For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VGTRAPRRSTVALAAAATLASTGSAVIGARNLLNGQAAQARRTIPKAWAIPPRADGVYSPGGGPVERWER GVPFDLHLMIFGDSTATGYGCHSADEVPGVLLARGVAEQFGQRIRLSTKAIVGATSKGLSGQIDAMFVAG PPPDAAVIMIGANDITKPNGIGPSARRLGAGVSRLRSAGAAVVVGTCPDFGVITAIPQPLRWYARSRGLR LARAQAHAVRAAGGVPVPFSDLLTPEFHKTPELLFSEDMFHPSAAGYALAASQLLPALCNALGQWDGDDA LETGAADTSSLWSRLGAMSRLWRRSTGVPAPVVAAAS
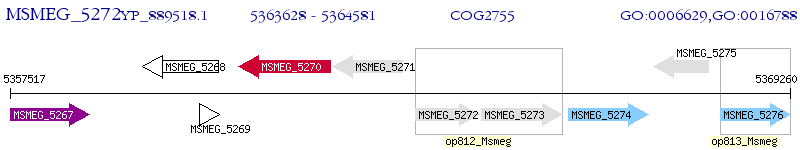
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_5272 | - | - | 100% (317) | hypothetical protein MSMEG_5272 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1104c | - | 1e-108 | 68.21% (302) | hypothetical protein Mb1104c |
| M. gilvum PYR-GCK | Mflv_2041 | - | 1e-143 | 81.21% (314) | GDSL family lipase |
| M. tuberculosis H37Rv | Rv1075c | - | 1e-108 | 68.21% (302) | hypothetical protein Rv1075c |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_1193c | - | 1e-106 | 63.99% (311) | hypothetical protein MAB_1193c |
| M. marinum M | MMAR_4392 | - | 1e-110 | 65.38% (312) | hypothetical protein MMAR_4392 |
| M. avium 104 | MAV_1199 | - | 1e-108 | 62.93% (321) | hypothetical protein MAV_1199 |
| M. thermoresistible (build 8) | TH_1369 | - | 9e-66 | 80.54% (149) | CONSERVED EXPORTED PROTEIN |
| M. ulcerans Agy99 | MUL_0206 | - | 4e-97 | 70.08% (244) | hypothetical protein MUL_0206 |
| M. vanbaalenii PYR-1 | Mvan_4676 | - | 1e-144 | 81.56% (320) | GDSL family lipase |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_2041|M.gilvum_PYR-GCK MGIPR-RRSTV--ALAAAATLASTGTAYVGARNLLTGQADQARQIIPKAW
Mvan_4676|M.vanbaalenii_PYR-1 MGIRA-SRRSV--ALAAAATLASTGTAYVGARNLLSGQADQARRVIPKSW
MSMEG_5272|M.smegmatis_MC2_155 MGTRAPRRSTV--ALAAAATLASTGSAVIGARNLLNGQAAQARRTIPKAW
MAB_1193c|M.abscessus_ATCC_199 MSILATRRRTIWAALSTGAAGAAMGTGVWGAYSFLNAQARQVRRVIPKTH
TH_1369|M.thermoresistible__bu --------------------------------------------------
Mb1104c|M.bovis_AF2122/97 ----MPRRSTI--ALATAGALASTGTAYLGARNLLVGQATHARTVIPKSF
Rv1075c|M.tuberculosis_H37Rv ----MPRRSTI--ALATAGALASTGTAYLGARNLLVGQATHARTVIPKSF
MAV_1199|M.avium_104 MTIRVPRRSAI--ALATAGALASTGTAYLGARSLLVGQATHARTVIPKSW
MMAR_4392|M.marinum_M ----MPRRSTI--ALATAGALASTGTAYLGARNLLVGQATHARTIIPKAW
MUL_0206|M.ulcerans_Agy99 ----MPRRSTI--ALATAGALASTGTAYLGARNLLVGQATHARTIIPKAW
Mflv_2041|M.gilvum_PYR-GCK DIPPRADGVYSPGGGPVERWHRGVPFDLHLMIFGDSTATGYGCTVADEVP
