For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
MGTDDRLPDRPGADDPGRGRRIGIDVGTVRIGVASSDPDGILATPVETVARDKRDKTGKHVRRLAALVKE YEAVEVIVGLPRTLGDRASASAHDAVDVAEQLARRIAPTPVRLADERLTTVSAQRSLREAGVRARGQKAV IDQVAAVGILQGWLDQRRAALAARREGTDG
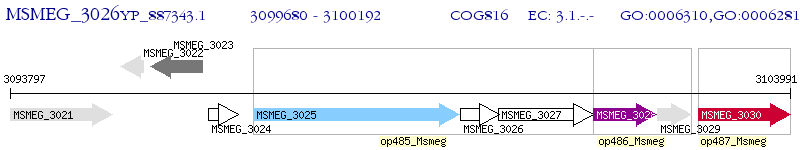
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_3026 | - | - | 100% (170) | Holliday junction resolvase-like protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb2584c | - | 2e-62 | 73.49% (166) | Holliday junction resolvase-like protein |
| M. gilvum PYR-GCK | Mflv_3762 | - | 2e-64 | 73.81% (168) | Holliday junction resolvase YqgF |
| M. tuberculosis H37Rv | Rv2554c | - | 2e-62 | 73.49% (166) | Holliday junction resolvase-like protein |
| M. leprae Br4923 | MLBr_00513 | - | 7e-54 | 69.18% (159) | Holliday junction resolvase-like protein |
| M. abscessus ATCC 19977 | MAB_2850c | - | 6e-53 | 66.23% (151) | putative Holliday junction resolvase |
| M. marinum M | MMAR_2171 | - | 2e-60 | 71.26% (167) | hypothetical protein MMAR_2171 |
| M. avium 104 | MAV_3430 | - | 3e-63 | 74.12% (170) | Holliday junction resolvase-like protein |
| M. thermoresistible (build 8) | TH_0113 | - | 2e-60 | 69.23% (169) | CONSERVED HYPOTHETICAL PROTEIN |
| M. ulcerans Agy99 | MUL_1755 | - | 1e-60 | 71.26% (167) | Holliday junction resolvase-like protein |
| M. vanbaalenii PYR-1 | Mvan_2641 | - | 2e-64 | 76.19% (168) | Holliday junction resolvase YqgF |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_3762|M.gilvum_PYR-GCK MPDTDWTSRVPDRPGDPNA--APDP--GRGRRLGIDVGSVRIGVAVSDPD
Mvan_2641|M.vanbaalenii_PYR-1 MPDTDWTSRVPDRPGDPNT--APDP--GRGRRLGVDVGTVRIGVAVSDPD
MSMEG_3026|M.smegmatis_MC2_155 MGTDD---RLPDRPG------ADDP--GRGRRIGIDVGTVRIGVASSDPD
Mb2584c|M.bovis_AF2122/97 --MVPAQHRPPDRPGDPA----HDP--GRGRRLGIDVGAARIGVACSDPD
Rv2554c|M.tuberculosis_H37Rv --MVPAQHRPPDRPGDPA----HDP--GRGRRLGIDVGAARIGVACSDPD
MAV_3430|M.avium_104 --MT--QHRVPDRPGDPD----QDP--GRGRRLGIDVGSVRIGVACSDPD
MMAR_2171|M.marinum_M --MVAASHRSPDRPGDPEG---LEPGTGRGRRLGIDVGTVRIGVASSDPD
MUL_1755|M.ulcerans_Agy99 --MVAASHRSPDRPGDPEG---LEPGTGRGRRLGIDVGTVRIGVASSDPD
MLBr_00513|M.leprae_Br4923 --MFSSQHRLLYQPSGPDLSKNLDP--ERGRRLGIDVGSVRIGVAFSDPD
TH_0113|M.thermoresistible__bu --VDDSRERRPDRPGDALQP--PDP--GPGRRLGVDVGKVRIGVAASDPD
MAB_2850c|M.abscessus_ATCC_199 --MPDTAAPTPDRPGP------DDP--GRGRRLGVDVGTVRIGISSSDPD
:*. :* ***:*:*** .***:: ****
Mflv_3762|M.gilvum_PYR-GCK AVLATPVETVTRDRR--SGRHLRRLTTLVGEYEAVEVVIGLPRTLGNRAG
Mvan_2641|M.vanbaalenii_PYR-1 AILATPVETVPRDKR--GTKHVRRLAALVDEYQVVEVVVGLPRTLADRAG
MSMEG_3026|M.smegmatis_MC2_155 GILATPVETVARDKRDKTGKHVRRLAALVKEYEAVEVIVGLPRTLGDRAS
Mb2584c|M.bovis_AF2122/97 AILATPVETVRRDRS---GKHLRRLAALAAELEAVEVIVGLPRTLADRIG
Rv2554c|M.tuberculosis_H37Rv AILATPVETVRRDRS---GKHLRRLAALAAELEAVEVIVGLPRTLADRIG
MAV_3430|M.avium_104 AVLATPVETVRRDRS---GKHLRRLAALVTELGAVEVVVGLPRTLADRTG
MMAR_2171|M.marinum_M GILATPLETVRRERS---GKHLRRLATLVGETEAVEVIVGLPRTLADRIG
MUL_1755|M.ulcerans_Agy99 GILATPLETVRRERS---GKHLRRLATLVGETEAVEVIVGLPRTLADRIG
MLBr_00513|M.leprae_Br4923 GILATPVETVRRYRS---AKHLRRLAELVVELQVVEVVVGLPWTLTDRTG
TH_0113|M.thermoresistible__bu GVLATPLETVRRERS---GRHLRRLAALVAELSAVEVVVGLPRTLADRMG
MAB_2850c|M.abscessus_ATCC_199 GILATPVETVSRDAK--EDSDFRRIAELVEEMSVVEVVVGLPRNLREGTG
.:****:*** * ..**:: *. * .***::*** .* : .
