For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
TGLLDGKVALVTGAGHGIGRGHALELAKHGATVIVNDLGTSLSGEGTGKVADEVVAIIESRGGTAVSDFS DVGDEEQVDLAVERAYSQLGRLDIVVNNAGIVRDKAIWNMTADDFDLVMRVHVRGSWLTSRAVARKWRAE SKESGGKVYGRIINTTSGAGLHGHFGQTNYSAAKAAIVGLTQTLSLELASIGATVNAISPGGRTRMSASM PGAQAPIEPDERAEDEFDPKDPSLGSPVVAWLASPEAGHVSGQVIRAMGEHLQLLKGWHPVASVSNGEKR WDAQKLGAIMATDVFGTRNTGLRLGG*
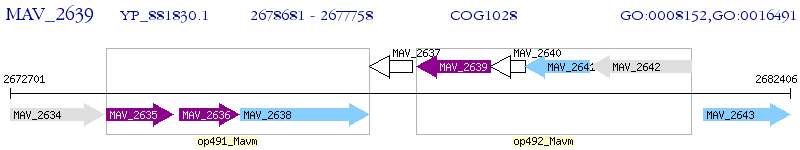
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. avium 104 | MAV_2639 | - | - | 100% (307) | short chain dehydrogenase |
| M. avium 104 | MAV_1937 | - | e-170 | 95.77% (307) | short chain dehydrogenase |
| M. avium 104 | MAV_0612 | - | 5e-61 | 44.44% (297) | short chain dehydrogenase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3578c | - | 3e-59 | 43.88% (294) | short chain dehydrogenase |
| M. gilvum PYR-GCK | Mflv_1492 | - | 2e-59 | 43.30% (291) | short chain dehydrogenase |
| M. tuberculosis H37Rv | Rv3548c | - | 4e-59 | 43.88% (294) | short chain dehydrogenase |
| M. leprae Br4923 | MLBr_02565 | fabG | 3e-30 | 36.40% (261) | 3-ketoacyl-(acyl-carrier-protein) reductase |
| M. abscessus ATCC 19977 | MAB_0608 | - | 6e-61 | 42.67% (307) | short chain dehydrogenase |
| M. marinum M | MMAR_5036 | - | 2e-61 | 46.58% (292) | short-chain type dehydrogenase/reductase |
| M. smegmatis MC2 155 | MSMEG_5999 | - | 2e-61 | 44.52% (292) | short chain dehydrogenase |
| M. thermoresistible (build 8) | TH_3887 | - | 7e-61 | 44.67% (291) | PUTATIVE short chain dehydrogenase |
| M. ulcerans Agy99 | MUL_4110 | - | 3e-61 | 46.58% (292) | short chain dehydrogenase |
| M. vanbaalenii PYR-1 | Mvan_5264 | - | 8e-60 | 43.30% (291) | short chain dehydrogenase |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_1492|M.gilvum_PYR-GCK ----------MGLLDGRVVIVTGAGGGIGRAHALAFAAEGARVVVNDIGV
Mvan_5264|M.vanbaalenii_PYR-1 ----------MGLLDGRVVIVTGAGGGIGRAHALAFAAEGARVVVNDIGV
MSMEG_5999|M.smegmatis_MC2_155 ----------MGLLDGRVVIVTGAGGGIGRAHALAFAAEGARVVVNDIGV
TH_3887|M.thermoresistible__bu ----------MGIVDGRVVIVTGAGNGIGRAHALAFAAEGARVVVNDIGV
Mb3578c|M.bovis_AF2122/97 ----------MGLVDGRVVIVTGAGGGIGRAHALAFAAEGARVVVNDIGV
Rv3548c|M.tuberculosis_H37Rv ----------MGLVDGRVVIVTGAGGGIGRAHALAFAAEGARVVVNDIGV
