For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
LSKLVALAAVIALSPITVIPAVLVLHAPRPRPASLAFLGGWVLGLVGLTTAFVGGSELLSGLHATPPKWA SWVRLVFGVALLALAAYHWFTRHRQRSMPRWMRSFAALSPVRAGVVGAVLVPLRPEVLILCAAAGLAVGN SSLSAGAQLVAAAIFVVLAASTVAVPILVYVGAGDRFDDALERLKAWMEANHDAMLAVILLLIGLIVLYN GIRALA*
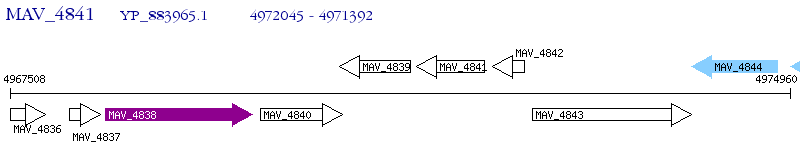
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. avium 104 | MAV_4841 | - | - | 100% (217) | hypothetical protein MAV_4841 |
| M. avium 104 | MAV_4839 | - | 3e-68 | 56.02% (216) | hypothetical protein MAV_4839 |
| M. avium 104 | MAV_1758 | - | 3e-14 | 31.70% (224) | hypothetical protein MAV_1758 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3511c | - | 8e-75 | 61.11% (216) | integral membrane protein |
| M. gilvum PYR-GCK | Mflv_0252 | - | 3e-60 | 51.61% (217) | hypothetical protein Mflv_0252 |
| M. tuberculosis H37Rv | Rv3481c | - | 9e-75 | 61.11% (216) | integral membrane protein |
| M. leprae Br4923 | MLBr_01233 | - | 6e-09 | 25.64% (234) | hypothetical protein MLBr_01233 |
| M. abscessus ATCC 19977 | MAB_0934 | - | 4e-15 | 27.98% (218) | hypothetical protein MAB_0934 |
| M. marinum M | MMAR_4961 | - | 4e-76 | 62.04% (216) | hypothetical protein MMAR_4961 |
| M. smegmatis MC2 155 | MSMEG_0729 | - | 4e-60 | 51.39% (216) | hypothetical protein MSMEG_0729 |
| M. thermoresistible (build 8) | TH_2356 | - | 1e-61 | 53.70% (216) | conserved hypothetical protein |
| M. ulcerans Agy99 | MUL_4034 | - | 3e-76 | 62.04% (216) | hypothetical protein MUL_4034 |
| M. vanbaalenii PYR-1 | Mvan_0642 | - | 1e-61 | 50.93% (216) | hypothetical protein Mvan_0642 |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_0252|M.gilvum_PYR-GCK ------MPGTWGSILIELIPLALVIALSPLSIIPAVLVLSAARPRPTGLA
Mvan_0642|M.vanbaalenii_PYR-1 ------MPGSWGPILTELIPLALVIALSPLSIIPAVLVLHSPRPRPTGLA
MSMEG_0729|M.smegmatis_MC2_155 ------MTGIWSSVLTELIPLAMVVALSPLSIIPAVLVLQTPRPRPTGLA
TH_2356|M.thermoresistible__bu ------MTGSWASLLTELIPLALVIALSPLSIIPGVLVLHTPRPRPTGLA
MMAR_4961|M.marinum_M MRKLLPVTGNSGSVLASLVPLALVIAVSPITIIPAVLVLHAPRPRPTGLA
MUL_4034|M.ulcerans_Agy99 MRKLLPVTGNSGSVLASLVPLALVIAVSPITIIPAVLVLHAPRPRPTGLA
Mb3511c|M.bovis_AF2122/97 MRGLLPVAGHWVSVLTGLVPLALVIALSPLSVIPAVLVVHSPQPRPSSLA
Rv3481c|M.tuberculosis_H37Rv MRGLLPVAGHWVSVLTGLVPLALVIALSPLSVIPAVLVVHSPQPRPSSLA
MAV_4841|M.avium_104 -------------MLSKLVALAAVIALSPITVIPAVLVLHAPRPRPASLA
MLBr_01233|M.leprae_Br4923 -------------MWITILLMAIAISLEPFRIGMTVLMLNRPRPTLQLLV
MAB_0934|M.abscessus_ATCC_1997 -------------MWITLLVMAIAVSFEPFRLGMTVLMLNRPRPMLQLLA
: :: :* .::..*: : **:: .:* *.