Mflv_3762|M.gilvum_PYR-GCK SSAQDATEVADQLAARIAPVPVRLADERFTTVTAQRELREAGVRARGQKS
Mvan_2641|M.vanbaalenii_PYR-1 SSAQDATEVAEVLAGRIAPVPVRLADERFTTVTAQRALREAGVRARGQRS
MSMEG_3026|M.smegmatis_MC2_155 ASAHDAVDVAEQLARRIAPTPVRLADERLTTVSAQRSLREAGVRARGQKA
Mb2584c|M.bovis_AF2122/97 RSAQDAIELAEALARRVSPTPVRLADERLTTVSAQRSLRQAGVRASEQRA
Rv2554c|M.tuberculosis_H37Rv RSAQDAIELAEALARRVSPTPVRLADERLTTVSAQRSLRQAGVRASEQRA
MAV_3430|M.avium_104 ASALDAIDLADQLARRIAPTPVRLADERLTTVAAQRSLRAAGVRAKEQRA
MMAR_2171|M.marinum_M SSAQDAIDLADELAARVAPIPVRLADERLTTVSAQRSLREAGVRAKNQRA
MUL_1755|M.ulcerans_Agy99 SSAQDAIDLADELAARVAPIPVRLADERLTTVSAQRSLREAGVRAKNQRA
MLBr_00513|M.leprae_Br4923 SSAKDAIDTAEALARRVAPVPVRLVDERLTTVSAQRLLRAAGVRAKDQRA
TH_0113|M.thermoresistible__bu PAAQDAIALADELARRIDPVPVRLADERLTTVTAQRSLRAAGVRSRDQRA
MAB_2850c|M.abscessus_ATCC_199 SSARDAAGFARELGERIAPVPVRLVDERFTTTTAQRSLREAGVRSRQQRG
:* ** * *. *: * ****.***:**.:*** ** ****: *:.
Mflv_3762|M.gilvum_PYR-GCK MIDQAAAVGILQNWLDQRRAALPAHGEVLDD----------
Mvan_2641|M.vanbaalenii_PYR-1 VIDQAAAVGILQNWLDQRRAALPAHGEVLDD----------
MSMEG_3026|M.smegmatis_MC2_155 VIDQVAAVGILQGWLDQRRAALAARREGTDG----------
Mb2584c|M.bovis_AF2122/97 VIDQAAAVAILQSWLDERLAAMAGTQEGSDA----------
Rv2554c|M.tuberculosis_H37Rv VIDQAAAVAILQSWLDERLAAMAGTQEGSDA----------
MAV_3430|M.avium_104 VIDQAAAVAILQSWLDQRRAAVREAGDG-------------
MMAR_2171|M.marinum_M VIDQAAAVAILQAWLDQRRTLAGGRADG-------------
MUL_1755|M.ulcerans_Agy99 VIDQAAAVAILQAWLDQRRTLAGGRADG-------------
MLBr_00513|M.leprae_Br4923 VIDQAAAVVILQNWLDQCRAATPARADEPTTGSVAGEVIDG
TH_0113|M.thermoresistible__bu MIDQVAAVDILQGWLDQRRAALSARGEAGDD----------
MAB_2850c|M.abscessus_ATCC_199 IIDQAAAVAILQDWLEQRRRCGSE-GDTP------------
:***.*** *** **:: :