For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
APATRYPAGRSHPGHALFAQFAYPPNELGYCGPQDAGALLRAGGQTVTVAREFDGAWPYLQAIADCAGVA DPLDVEVVRSYWVGGPRLGQVDAAMLLDRLRSAFAGQVTGLLADIDAQATVLAHHSFHVCVVYPWVRFLD RDPATPLRVLQSCRIRWGTVVSVDTEHVMLESPPLTFGTGRLSLGDPATERVRWRRDGISLTAAPRSGDV VAAHWDWVCGPLTGTEATALMTATQSTLDLVNTARMRS*
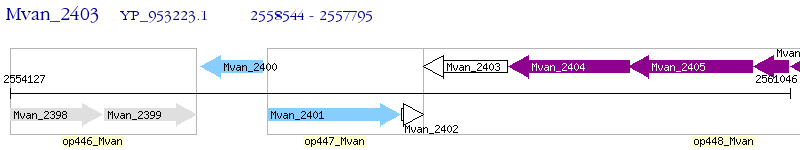
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. vanbaalenii PYR-1 | Mvan_2403 | - | - | 100% (249) | hypothetical protein Mvan_2403 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | - | - | - | - | - |
| M. gilvum PYR-GCK | - | - | - | - | - |
| M. tuberculosis H37Rv | - | - | - | - | - |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | - | - | - | - | - |
| M. marinum M | MMAR_1856 | - | 2e-36 | 37.18% (234) | hypothetical protein MMAR_1856 |
| M. avium 104 | - | - | - | - | - |
| M. smegmatis MC2 155 | MSMEG_2710 | - | 2e-71 | 57.76% (232) | hypothetical protein MSMEG_2710 |
| M. thermoresistible (build 8) | TH_4124 | - | 3e-48 | 41.30% (247) | PUTATIVE hypothetical protein MMAR_1856 |
| M. ulcerans Agy99 | - | - | - | - | - |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_1856|M.marinum_M --------MSVPDTPDIRGPEMFARYAYAPNALGYCGPPLGATLRDGSVA
TH_4124|M.thermoresistible__bu VTAVGPPTAAVPTGAESDGVLMFARYAYAPNQLGYCGPVESAALRSGSEQ
Mvan_2403|M.vanbaalenii_PYR-1 ----MAPATRYPAGRSHPGHALFAQFAYPPNELGYCGPQDAGALLRAG-G
MSMEG_2710|M.smegmatis_MC2_155 ----MVVHSPLPAG-----YELFTRYAFPPNELGYCGPD--GAAEPA---
* :*:::*:.** ****** .: .
MMAR_1856|M.marinum_M DVRAAARKFSGAWPYLRVLSRLTGIADPLDYRLVEAYWLGGGVGADVEPG
TH_4124|M.thermoresistible__bu RIRAAARRFSGAWPYLQVLARLTGIADPLDSRLVRAYWLGGGVCDQLDPA
Mvan_2403|M.vanbaalenii_PYR-1 QTVTVAREFDGAWPYLQAIADCAGVADPLDVEVVRSYWVGGPRLGQVDAA
MSMEG_2710|M.smegmatis_MC2_155 ELAFHAPEFDGAWPYLRAIAHAIG-AEPLAEDVVAGYWVGGPVLNRVDPA
* .*.******:.:: * *:** :* .**:** ::..
MMAR_1856|M.marinum_M RFFDELLAIIGPQAGRYWSHLTPELAHEAAGNHCFHVFGVYPWTRFLGRG
TH_4124|M.thermoresistible__bu EFYAELLAVIGPQAGHYWTHLTPELAAEATGNHCFHVFGVYPWSRLLGRG
Mvan_2403|M.vanbaalenii_PYR-1 MLLDRLRSAFAGQVTGLLADID--AQATVLAHHSFHVCVVYPWVRFLDR-
MSMEG_2710|M.smegmatis_MC2_155 DLLADLRTAFAGQVMGLLDDVP--VSESVLAHHSFHVFVVYPWIRFLDR-
: * : :. *. .: . .:*.*** **** *:*.*
MMAR_1856|M.marinum_M LDEHPLSVLDSCRITWATVVSRTGDAVEVSRQSLIWDGEALALAAPSSQL
TH_4124|M.thermoresistible__bu MDEHPVHVLDSCRITPAEVLDRDGDQIVVRSRPLLFEDRRLRLGEPTTRT
Mvan_2403|M.vanbaalenii_PYR-1 DPATPLRVLQSCRIRWGTVVSVDTEHVMLESPPLTFGTGRLSLGDPATER
MSMEG_2710|M.smegmatis_MC2_155 NPEPALHVLQNCRIRWGTVVEVEGEHVKLESRPLVRNGRRIELDSAVAEY
.: **:.*** . *:. : : : .* : * . :.
MMAR_1856|M.marinum_M LDVWADGYSAVPDVSVGEEVAVHWGRLCARLQPEQVVALAESTHRQLRVT
TH_4124|M.thermoresistible__bu VAVSVDGHSALPDVAAGDRIALHWDWICGRLEPEQIASLDTSTERQLELT
Mvan_2403|M.vanbaalenii_PYR-1 VRWRRDGISLTAAPRSGDVVAAHWDWVCGPLTGTEATALMTATQSTLDLV
MSMEG_2710|M.smegmatis_MC2_155 VRWSRGGASLIAAPRPGDTVSAHWDWVCDTLSPHDVTALTESTLTTLGVV
: .* * . *: :: **. :* * : .:* :* * :.
MMAR_1856|M.marinum_M NQRLARV---
TH_4124|M.thermoresistible__bu NRRLGVELS-
Mvan_2403|M.vanbaalenii_PYR-1 NTARMRS---
MSMEG_2710|M.smegmatis_MC2_155 NALREVKEEA
*