Mvan_4676|M.vanbaalenii_PYR-1 DIPPRADGVYAPGGGPVERWHRGVPFDLHLMIFGDSTATGYGCAAADEVP
MSMEG_5272|M.smegmatis_MC2_155 AIPPRADGVYSPGGGPVERWERGVPFDLHLMIFGDSTATGYGCHSADEVP
MAB_1193c|M.abscessus_ATCC_199 DAPPVADGVYERGGGPVTRYTRGVQYDLHLAIFGDSTATGYGCHIPDEVP
TH_1369|M.thermoresistible__bu --------VYSPDGGPVQPWNRDVPFDLHLMVFGDSTATGYGCREADEVP
Mb1104c|M.bovis_AF2122/97 DAPPRADGVYTRGGGPVQRWRREVPFDVHLMIFGDSTATGYGCASAEEVP
Rv1075c|M.tuberculosis_H37Rv DAPPRADGVYTRGGGPVQRWRREVPFDVHLMIFGDSTATGYGCASAEEVP
MAV_1199|M.avium_104 DIPPRADGVYTRGGGPVQRWHRGMAVDLHLMIFGDSTAAGYGCMCADEVP
MMAR_4392|M.marinum_M DVPPRADGVYSRGGGPVQRWRRGTPFDLHLMMFGDSTACGYGCGSPEEVP
MUL_0206|M.ulcerans_Agy99 DVPPRADGVYSRGGGPVQRWRRGTPFDLHLMMFGDSTACGYGCGSPEEVP
** .**** : * *:** :****** **** .:***
Mflv_2041|M.gilvum_PYR-GCK GVLIARGLAEESGKRIRLSTKAIVGATSKGLSGQIDAMYVAGPPPDAAVI
Mvan_4676|M.vanbaalenii_PYR-1 GVLIARGLAEESGLRIRLSTKAIVGATSKGLSGQIDAMFVAGAPPDAAVI
MSMEG_5272|M.smegmatis_MC2_155 GVLLARGVAEQFGQRIRLSTKAIVGATSKGLSGQIDAMFVAGPPPDAAVI
MAB_1193c|M.abscessus_ATCC_199 GVLLARGLAEESGKRIRLSTKAISGATSKGLAAQIDAMSIIGPPPDAAVI
TH_1369|M.thermoresistible__bu GVLLARALASASGKRVRLSTKAIVGATSKGLQGQIDAMFVADQPPDAAVI
Mb1104c|M.bovis_AF2122/97 GVLIARGLAEQTGKRIRLSTKAIVGATSKGVCGQVDAMFVVGPPPDAAVI
Rv1075c|M.tuberculosis_H37Rv GVLIARGLAEQTGKRIRLSTKAIVGATSKGVCGQVDAMFVVGPPPDAAVI
MAV_1199|M.avium_104 GVRIARGLAEQTGKRIRLSTKAIVGATSKGVCGQVDAMFVAGPPPDVAVI
MMAR_4392|M.marinum_M GVVMARKLAEQTGKRVRLSTKALVGATSKGVCGQVDAMFVAGPPPDAAII
MUL_0206|M.ulcerans_Agy99 GVVMARKLAEQTGKRVRLSTKALVGATSKGVCGQVDAMFVARPPPDATII
** :** :*. * *:******: ******: .*:*** : ***.::*
Mflv_2041|M.gilvum_PYR-GCK MIGANDITKPNGIGPSARRLGRAVRRLRGSDAVVVVGTCPDFGVITAIPQ
Mvan_4676|M.vanbaalenii_PYR-1 MIGANDITKPNGIGPSARRLGRAVRRLRGSDAVVVVGTCPDFGVITAIPQ
MSMEG_5272|M.smegmatis_MC2_155 MIGANDITKPNGIGPSARRLGAGVSRLRSAGAAVVVGTCPDFGVITAIPQ
MAB_1193c|M.abscessus_ATCC_199 MIGANDVTSLNAVRASAQRLGDAVRRLRAAGSVVVVGTCPDFGVITAIPQ
TH_1369|M.thermoresistible__bu MIGANDVTRLNGIRPSAQRLGAGVRRLREAGAVVVVGTCPDFGVVTAIPQ
Mb1104c|M.bovis_AF2122/97 MIGANDITALNGIGPSAQRLADCVRRLRTRGAVVVVGTCPDLGVITAIPQ
Rv1075c|M.tuberculosis_H37Rv MIGANDITALNGIGPSAQRLADCVRRLRTRGAVVVVGTCPDLGVITAIPQ
MAV_1199|M.avium_104 MVGANDVTALNGVSQSAHRLGLCVRKLRNRGAVVIVGTCPDLGFISAIPQ
MMAR_4392|M.marinum_M IIGANDITALNGIGPSAERLGSVVRKLRARGAIVVVGTCPDFSVITAIPQ
