For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
TGFTEIAAAGPLLVALGVCMLAGLVSFVSPCVVPLVPGYLSYLAAVVGVSHEETQPGAGVIKTPPAARWR VAGSAVLFVAGFTTVFVLDTVAVLGMTTVLTTHQVLLQRVGGVLTIVMGLVFVGLLPALQRQVQFSLRQL TTVAGAPVLGTVFALGWTPCLGPTLSGVITVAAATDGVNVTRGILLVIAYCMGLGIPFVLLASGSAQAVA GLRWLRQYGRAIQVFGGMLLIAIGAALIAGVWDDFVSWLRDAVVSDMRVSI*
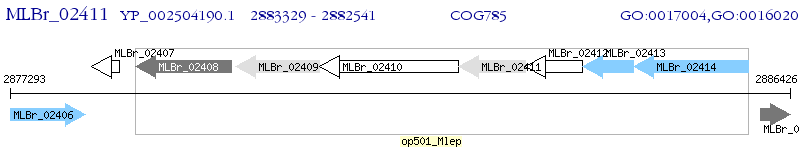
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. leprae Br4923 | MLBr_02411 | - | - | 100% (262) | putative cytochrome C-type biogenesis protein |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0540 | ccdA | 1e-113 | 78.63% (262) | cytochrome C-type biogenesis protein CcdA |
| M. gilvum PYR-GCK | Mflv_0051 | - | 3e-98 | 67.70% (257) | cytochrome c biogenesis protein, transmembrane region |
| M. tuberculosis H37Rv | Rv0527 | ccdA | 1e-113 | 78.63% (262) | cytochrome C-type biogenesis protein CcdA |
| M. abscessus ATCC 19977 | MAB_3975c | - | 8e-96 | 65.90% (261) | putative cytochrome C biogenesis protein CcdA |
| M. marinum M | MMAR_0873 | ccsA | 1e-112 | 76.81% (263) | cytochrome C-type biogenesis protein, CcsA |
| M. avium 104 | MAV_4618 | - | 1e-115 | 78.24% (262) | cytochrome C biogenesis protein transmembrane region |
| M. smegmatis MC2 155 | MSMEG_0972 | - | 1e-102 | 69.85% (262) | cytochrome C biogenesis protein transmembrane region |
| M. thermoresistible (build 8) | TH_2250 | ccsA | 1e-103 | 67.64% (275) | POSSIBLE CYTOCHROME C-TYPE BIOGENESIS PROTEIN CCDA |
| M. ulcerans Agy99 | MUL_0626 | ccsA | 1e-112 | 76.05% (263) | cytochrome C-type biogenesis protein, CcsA |
| M. vanbaalenii PYR-1 | Mvan_0867 | - | 5e-98 | 67.69% (260) | cytochrome c biogenesis protein, transmembrane region |
CLUSTAL 2.0.9 multiple sequence alignment
Mb0540|M.bovis_AF2122/97 ------MTGFTEIAAVGPLLVAVGVCLLAGLVSFASPCVVPLVPGYLSYL
Rv0527|M.tuberculosis_H37Rv ------MTGFTEIAAVGPLLVAVGVCLLAGLVSFASPCVVPLVPGYLSYL
MLBr_02411|M.leprae_Br4923 ------MTGFTEIAAAGPLLVALGVCMLAGLVSFVSPCVVPLVPGYLSYL
MAV_4618|M.avium_104 ------MTGLTQIAAAGPLLAALGVCLLAGLVSFASPCVVPLVPGYLSYL
MMAR_0873|M.marinum_M ------MTGFAEVAAAGPLLVALGLCVLAGLVSFASPCVVPLVPGYLSYL
MUL_0626|M.ulcerans_Agy99 ------MTGFAEVAAAGPLLVALGLCVLAGLVSFASPCVVPLVPGYLSYL
MSMEG_0972|M.smegmatis_MC2_155 ------MTGFAEIAAAGPVLLALGVSVLAGLVSFASPCVVPLVPGYLSYL
