For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
GAQRASMQRPAADTPDGFGVAVVREEGRWRCSPMGPKALTSLRAAETELRELRSAGAVFGLLDVDDEFFV IVRPAPSGTRLLLSDATAALDYDIAAEVLDNLDAEIDPEDLEDADPFEEGDLGLLSDIGLPEAVLGVILD ETDLYADEQLGRIAREMGFADQLSAVIDRLGR*
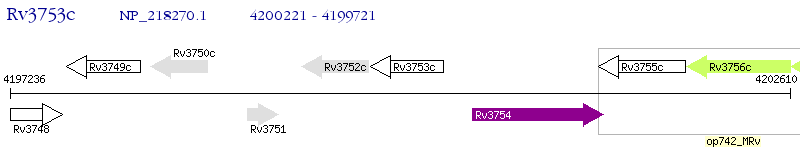
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. tuberculosis H37Rv | Rv3753c | - | - | 100% (166) | hypothetical protein Rv3753c |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3779c | - | 2e-90 | 100.00% (165) | hypothetical protein Mb3779c |
| M. gilvum PYR-GCK | Mflv_1213 | - | 2e-72 | 80.72% (166) | hypothetical protein Mflv_1213 |
| M. leprae Br4923 | MLBr_02473 | - | 3e-77 | 88.61% (158) | hypothetical protein MLBr_02473 |
| M. abscessus ATCC 19977 | MAB_0267 | - | 3e-62 | 69.88% (166) | hypothetical protein MAB_0267 |
| M. marinum M | MMAR_5296 | - | 5e-81 | 88.96% (163) | hypothetical protein MMAR_5296 |
| M. avium 104 | MAV_0323 | - | 1e-83 | 93.83% (162) | hypothetical protein MAV_0323 |
| M. smegmatis MC2 155 | MSMEG_6328 | - | 2e-75 | 82.42% (165) | hypothetical protein MSMEG_6328 |
| M. thermoresistible (build 8) | TH_1654 | - | 3e-69 | 76.36% (165) | CONSERVED HYPOTHETICAL PROTEIN |
| M. ulcerans Agy99 | MUL_4371 | - | 3e-81 | 88.96% (163) | hypothetical protein MUL_4371 |
| M. vanbaalenii PYR-1 | Mvan_5595 | - | 4e-75 | 83.73% (166) | hypothetical protein Mvan_5595 |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_1213|M.gilvum_PYR-GCK MGAQSAQKQERVTDVPDGFGVAVVREDGKWRCTAMQKAALTSLTAAETEL
Mvan_5595|M.vanbaalenii_PYR-1 MGAQSAQKQERVTDVPDGFAVAVVREDGKWRCTPMRPATLTSLTAAETEL
Rv3753c|M.tuberculosis_H37Rv -------MQRPAADTPDGFGVAVVREEGRWRCSPMGPKALTSLRAAETEL
Mb3779c|M.bovis_AF2122/97 MGAQRASTQRPAADTPDGFGVAVVREEGRWRCSPMGPKALTSLRAAETEL
MMAR_5296|M.marinum_M MGAQRAAAKGPSAETPDGFGVAVVREEGKWRCAAMSPKALATLSEAETEL
MUL_4371|M.ulcerans_Agy99 MGAQRAAAKGPSAETPDGFGVAVVREEGKWRCAAMSPKALATLSEAETEL
MLBr_02473|M.leprae_Br4923 --------------MPDGFGVAVVREEGQWRCSAMASKSLTSLTAAETEL
MAV_0323|M.avium_104 MGAQRASAQGLSADTPDGFGVAVVREEGQWRCAPMAPKSLTSLSAAETEL
MSMEG_6328|M.smegmatis_MC2_155 MGAQRAPADLP-----DGFGVAVVREDGKWRCAPMRKSALNSLSAAETEL
TH_1654|M.thermoresistible__bu MGAQGAPARKPGADDIEGFGVAVVREDGRWQCFPMRRAALRSLAAAEREL
MAB_0267|M.abscessus_ATCC_1997 MDAQRAKAAGPSAESLEGFGVAVVREEGRWRVTSLKARALTDLAAAETEL
:**.******:*:*: .: :* * ** **
Mflv_1213|M.gilvum_PYR-GCK RELRSAGAVFGILDIDDEFFVILRPAPSGTRLLLSDATAALDYDIAAEVL