Mflv_0252|M.gilvum_PYR-GCK FLA-GWVLGLAA-LTAVFLQLSDLVGGIGTKPPSWASWLRIVVGVLLILW
Mvan_0642|M.vanbaalenii_PYR-1 FLV-GWLVGLAA-LTAIFLQLSDLVGGIGTQPPSWASWLRIVVGVGLIGF
MSMEG_0729|M.smegmatis_MC2_155 FLG-GWLVGMAA-LTALFIEVSNLFGRLD-KPPVWASWLRIVLGTALIIF
TH_2356|M.thermoresistible__bu FLL-GWLIGLAV-LAAAFVALSGLLGGGLEHPPGWASWLRVVIGALLIVF
MMAR_4961|M.marinum_M FLG-GWVLGLIA-VTAMFVAGSELFGGLRRSPPAWASWARIVVGSLLILF
MUL_4034|M.ulcerans_Agy99 FLG-GWVLGLIA-VTAMFVAGSELFGGLRRSPPAWASWARIVVGSLLILF
Mb3511c|M.bovis_AF2122/97 FLG-GWLLGLAV-VTAVFVAASGALGGLSTTSPAWASWLRVVLGSALIVF
Rv3481c|M.tuberculosis_H37Rv FLG-GWLLGLAV-VTAVFVAASGALGGLSTTSPAWASWLRVVLGSALIVF
MAV_4841|M.avium_104 FLG-GWVLGLVG-LTTAFVGGSELLSGLHATPPKWASWVRLVFGVALLAL
MLBr_01233|M.leprae_Br4923 FLCSGFTMGMTVGFVVLFVFRRRLMASMQLTLPKVQILIGVLALLVAAVL
MAB_0934|M.abscessus_ATCC_1997 FLSGGFVMGMAVGLVVLFAFRNVSTGSAHFTLPRVQIGIGVAALLVAALV
** *: :*: ... * . * :
Mflv_0252|M.gilvum_PYR-GCK GAYRWLKRASSEHMPGWMTRISALTPAKAAATAAALTVLNVKVLFICAAA
Mvan_0642|M.vanbaalenii_PYR-1 GGYRWVTRKKSEHMPGWMTRMSTLTPAKAGATAAALTVLNPKVLFICAAA
MSMEG_0729|M.smegmatis_MC2_155 GVYRWLNRSKAAHTPKWMQSMGKLTPPRAGVTGVVLTIVNPKVLFICAAA
TH_2356|M.thermoresistible__bu GGYRWATRHRSTEMPRWMRSLTTASPPRAAITAAVLTVANPKVLFICAAA
MMAR_4961|M.marinum_M GIYRWLNRHGEVESPRWMRSFATITPRRAGITAAVLVLVRPEVVIICLAA
MUL_4034|M.ulcerans_Agy99 GIYRWLNRHGEVESPRWMRSFATITPRRAGITAAVLVLVRPEVVIICLAA
Mb3511c|M.bovis_AF2122/97 GVLRWLTRHRHTEMPGWMRAFASFTPARAGLVGAVLVVVRPEVLIICAAA
Rv3481c|M.tuberculosis_H37Rv GVLRWLTRHRHTEMPGWMRAFASFTPARAGLVGAVLVVVRPEVLIICAAA
MAV_4841|M.avium_104 AAYHWFTRHRQRSMPRWMRSFAALSPVRAGVVGAVLVPLRPEVLILCAAA
MLBr_01233|M.leprae_Br4923 TVQVCISSEPPAESP--VDSASGPPKPSKWAPRPLARLLNGDSLWVAGVA
MAB_0934|M.abscessus_ATCC_1997 ASRLRG---PASHAQ--LDKVSAR----------IQSIATGGSLWVAGVA