TH_2250|M.thermoresistible__bu ------MTGFAELAAAGPVLVAVAISILAGLVSFASPCVVPLVPGYLSYL
Mflv_0051|M.gilvum_PYR-GCK MNVD----QIDQLTASGPLLLAMGLAALAGLVSFASPCVVPLVPGYLSYL
Mvan_0867|M.vanbaalenii_PYR-1 MTLD----QIDQLTATGPLLLATGLAALAGLVSFASPCVIPLVPGYLSYL
MAB_3975c|M.abscessus_ATCC_199 MILADAGSAFQDAATSGPLLLALGACALAGLVSFASPCVIPLVPGYLSYL
: : :: **:* * . . *******.****:**********
Mb0540|M.bovis_AF2122/97 AAVVGV---DEQLP-AGVVK----------PPVAARWRVAGSAALFVAGF
Rv0527|M.tuberculosis_H37Rv AAVVGV---DEQLP-AGVVK----------PPVAARWRVAGSAALFVAGF
MLBr_02411|M.leprae_Br4923 AAVVGVS-HEETQPGAGVIK----------TPPAARWRVAGSAVLFVAGF
MAV_4618|M.avium_104 AALVGV----EEQPQAGAVQ----------APPGARWRVAGSAALFVAGF
MMAR_0873|M.marinum_M AAVVGVDGSGDGRAAGVGVK----------APAGVRWRVAGSALLFVAGF
MUL_0626|M.ulcerans_Agy99 AAVVGVDGSGDGRAAGVGVK----------APAGVRWRVAGSALLFVAGF
MSMEG_0972|M.smegmatis_MC2_155 AAVVGVDDD--PVRTGGER----------VALRSARLRVAGAAALFVAGF
TH_2250|M.thermoresistible__bu AAVVGVPKEAAPVVTGGPRRDPPADSGAGMSPRSARLRVTGAAALFVAGF
Mflv_0051|M.gilvum_PYR-GCK AAVVGVD----DGPAGS----------GAVAVKGARLRVAGAAALFVAGF
Mvan_0867|M.vanbaalenii_PYR-1 AAVVGVD----QSAAGSAAP-------GAVAVKGARLRVAGAAGLFVAGF
MAB_3975c|M.abscessus_ATCC_199 AAVVGAD----NGPENG------------AGAQPHRLRVTGSALLFVAGF
**:**. * **:*:* ******
Mb0540|M.bovis_AF2122/97 TTVFVLGTVAVLGMTTTLITNQLLLQRVGGVLIVVMGLVFVGFIGALQRQ
Rv0527|M.tuberculosis_H37Rv TTVFVLGTVAVLGMTTTLITNQLLLQRVGGVLIVVMGLVFVGFIGALQRQ
MLBr_02411|M.leprae_Br4923 TTVFVLDTVAVLGMTTVLTTHQVLLQRVGGVLTIVMGLVFVGLLPALQRQ
MAV_4618|M.avium_104 TAVFVLGTVAVLGMTTTLITNQLLLQRAGGVLTIVMGLVFVGLIPALQRQ
MMAR_0873|M.marinum_M TTVFVLGTVAVLGMTTTLISNQLVLQRAGGVLTIIMGLVFVGLIPALQRQ
MUL_0626|M.ulcerans_Agy99 TTVFVLGTVAVLGMTTTLISNQLVLQRAGGVLTIIMGLVFVGLIPALQRQ
MSMEG_0972|M.smegmatis_MC2_155 TVVFLLGAVAVLGMTTTLIANQLLLQRIGGVVTIVMGLVFVGFIPALQRQ
TH_2250|M.thermoresistible__bu TVVFLLGTVAVLGMTTTLITNQVLLQRIGGVITIVMGLVFIGFIPALQRQ
Mflv_0051|M.gilvum_PYR-GCK TVVFLLGTVAVLGMTTTLITNQLLLQRIGGVITILMGLVFIGLVPVLQRD
Mvan_0867|M.vanbaalenii_PYR-1 TVVFLLGTVAVLGMTTALITNQLLLQRLGGVVTILMGLVFIGLVPVLQRD
MAB_3975c|M.abscessus_ATCC_199 TAVFLLGTVAVMGMTTSLITNQVLLQRIGGVVTIAMGLVFIGLVPALQRD
*.**:*.:***:**** * ::*::*** ***: : *****:*:: .***:
Mb0540|M.bovis_AF2122/97 ARFTPRQLTSVAGAPVLGAVFALGWTPCLGPTLTGVITVASATEGASVAR
Rv0527|M.tuberculosis_H37Rv ARFTPRQLTSVAGAPVLGAVFALGWTPCLGPTLTGVITVASATEGASVAR
