRRISAGLKAGGPP--AKIGRV
Mvan_5614|M.vanbaalenii_PYR-1 PQ-VTAMGENCRRISVALKNGGPS--AKIGRV
MSMEG_6361|M.smegmatis_MC2_155 PQ-IAAMGENCRRISAALKNGAPP--AKIGRV
TH_1664|M.thermoresistible__bu PQ-IAAMGENCRRISAGLRADGPP--ARLGRV
MAB_0208|M.abscessus_ATCC_1997 PQ-QAAIAESFRRVGEGAEEALHDGDGRGGE-
.: .: *. ** .