MMAR_5036|M.marinum_M ----------MGLLDGRVVIVTGAGGGIGRAHALAFAAEGARVVVNDIGV
MUL_4110|M.ulcerans_Agy99 ----------MGLLDGRVVIVTGAGGGIGRAHALAFAVEGARVVVNDIGV
MAB_0608|M.abscessus_ATCC_1997 ---------MPGLLDGRVVIITGAGRGIGRAHALAFAAEGARVVVNDIGV
MAV_2639|M.avium_104 ---------MTGLLDGKVALVTGAGHGIGRGHALELAKHGATVIVNDLGT
MLBr_02565|M.leprae_Br4923 MAPKCSSDLFSQIVSSGPGSFVAKQLGVPRPETLRRYRPSDAPLCGSLLI
::.. ... *: * .:* . : ..:
Mflv_1492|M.gilvum_PYR-GCK GLDGSPAGGGSAAQTVVDEIIAA--GGEAVTSG----------ANVADWG
Mvan_5264|M.vanbaalenii_PYR-1 GLDGSPAGGGSAAQAVVDEIVGA--GGEAVTSG----------ANVADWA
MSMEG_5999|M.smegmatis_MC2_155 GLDGSPASGGSAAQTVVDEIVAA--GGEAVANG----------SNVADWA
TH_3887|M.thermoresistible__bu GLDGSPASGGSAAQAVVDEIKAA--GGEAVANG----------SNVADWA
Mb3578c|M.bovis_AF2122/97 GLDGSPASGGSAAQDVVDEILAA--GGQAVADG----------SDISDWD
Rv3548c|M.tuberculosis_H37Rv GLDGSPASGGSAAQDVVDEILAA--GGQAVADG----------SDISDWD
MMAR_5036|M.marinum_M GLDGSPAGGGSAAHGVVDEITAA--GGEAVANG----------SDVSNWQ
MUL_4110|M.ulcerans_Agy99 GLDGSPAGGGSAAHGVVDEITAA--GGEAVANG----------SDVSNWQ
MAB_0608|M.abscessus_ATCC_1997 GLDGS--AGISPAQEVVEEITAA--GGEAVTNG----------DDVADWS
MAV_2639|M.avium_104 SLSGE--GTGKVADEVVAIIESR--GGTAVSDF----------SDVGDEE
MLBr_02565|M.leprae_Br4923 GGDGRLVEPLRATLEKDYDLVSNNLGGRWTDSFGGLVFDATGITAPAELK
. .* : : . ** . . .:
Mflv_1492|M.gilvum_PYR-GCK QAEGLIQTAVDAFGGLDVLVNN-----AGIVRDRMFANTSEEEFDAVTAV
Mvan_5264|M.vanbaalenii_PYR-1 QAEGLIQTAVDSFGTLDVLVNN-----AGIVRDRMFANTSEEEFDAVTAV
MSMEG_5999|M.smegmatis_MC2_155 QAESLVQTAVDTFGGLDVLVNN-----AGIVRDRMFANTSEEEFDAVVAV
TH_3887|M.thermoresistible__bu QAEELVKTAIDTYGGLDVLVNN-----AGIVRDRMIANTSEEEFDAVVAV
Mb3578c|M.bovis_AF2122/97 QAANLIQAAVETYGGVDVLVNN-----AGIVRDRMIANTSEEEFDAVIAV
Rv3548c|M.tuberculosis_H37Rv QAANLIQAAVETYGGVDVLVNN-----AGIVRDRMIANTSEEEFDAVIAV
MMAR_5036|M.marinum_M QAADLISTAVDTFGGLDVLVNN-----AGIVRDRMMANTSEEEFDAVIAV
MUL_4110|M.ulcerans_Agy99 QAADLISTAVDTFGGLDVLVNN-----AGIVRDRMMANTSEEEFDAVIAV
MAB_0608|M.abscessus_ATCC_1997 GAKNLIDTAVNSFGRLDVLVNN-----AGFLRDRMLANMSEEEWDAVIRV
MAV_2639|M.avium_104 QVDLAVERAYSQLGRLDIVVNN-----AGIVRDKAIWNMTADDFDLVMRV
MLBr_02565|M.leprae_Br4923 GLHNFFTPLLRNLGRCARVVVIGTTPDATTSTDERIAQRALEGFSRSLGK
. * :* * *. : : : : :.
Mflv_1492|M.gilvum_PYR-GCK HLKGHFATMRHAAAYWRAKSKAGDT---VDARIINTSSGAGLQGSVGQAN
Mvan_5264|M.vanbaalenii_PYR-1 HLKGHFATMKHAAAYWRAKSKAGES---VDARIINTSSGAGLQGSVGQAN
MSMEG_5999|M.smegmatis_MC2_155 HLKGHFATMKHAAAYWRAKSKAGET---VDARIINTSSGAGLQGSVGQAN
TH_3887|M.thermoresistible__bu HLKGHFATTRHAAAYWRGLAKEGKA---VDARIINTSSGAGLQGSVGQGN
Mb3578c|M.bovis_AF2122/97 HLKGHFATMRHAASHWRGLSKAGKAPKDIDARIINTSSGAGLQGSVGQGN
Rv3548c|M.tuberculosis_H37Rv HLKGHFATMRHAASHWRGLSKAGKAPKDIDARIINTSSGAGLQGSVGQGN
MMAR_5036|M.marinum_M HLKGHFATMRHAASYWRGLSKAGKA---VDARIINTSSGAGLQGSVGQAN
MUL_4110|M.ulcerans_Agy99 HLKGHFATMRHAASYWRGLSKVGKA---VDARIINTSSGAGLQGSVGQAN
MAB_0608|M.abscessus_ATCC_1997 HLKGHFAPLRHAAAYWRDEFKAGNA---VDARIINTSSAAGLQGSVGQGN
MAV_2639|M.avium_104 HVRGSWLTSRAVARKWRAESKESGG--KVYGRIINTTSGAGLHGHFGQTN
MLBr_02565|M.leprae_Br4923 ELR---RGATVALVYLSPDAKPAATGLESTMRFILSAKSAYVNGQVFYVG
.:: . * . *:* ::..* ::* . .
Mflv_1492|M.gilvum_PYR-GCK YSAAKAG------IAALTLVAAAEMGRYGVSVNAIAPSARTRMTETVFAD
Mvan_5264|M.vanbaalenii_PYR-1 YSASKAG------IAALTLVAAAEMGRYGVSVNAIAPSARTRMTESVFAD
MSMEG_5999|M.smegmatis_MC2_155 YSAAKAG------IAALTLVAAAEMGRYGVTVNAIAPSARTRMTETVFAD
TH_3887|M.thermoresistible__bu YSAAKAG------IAALTLVAAAELGRYGVTVNAIAPSARTRMTETVFAE
Mb3578c|M.bovis_AF2122/97 YSAAKAG------IAALTLVGAAEMRRYGVTVNAIAPAARTRMTETVFAE
Rv3548c|M.tuberculosis_H37Rv YSAAKAG------IAALTLVGAAEMRRYGVTVNAIAPAARTRMTETVFAE
MMAR_5036|M.marinum_M YSAAKAG------IAALTLVGAAEMGRYGVTVNAIAPAARTRMTETVFAE
MUL_4110|M.ulcerans_Agy99 YSAAKAG------IAALTLVGAAEMGRYGVTVNAIAPAARTRMTETVFAE
MAB_0608|M.abscessus_ATCC_1997 YAAAKAG------IATLTLQAAAEMGRYGITVNAIAPAARTRMTEAVFAE
MAV_2639|M.avium_104 YSAAKAA------IVGLTQTLSLELASIGATVNAISPGGRTRMSASMPGA
MLBr_02565|M.leprae_Br4923 AADSTPPTNWDRPLENKVAIVTGAARGIGAAIAEVVARDGARVVAIDVES
: :.. : . : * :: : . :*:
Mflv_1492|M.gilvum_PYR-GCK MMATQDK----------DFDAMAPENVSPLVVWLGSVE----------SR
Mvan_5264|M.vanbaalenii_PYR-1 MMATQDK----------DFDSMAPENISPLVVWLGSVE----------SR
MSMEG_5999|M.smegmatis_MC2_155 MMATQDQ----------EFDAMAPENISPIVVWLGSAE----------AR
TH_3887|M.thermoresistible__bu MMATQDQ----------EFDSMAPENISPLVVWLGSAE----------SK
Mb3578c|M.bovis_AF2122/97 VMAKPQE----------GFDAMAPENVSPLVVWLGSAE----------SR
Rv3548c|M.tuberculosis_H37Rv MMAKPQE----------GFDAMAPENVSPLVVWLGSAE----------SR
MMAR_5036|M.marinum_M MMAKPDE----------GFDAMAPENVSPLVVWLASAE----------AG
MUL_4110|M.ulcerans_Agy99 MMAKPDE----------GFDAMAPENVSPLVVWLASAE----------AG
MAB_0608|M.abscessus_ATCC_1997 TMAKPEDG---------AFDAMAPENVSPLVVWLGSPE----------SR
MAV_2639|M.avium_104 QAPIEPDERAED-----EFDPKDPSLGSPVVAWLASPE----------AG
MLBr_02565|M.leprae_Br4923 AAEALDETANRVGGTTLSLDVTADDAVDKITEHLHDYQGGHADILVNNAG
. :* . . :. * . : :
Mflv_1492|M.gilvum_PYR-GCK DVTGKMFEVEGGK-------------IRVAEGWAHGPEIDKG--------
Mvan_5264|M.vanbaalenii_PYR-1 DVTGRVFEVEGGK-------------IRVAEGWAHGPEIDKG--------
MSMEG_5999|M.smegmatis_MC2_155 DVTGKMFEVEGGK-------------IRIAEGWAHGPEIDKG--------
TH_3887|M.thermoresistible__bu DVSGRVFEVEGGK-------------VRVAEGWAHGPEADKG--------
Mb3578c|M.bovis_AF2122/97 DVTGKVFEVEGGI-------------IRVAEGWAHGPQVDKG--------
Rv3548c|M.tuberculosis_H37Rv DVTGKVFEVEGGI-------------IRVAEGWAHGPQVDKG--------
MMAR_5036|M.marinum_M AVTGKVFEVEGGV-------------IRVAEGWAHGPEVDKG--------
MUL_4110|M.ulcerans_Agy99 AVTGKVFEVEGGV-------------IRVAEGWAHGPEVDKG--------
MAB_0608|M.abscessus_ATCC_1997 EVTGKVFEVEAGI-------------IRVAEGWAHGPQVDKG--------
MAV_2639|M.avium_104 HVSGQVIRAMGEH-------------LQLLKGWHPVASVSNGE-------
MLBr_02565|M.leprae_Br4923 ITRDKLLANMDDCRWDAVLEVNLLAPQRLTEGLVGNGSISKGGRVIVLSS
. .::: :: :* . .:*
Mflv_1492|M.gilvum_PYR-GCK -----ARWDPAELGPVVGDLLAKARTPVPVYGA-----------------
Mvan_5264|M.vanbaalenii_PYR-1 -----ARWDPAELGSVVPDLLAKARTPVPVYGA-----------------
MSMEG_5999|M.smegmatis_MC2_155 -----ARWDPAELGPVVADLLAKARPAVPVYGA-----------------
TH_3887|M.thermoresistible__bu -----ARWDPAELGPVVRDLLAKARQPVPVYGAXXAPPSTACSRSTSALS
Mb3578c|M.bovis_AF2122/97 -----VKWDPAELGPVVSDLLAKSRPPVPVYGA-----------------
Rv3548c|M.tuberculosis_H37Rv -----VKWDPAELGPVVSDLLAKSRPPVPVYGA-----------------
MMAR_5036|M.marinum_M -----AKWDPAELRLVVADLLAKSRPSVPVYGA-----------------
MUL_4110|M.ulcerans_Agy99 -----AKWDPAELRLVVADLLAKSRPSVPVYGA-----------------
MAB_0608|M.abscessus_ATCC_1997 -----ARWDPSELGPVVTDLLDKARTPVPVYGSQG---------------
MAV_2639|M.avium_104 -----KRWDAQKLGAIMATDVFGTRNTGLRLGG-----------------
MLBr_02565|M.leprae_Br4923 MAGIAGNRGQTNYATAKAGVIGLTQALAPGLAEKGITINAVAPGFIETQM
. . : : :: .
Mflv_1492|M.gilvum_PYR-GCK --------------------------------------------------
Mvan_5264|M.vanbaalenii_PYR-1 --------------------------------------------------
MSMEG_5999|M.smegmatis_MC2_155 --------------------------------------------------
TH_3887|M.thermoresistible__bu LRQLVANV------------------------------------------
Mb3578c|M.bovis_AF2122/97 --------------------------------------------------
Rv3548c|M.tuberculosis_H37Rv --------------------------------------------------
MMAR_5036|M.marinum_M --------------------------------------------------
MUL_4110|M.ulcerans_Agy99 --------------------------------------------------
MAB_0608|M.abscessus_ATCC_1997 --------------------------------------------------
MAV_2639|M.avium_104 --------------------------------------------------
MLBr_02565|M.leprae_Br4923 TAAIPLATREVGRRLNSLLQGGLPVDVAETIAYFAIPASNAVTGNVIRVC
Mflv_1492|M.gilvum_PYR-GCK -------
Mvan_5264|M.vanbaalenii_PYR-1 -------
MSMEG_5999|M.smegmatis_MC2_155 -------
TH_3887|M.thermoresistible__bu -------
Mb3578c|M.bovis_AF2122/97 -------
Rv3548c|M.tuberculosis_H37Rv -------
MMAR_5036|M.marinum_M -------
MUL_4110|M.ulcerans_Agy99 -------
MAB_0608|M.abscessus_ATCC_1997 -------
MAV_2639|M.avium_104 -------
MLBr_02565|M.leprae_Br4923 GQAMLGA