MLBr_02411|M.leprae_Br4923 VQFSLRQLTTVAGAPVLGTVFALGWTPCLGPTLSGVITVAAATDGVNVTR
MAV_4618|M.avium_104 ARFSPRQLTTVAGAPLLGAVFALGWTPCLGPTLAGVITVASATDGASVAR
MMAR_0873|M.marinum_M ARFSPRQVTTVAGAPLLGAVFALGWTPCLGPTLTGVITVASATDGASVAR
MUL_0626|M.ulcerans_Agy99 ARFSPRQVTTVAGAPLLGAVFALGWTPCLGPTLTGVITVASATNGASVAR
MSMEG_0972|M.smegmatis_MC2_155 ARFTPRQLSTVAGAPLLGAVFALGWTPCLGPTLTGVIAVASATDGATVAR
TH_2250|M.thermoresistible__bu ARFAPRQWSTVAGAPLLGAVFALGWTPCLGPTLTGVITVASATDGASVAR
Mflv_0051|M.gilvum_PYR-GCK TRFTPRRISTVGGAPLLGAVFALGWTPCLGPTLTGVIAVASATEGSNVAR
Mvan_0867|M.vanbaalenii_PYR-1 TRFTPRQISTLGGAPLLGAVFALGWTPCLGPTLTGVIAVASATEGSNVAR
MAB_3975c|M.abscessus_ATCC_199 IRFTPRRVSTVAGAPLLGAVFALGWTPCLGPTLTGVVAVASATEGADVAR
:*: *: :::.***:**:**************:**::**:**:* *:*
Mb0540|M.bovis_AF2122/97 GIVLVIAYCLGLGIPFVLLAFGSAWAVAGLGWLRRHTRAIQIFGGALLIA
Rv0527|M.tuberculosis_H37Rv GIVLVIAYCLGLGIPFVLLAFGSAWAVAGLGWLRRHTRAIQIFGGALLIA
MLBr_02411|M.leprae_Br4923 GILLVIAYCMGLGIPFVLLASGSAQAVAGLRWLRQYGRAIQVFGGMLLIA
MAV_4618|M.avium_104 GIVLVIAYCLGLGIPFVLLAFGSAGAVAGLGWLRRHTRAIQIFGGVLLIT
MMAR_0873|M.marinum_M GIVLVIAYCLGLGIPFVLLAFGSAGAVAGLGWLRRHTRAIQIFGGVLLIA
MUL_0626|M.ulcerans_Agy99 GIVLVIAYCLGLGIPFVLLAFGSAGAVAGLGWLRRHTRAIQIFGGVLLIV
MSMEG_0972|M.smegmatis_MC2_155 GVVLVLAYCLGLGIPFVLLALGSARAVQGLGWLRRHTRTIQIFGGVLLIL
TH_2250|M.thermoresistible__bu GVVLVIAYCLGLGIPFILLAFGSARAVQGLGWLRRNTRTIQIFGGVLLIL
Mflv_0051|M.gilvum_PYR-GCK GVVLVIAYCLGLGIPFVLLAFGSARAVAGLGWLRRNTRTIQILGGVLMIL
Mvan_0867|M.vanbaalenii_PYR-1 GVVLVIAYCLGLGIPFVLLAFGSARAVAGLGWLRRHTRTIQVLGGVLMIL
MAB_3975c|M.abscessus_ATCC_199 GIALVIAYCLGLGMPFLLLAFGSARAVRGMDWLRRHTRMIQVIGGIALLA
*: **:***:***:**:*** *** ** *: ***: * **::** ::
Mb0540|M.bovis_AF2122/97 VGAALVTGVWNDVVSWLRDAFVSDVRLPI
Rv0527|M.tuberculosis_H37Rv VGAALVTGVWNDVVSWLRDAFVSDVRLPI
MLBr_02411|M.leprae_Br4923 IGAALIAGVWDDFVSWLRDAVVSDMRVSI
MAV_4618|M.avium_104 VGALLVTGLWNDFVSWLRDAFVSDVRLPI
MMAR_0873|M.marinum_M VGLALVTGVWNDVVSWLRDAFVSDVRLPI
MUL_0626|M.ulcerans_Agy99 VGLALVTGVWNDVVSWLRDAFVSDVRLPI
MSMEG_0972|M.smegmatis_MC2_155 VGAALVTGLWNEFVSYVRDAFVSDVRLPI
TH_2250|M.thermoresistible__bu VGAALVTGLWNEFISWVRDAFVSDVRLPI
Mflv_0051|M.gilvum_PYR-GCK VGAALVTGLWNEFVSWVRDAFVSDVRLPI
Mvan_0867|M.vanbaalenii_PYR-1 VGAALVTGLWNDFVSWIRDAFVSDVRLPI
MAB_3975c|M.abscessus_ATCC_199 VGVALITGVWADFVAWVRDSFVSDARLPI
:* *::*:* :.::::**:.*** *:.*