Mvan_5595|M.vanbaalenii_PYR-1 RELRSAGAVFGILDIDDEFFVIVRPAPSGTRLLLSDATAALDYDIAAEVL
Rv3753c|M.tuberculosis_H37Rv RELRSAGAVFGLLDVDDEFFVIVRPAPSGTRLLLSDATAALDYDIAAEVL
Mb3779c|M.bovis_AF2122/97 RELRSAGAVFGLLDVDDEFFVIVRPAPSGTRLLLSDATAALDYDIAAEVL
MMAR_5296|M.marinum_M RELRSSGAVFGLLDIDDEFFVILRPAPSGTRLLLSDATAALDYDIAAEVL
MUL_4371|M.ulcerans_Agy99 RELRSSGAVFGLLDIDDEFFVILRPAPSGTRLLLSDATAALDYDIAAEVL
MLBr_02473|M.leprae_Br4923 RELRSVGAVFGLLDVDDEFFIILRPAPSGTRLLLSDATAALDYDIAAEIL
MAV_0323|M.avium_104 RELRSSGAVFGLLDVDDEFFVIVRPAPSGTRLLLSDATAALDYDIAAEVL
MSMEG_6328|M.smegmatis_MC2_155 REIRSAGAVFGLLDVDDEFFVIVRPAPAGTRLLLSDATAALDYDIAAEAL
TH_1654|M.thermoresistible__bu CEMRAAGPVFGLLDIDDEFFIIVRPAPSGTRLLLSDATAALDYDIAAEAL
MAB_0267|M.abscessus_ATCC_1997 RELRSAGAVFGLLDVDEEFFIIIRPAPSGTHLLISDATAAIDYDIAVEVL
*:*: *.***:**:*:***:*:****:**:**:******:*****.* *
Mflv_1213|M.gilvum_PYR-GCK EKLDA---DIDDDELERCDPFEEGDLAVLSDIGLPEAVLGVILDESDDLY
Mvan_5595|M.vanbaalenii_PYR-1 EKLDA---DIDDEELERCDPFEEGDLGLLSDIGLPEAVLGVILDESDDLY
Rv3753c|M.tuberculosis_H37Rv DNLDA---EIDPEDLEDADPFEEGDLGLLSDIGLPEAVLGVILDET-DLY
Mb3779c|M.bovis_AF2122/97 DNLDA---EIDPEDLEDADPFEEGDLGLLSDIGLPEAVLGVILDET-DLY
MMAR_5296|M.marinum_M DSLDS---EIDPEDLEDADPFEEGDLGLLADIGLPEAVLGVILDET-DLY
MUL_4371|M.ulcerans_Agy99 DSLDS---EIDPEDLEDADPFEEGDLGLLADIGLPEAVLGVILDET-DLY
MLBr_02473|M.leprae_Br4923 DSLDA---EIDPEDLEDAYPFEEGDLGLLSDVGLPEATLGVILDQT-DLY
MAV_0323|M.avium_104 DKLDA---DIDPEDLEDSDPFEEGDLGLLSDIGLPEAVLGVILDET-DLY
MSMEG_6328|M.smegmatis_MC2_155 EKLDA---DIEADDLDDADPFEEGDLGLLSDLGLPDAVLSVILDET-DLY
TH_1654|M.thermoresistible__bu ERLDA---DIDEDDLESADPFEEGDLGVLADIGLPDAVLSVILDEI-DLY
MAB_0267|M.abscessus_ATCC_1997 ERLNIPVPELELDELDEVDPWEEGDLGLLSDIGLPEPVLSVILSET-DLY
: *: ::: ::*: *:*****.:*:*:***:..*.***.: ***
Mflv_1213|M.gilvum_PYR-GCK ADEQVGRIAREMGFADELSALLDRLGR
Mvan_5595|M.vanbaalenii_PYR-1 ADEQVGRIAREMGFADELSAVLDRLGR
Rv3753c|M.tuberculosis_H37Rv ADEQLGRIAREMGFADQLSAVIDRLGR
Mb3779c|M.bovis_AF2122/97 ADEQLGRIAREMGFADQLSAVIDRLGR
MMAR_5296|M.marinum_M ADEQLGRIAREMGFAEQLSAIIDRLGR
MUL_4371|M.ulcerans_Agy99 ADEQLGRIAREMGFAEQLSAIIDRLGR
MLBr_02473|M.leprae_Br4923 ADEQLGHIAREMGFAEQLSAVINRLGR
MAV_0323|M.avium_104 ADEQLGRIAREMGFADQLSAVIDRLGR
MSMEG_6328|M.smegmatis_MC2_155 ADEQIGRIAREMGFAEELSAVLDRLNR
TH_1654|M.thermoresistible__bu ADEQLGRIAGEMGFAEELAAVLERLGR
MAB_0267|M.abscessus_ATCC_1997 PDEQLGMIAARMGFADEFGAVLDKLDR
.***:* ** .****:::.*::::*.*