MUL_0206|M.ulcerans_Agy99 IIGANDMTALNGIGPSTERLGSVVRKLRARGAIVVVGTCPDFSVITAIPQ
::****:* *.: *:.**. * :** .: *:******:..::****
Mflv_2041|M.gilvum_PYR-GCK PLRWVARSRGLRLAHAQAAAVRAAGGVPVPFSDLLAPHFYKTPELLFSPD
Mvan_4676|M.vanbaalenii_PYR-1 PLRWVARSRGLRLARAQAAAVRAAGGVPVPFSDLLAPHFYKTPELLFSPD
MSMEG_5272|M.smegmatis_MC2_155 PLRWYARSRGLRLARAQAHAVRAAGGVPVPFSDLLTPEFHKTPELLFSED
MAB_1193c|M.abscessus_ATCC_199 PLRAVVRSRGLRLAHFQAAAVRATGGVPVPLADLLAPEFLQAPEEMFADD
TH_1369|M.thermoresistible__bu PLRSVARXX-----------------------------------------
Mb1104c|M.bovis_AF2122/97 PLRALAHTRGVRLARAQTAAVKAAGGVPVPLGHLLAPKFRAMPELMFSAD
Rv1075c|M.tuberculosis_H37Rv PLRALAHTRGVRLARAQTAAVKAAGGVPVPLGHLLAPKFRAMPELMFSAD
MAV_1199|M.avium_104 PLRSLAHERCLQLARAQTATVRAAGGVPVPLAQLMAPQFRATPEAMFSAD
MMAR_4392|M.marinum_M PLRSLAHTRGSQLARAQTAAVKAAGGVPVPLANLLARQFRAEPERMFSAD
MUL_0206|M.ulcerans_Agy99 PLRSLANTRGSQLARAQTAAVKAAGGVPVPLANLLARQFRAEPERMFSAD
*** ..
Mflv_2041|M.gilvum_PYR-GCK MFHPSAAGYELAATQLLPALCEELGEFITGEPLAEALESRTGEGTSLLSR
Mvan_4676|M.vanbaalenii_PYR-1 MFHPSAAGYELAAKQLLPALGNALGAFVTEDSAEAALESRTGEDGSLLGR
MSMEG_5272|M.smegmatis_MC2_155 MFHPSAAGYALAASQLLPALCNALGQWDGDD----ALETGAADTSSLWSR
MAB_1193c|M.abscessus_ATCC_199 HYHPSAAGYALAARLLLPAVAHALGEWTGGPIPDLPWVSAAAESQTMSAR
TH_1369|M.thermoresistible__bu ------------------------XAWTR---------------------
Mb1104c|M.bovis_AF2122/97 RYHPSAPAYALAADLLFLALRDALTEKLDIPIHETPSRPGTATLEPGHTR
Rv1075c|M.tuberculosis_H37Rv RYHPSAPAYALAADLLFLALRDALTEKLDIPIHETPSRPGTATLEPGHTR
MAV_1199|M.avium_104 GYHPSAPAYALAADALLLALCEALGEQVERPPLNQPVPSAQPVLGQRHTR
MMAR_4392|M.marinum_M RYHPSATAYALAADQLLLSLRDALREKIGSAMLPVPSRPSFRTSSQERSR
MUL_0206|M.ulcerans_Agy99 -LDITHPQRRTPWRPISRSSVCAMRYAKRSGAQCFPCRRGLASAPAARSV
Mflv_2041|M.gilvum_PYR-GCK VSGLTRLWRR-STGVPAPIVVAAG-----------
Mvan_4676|M.vanbaalenii_PYR-1 LGGLSRLWRR-STGVPAPIVVAAG-----------
MSMEG_5272|M.smegmatis_MC2_155 LGAMSRLWRR-STGVPAPVVAAAS-----------
MAB_1193c|M.abscessus_ATCC_199 LRVLSRIWRRNATEIATSITEPADRPDNLPVASLH
TH_1369|M.thermoresistible__bu -----------------------------------
Mb1104c|M.bovis_AF2122/97 HSMMSRLRRPRPARAVPTGG---------------
Rv1075c|M.tuberculosis_H37Rv HSMMSRLRRPRPARAVPTGG---------------
MAV_1199|M.avium_104 SSVMSRLWRRPAPGAAPSSCPEVGSD---------
MMAR_4392|M.marinum_M GAIVTRLWRRPVPVVGGPVAIPASG----------
MUL_0206|M.ulcerans_Agy99 PGAPS------SPGCGGARSRWLVDPS--------