: : :. .*
Mflv_0252|M.gilvum_PYR-GCK GLAIGS--------------SGLDTPTVWAAVLWFVLVAASTVAAPIVAY
Mvan_0642|M.vanbaalenii_PYR-1 GLAIGS--------------AGLDSPTVWGAMIWFVAIAGLTVALPVLGY
MSMEG_0729|M.smegmatis_MC2_155 GLAIGS--------------AGLGQPQVWAAAIWFVVFASSTVALPILAY
TH_2356|M.thermoresistible__bu GLAIGT--------------AGVGVG-AWPAVLYYVLVAGSTVALPILAY
MMAR_4961|M.marinum_M GLAIGT--------------GGLGVAMRWVCGAIFVAISASTVAGPILAF
MUL_4034|M.ulcerans_Agy99 GLAIGT--------------GGLGVAMRWVCGAIFVAISASTVAGPILAF
Mb3511c|M.bovis_AF2122/97 GLAIGS--------------GGHGAAGSWIYTAFFAMLAASTVAIPILAY
Rv3481c|M.tuberculosis_H37Rv GLAIGS--------------GGHGAAGSWIYTAFFAMLAASTVAIPILAY
MAV_4841|M.avium_104 GLAVGN--------------SSLSAGAQLVAAAIFVVLAASTVAVPILVY
MLBr_01233|M.leprae_Br4923 GLGIALPSVDYLAALAVILTSGAAATTQVGALLMFNVVAFALIEIPLAAY
MAB_0934|M.abscessus_ATCC_1997 GLGIALPSVDFLAALAVIVASGSPPATQISALLMFNVIAFALVEIPFLAH
**.:. .. : .: : *. .
Mflv_0252|M.gilvum_PYR-GCK AISGERLDPTLNRVQAWMERQHATLIAAILVVIGLLVLYKGIHALAG
Mvan_0642|M.vanbaalenii_PYR-1 AVSGERLDPTLNRLKAWMERQHAVLVAAILVAIGLLVLYKGIHALIG
MSMEG_0729|M.smegmatis_MC2_155 AVSGSKLDPTLARLGEWMERQHAVLVAVILVVIGLLVLYKGLHAL--
TH_2356|M.thermoresistible__bu ALAPDRLGQPLERLREWMERQHGALVAAILVVIGLMVLFKGIHAL--
MMAR_4961|M.marinum_M VGAGDRLEDSLDRLKEWMEKNHAGMLAAVLILIGVMVLYNGIHAL--
MUL_4034|M.ulcerans_Agy99 VGAGDRLEDSLDRLKEWMEKNHAGMLAAVLILIGVMVLYNGIHAL--
Mb3511c|M.bovis_AF2122/97 VAAGDRLDDSLERLKDWMEKNHAGMVAAILVVIGLLLLYNGVHAM--
Rv3481c|M.tuberculosis_H37Rv VAAGDRLDDSLERLKDWMEKNHAGMVAAILVVIGLLLLYNGVHAM--
MAV_4841|M.avium_104 VGAGDRFDDALERLKAWMEANHDAMLAVILLLIGLIVLYNGIRALA-
MLBr_01233|M.leprae_Br4923 LLAPDTTRAWTAALNNWIRSRRRLEVATLLAGVSCLLLAVGIAGL--
MAB_0934|M.abscessus_ATCC_1997 AVAPERTAEAMARLNSWIQSNRRRNIALVLTAVGCVLLAVGLAGI--
: . : *:. .: :* :* :. ::* *: .: