For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VSYDAHLRVWRGDDDGGELIDFTVPVSEGEVVLDIIHRIQATQASDLAVRWNCKAGKCGSCSAEINGRPR LMCMTRMSTFDESETITVTPLRAFPVIRDLVTDVSFNYQKAREVQSFTPPKGLKPGDYRMQQEDVERSQE FRKCIECFLCQNTCHAVRDHEENKKAFAGPRYLMRVAELEMHPLDVADRREVAQEELGLGYCNITKCCTE VCPEGIHITDNALIPMKERVADRKYDPIVWLGNKLFRR
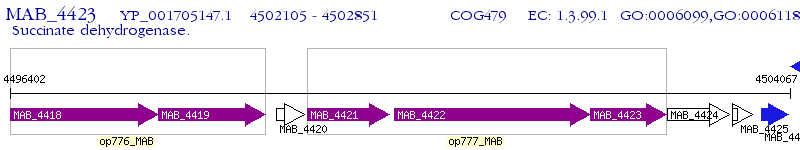
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. abscessus ATCC 19977 | MAB_4423 | - | - | 100% (248) | fumarate reductase iron-sulfur subunit |
| M. abscessus ATCC 19977 | MAB_3676 | sdhB | 5e-18 | 27.91% (215) | succinate dehydrogenase iron-sulfur subunit |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0253c | - | 1e-121 | 80.65% (248) | fumarate reductase iron-sulfur subunit |
| M. gilvum PYR-GCK | Mflv_0395 | - | 1e-123 | 82.66% (248) | fumarate reductase iron-sulfur subunit |
| M. tuberculosis H37Rv | Rv0247c | - | 1e-121 | 80.65% (248) | fumarate reductase iron-sulfur subunit |
| M. leprae Br4923 | MLBr_00696 | sdhB | 1e-16 | 28.64% (206) | succinate dehydrogenase iron-sulfur subunit |
| M. marinum M | MMAR_0510 | sdhB_1 | 1e-123 | 81.45% (248) | succinate dehydrogenase (iron-sulfur subunit), SdhB_1 |
| M. avium 104 | MAV_4910 | - | 1e-124 | 81.45% (248) | fumarate reductase iron-sulfur subunit |
| M. smegmatis MC2 155 | MSMEG_0417 | - | 1e-126 | 83.40% (247) | fumarate reductase iron-sulfur subunit |
| M. thermoresistible (build 8) | TH_1916 | - | 1e-122 | 80.72% (249) | PROBABLE SUCCINATE DEHYDROGENASE |
| M. ulcerans Agy99 | MUL_1171 | sdhB_1 | 1e-123 | 81.85% (248) | fumarate reductase iron-sulfur subunit |
| M. vanbaalenii PYR-1 | Mvan_0283 | - | 1e-120 | 80.65% (248) | fumarate reductase iron-sulfur subunit |
CLUSTAL 2.0.9 multiple sequence alignment
Mb0253c|M.bovis_AF2122/97 ----------------------MTYSASMRVWRGDE-----SCGELREFT
Rv0247c|M.tuberculosis_H37Rv ----------------------MTYSASMRVWRGDE-----SCGELREFT
MMAR_0510|M.marinum_M ----------------------MGYNASLRVWRGDE-----SNGELRDFT
MUL_1171|M.ulcerans_Agy99 ----------------------MGYNASLRVWRGDE-----SNGELRDFT
MAV_4910|M.avium_104 ----------------------MTYNAKMRVWRGDD-----ANGALQDFT
TH_1916|M.thermoresistible__bu ----------------------MGYQAKLRVWRGDD-----DGGALQDYS
Mflv_0395|M.gilvum_PYR-GCK ---------------------MAAYNAKLRVWRGDD-----TGGELKDFT
Mvan_0283|M.vanbaalenii_PYR-1 ---------------------MAAYQGKLRVWRGDD-----TGGGLQDFT
MSMEG_0417|M.smegmatis_MC2_155 ---------------------MATYDAKLRVWRGDD-----TGGELHDYT
MAB_4423|M.abscessus_ATCC_1997 ----------------------MSYDAHLRVWRGDD-----DGGELIDFT
MLBr_00696|M.leprae_Br4923 MAVEPVEVKSLEPPRPPISDDAVMVTVKIARFNPDNPDAFAETGGWQSFR
: :. *: * .:
Mb0253c|M.bovis_AF2122/97 VEVNEGEVVLDVILRLQQTQTPDLAVRWNCKAGKCGSCSAEINGKPRLMC
Rv0247c|M.tuberculosis_H37Rv VEVNEGEVVLDVILRLQQTQTPDLAVRWNCKAGKCGSCSAEINGKPRLMC
MMAR_0510|M.marinum_M VEVNEGEVVLDVIHRLQQTQTPDLAVRWNCKAGKCGSCSAEINGKPRLMC
MUL_1171|M.ulcerans_Agy99 VEVNEGEVVLDVIHRLQQTQTPDLAVRWNCKAGKCGSCSAEINGKPRLMC
MAV_4910|M.avium_104 VEVNEGEVVLDIIHRLQQTQTPDLAVRWNCKAGKCGSCSAEINGYPRLLC
TH_1916|M.thermoresistible__bu VEVNEGEVVLDIIHRLQQTQTPDLAVRWNCKAGKCGSCSAEINGRPRLMC
Mflv_0395|M.gilvum_PYR-GCK VEVNEGEVVLDIIHRLQATQTPDLAVRWNCKAGKCGSCSAEINGRPRLLC
Mvan_0283|M.vanbaalenii_PYR-1 VEVNDGEVVLDIIHRLQATQTPDLAVRWNCKAGKCGSCSAEINGRPRLLC
MSMEG_0417|M.smegmatis_MC2_155 VEVNDGEVVLDIIHRLQATQTPDLAVRWNCKAGKCGSCSAEINGRPRLMC
MAB_4423|M.abscessus_ATCC_1997 VPVSEGEVVLDIIHRIQATQASDLAVRWNCKAGKCGSCSAEINGRPRLMC
MLBr_00696|M.leprae_Br4923 VPCLPTDRLLNLLIYIKGYLDGTLTFRRSCAHGVCGSDAMRVNGVNRLAC
* : :*::: :: *:.* .* * *** : .:** ** *
Mb0253c|M.bovis_AF2122/97 MTRMSTFD-----EDEIVTVTPMRTFPVIRDLVTDVSFNYQKAREIPSFA
Rv0247c|M.tuberculosis_H37Rv MTRMSTFD-----EDEIVTVTPMRTFPVIRDLVTDVSFNYQKAREIPSFA
MMAR_0510|M.marinum_M MTRMSTFA-----EDEIVTVTPLRTFPVIRDLVTDVSFNYEKAREIPSFA
MUL_1171|M.ulcerans_Agy99 MTRMSTFA-----EDEIVTVTPLRTFPVIRDLVTDVSFNYEKAREIPSFA
MAV_4910|M.avium_104 MTRMSTFA-----EDEVVTVTPLRTFPVIRDLVTDVSFNYEKAREIPSFA
TH_1916|M.thermoresistible__bu MTRMSTFA-----ENEVITVTPMRTFPVIRDLVNDVSFNFEKAREIPSFR
Mflv_0395|M.gilvum_PYR-GCK MTRMSTFG-----EDEVVTVTPLRTFPVMRDLVTDVSFNYEKAREIPAFT
Mvan_0283|M.vanbaalenii_PYR-1 MTRMSTFG-----EDEVVTVTPLRTFPVMRDLVTDVSFNYEKAREIPSFA
MSMEG_0417|M.smegmatis_MC2_155 MTRMSTFG-----EDEVVTVTPLRTFPVMRDLVTDVSFNYEKARQIPSFT
MAB_4423|M.abscessus_ATCC_1997 MTRMSTFD-----ESETITVTPLRAFPVIRDLVTDVSFNYQKAREVQSFT
MLBr_00696|M.leprae_Br4923 KVLMRDLLPKKAGKNLTITVEPIRGLPVEKDLVVDMEPFFDAYRAVKPYL
. * : :. :** *:* :** :*** *:. :: * : .:
Mb0253c|M.bovis_AF2122/97 PPKELQPSEYRMAQ-VDVARSQEFRKCIECFLCQNVCHVVRDHEENKDAF
Rv0247c|M.tuberculosis_H37Rv PPKELQPSEYRMAQ-VDVARSQEFRKCIECFLCQNVCHVVRDHEENKDAF
MMAR_0510|M.marinum_M PPKELQPGEYRMAQ-VDVARSQEFRKCIECFLCQNVCHVIRDHEENKKAF
MUL_1171|M.ulcerans_Agy99 PPKELQPGEYRMAQ-VDVARSQEFRKCIECFLCQNVCHVIRDHEENKKAF
MAV_4910|M.avium_104 PPKDLQPGEYRMAQ-EDVQRSQEFRKCIECFLCQNVCHVIRDHDENKKAF
TH_1916|M.thermoresistible__bu PPKDLQPGEYRMQQ-EDVNRSQEFRKCIECFLCQNVCHVIRDHEDNKKAF
Mflv_0395|M.gilvum_PYR-GCK PPKDLQPGEYRMQQ-EDVNRSQEFRKCIECFLCQNVCHVVRDHEENKLAF
Mvan_0283|M.vanbaalenii_PYR-1 PPKNLQPGEYRMAQ-EDVNRSQEFRKCIECFLCQNVCHVNRDHEENKLAF
MSMEG_0417|M.smegmatis_MC2_155 PPKDLQPGEYRMQQ-EDVNRSQEFRKCIECFLCQNVCHVVRDHEENKENF
MAB_4423|M.abscessus_ATCC_1997 PPKGLKPGDYRMQQ-EDVERSQEFRKCIECFLCQNTCHAVRDHEENKKAF
MLBr_00696|M.leprae_Br4923 ITSGNPPTRERIQSPTDRARYDDTTKCILCACCTTSCPVFWSEG----TY
.. * *: . * * :: *** * * . * . .. :
Mb0253c|M.bovis_AF2122/97 AGPRFLMRIAELEMHPLDTR--DRRSQAQEEHGLGYCNITKCCTEVCPEN
Rv0247c|M.tuberculosis_H37Rv AGPRFLMRIAELEMHPLDTR--DRRSQAQEEHGLGYCNITKCCTEVCPEN
MMAR_0510|M.marinum_M AGPRFLMRIAELEMHPLDTR--DRRNEAQEAHGLGYCNITKCCTEVCPEN
MUL_1171|M.ulcerans_Agy99 AGPRFLMRVAELEMHPLDTR--DRRNEAQEAHGLGYCNITKCCTEVCPEN
MAV_4910|M.avium_104 AGPRFLMRIAELEMHPLDTR--DRRNDAQDEHGLGYCNITKCCTEVCPEN
TH_1916|M.thermoresistible__bu AGPRFLMRMAELDMHPLDTHP-NRAEEVQDEHGLGYCNITKCCTEVCPEH
Mflv_0395|M.gilvum_PYR-GCK SGPRFLMRQAELDMHPLDTHG-HRSEQAQEENGLGYCNITKCCTEVCPEH
Mvan_0283|M.vanbaalenii_PYR-1 SGPRYLMRQAELDMHPLDTHG-HRAEQAQDENGLGYCNITKCCTEVCPEH
MSMEG_0417|M.smegmatis_MC2_155 AGPRFHMRIAELDMHPLDTVD--RKEMAQDEFGLGYCNITKCCTEVCPEH
MAB_4423|M.abscessus_ATCC_1997 AGPRYLMRVAELEMHPLDVA--DRREVAQEELGLGYCNITKCCTEVCPEG
MLBr_00696|M.leprae_Br4923 YGPAAIVNAHRFIFDSRDEGAGERLTLLNEVDGVWRCRTTFNCTESCPRN
** :. .: :.. * * :: *: *. * *** **.
Mb0253c|M.bovis_AF2122/97 IKITDNALIPMKERVADRKYDPVVWLGSKLFRR-
Rv0247c|M.tuberculosis_H37Rv IKITDNALIPMKERVADRKYDPVVWLGSKLFRR-
MMAR_0510|M.marinum_M IKITDNALIPMKERVVDRKYDPVVWLGNKLFRR-
MUL_1171|M.ulcerans_Agy99 IKITDNALIPMKERVVDRKYDPVVWLGNKLFRR-
MAV_4910|M.avium_104 IKITDNALIPMKERVADRKYDPVVWLGNKLFRR-
TH_1916|M.thermoresistible__bu IKITDNALIPMKERVADRKYDPVVWLGNKLFRRQ
Mflv_0395|M.gilvum_PYR-GCK IKITDNALIPMKERAADRKYDPVVWLGNKLFRRK
Mvan_0283|M.vanbaalenii_PYR-1 IKITDNALIPMKERAADRKYDPVVWLGNKLFRRR
MSMEG_0417|M.smegmatis_MC2_155 IKITDNALIPMKERVADRKYDPIVWLGNKLFRR-
MAB_4423|M.abscessus_ATCC_1997 IHITDNALIPMKERVADRKYDPIVWLGNKLFRR-
MLBr_00696|M.leprae_Br4923 IEVTKAIQEVKRALMFSR----------------
*.:*. : .*