For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VISAPDLIETFPFPFAADTYRYTTNVEPAGSSVHTPAGRWGERVVDIDSEYETELAERAEILAADPTRYA VLPHMRVACWDAMLTLMREMSSAYPDTMSLERHGAGWRWRNDRLGITQDFVIGDETTLPAEPLAYIGGQV QEDIVLLDQRDGDLFGDAGVVTFAADWSFGFDVGMTFLEIHGPVPRLRQTGVITRAREFLMRLQPGETYR RTNWTLTIDRRLDVSTERYCEWGPDRTLIQTVPDDEFGRRVHLRVEVQHLIRLPESGSICFLIRSYMLPL ADLAAFEPWRQRAAAVLAELPDDIADYKGIIAYRDRAVAWLRNAG
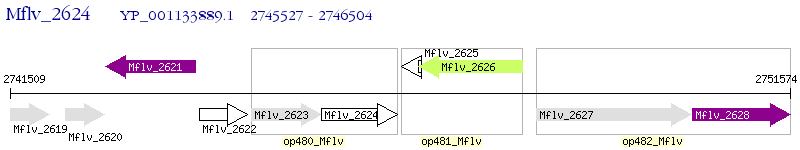
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. gilvum PYR-GCK | Mflv_2624 | - | - | 100% (325) | hypothetical protein Mflv_2624 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | - | - | - | - | - |
| M. tuberculosis H37Rv | - | - | - | - | - |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | - | - | - | - | - |
| M. marinum M | - | - | - | - | - |
| M. avium 104 | - | - | - | - | - |
| M. smegmatis MC2 155 | MSMEG_4637 | - | 1e-156 | 80.94% (320) | hypothetical protein MSMEG_4637 |
| M. thermoresistible (build 8) | - | - | - | - | - |
| M. ulcerans Agy99 | - | - | - | - | - |
| M. vanbaalenii PYR-1 | Mvan_3957 | - | 1e-176 | 91.08% (325) | hypothetical protein Mvan_3957 |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_2624|M.gilvum_PYR-GCK ------MISAPDLIETFPFPFAADTYRYTTNVEPAGSSVHTPAGRWGERV
Mvan_3957|M.vanbaalenii_PYR-1 ------MISAPDLIETFPFPFAADEYRYSTNVEPAGAAVPTPAGRWGERV
MSMEG_4637|M.smegmatis_MC2_155 MTITPERLSAPDLVETFPFPFPTDRYRYSTNVEPAGGAVSTPAGQWGERV
:*****:*******.:* ***:*******.:* ****:*****
Mflv_2624|M.gilvum_PYR-GCK VDIDSEYETELAERAEILAADPTRYAVLPHMRVACWDAMLTLMREMSSAY
Mvan_3957|M.vanbaalenii_PYR-1 VDLDSEYETELAERAAILAADPTRYAVLPHMRPACWDAMLTLMRAMASAY
MSMEG_4637|M.smegmatis_MC2_155 VDIDSEYEHELAQRAEILAADPTRYAVLPHMRAACWDAMVTLMREMATAY
**:***** ***:** **************** ******:**** *::**
Mflv_2624|M.gilvum_PYR-GCK PDTMSLERHGAGWRWRNDRLGITQDFVIGDETTLPAEPLAYIGGQVQEDI
Mvan_3957|M.vanbaalenii_PYR-1 PDTMSLQRHGTGWRWRNDRLGLAQDFVIGDEASLPAEPLAYIAGQVQEDI
MSMEG_4637|M.smegmatis_MC2_155 PDTMSLTESDGVWHWENKRLGISQSFTYGDESTLPSEPLAYIAGQVQDDI
****** . . *:*.*.***::*.*. ***::**:******.****:**
Mflv_2624|M.gilvum_PYR-GCK VLLDQRDGDLFGDAGVVTFAADWSFGFDVGMTFLEIHGPVPRLRQTGVIT
Mvan_3957|M.vanbaalenii_PYR-1 VLLDQRDGDLFGDAGVVTFAADWSFGFDVGMTFLEIHGPVPRLRETGVIT
MSMEG_4637|M.smegmatis_MC2_155 VLLDQRDDDLFGDAGVVTFAADWSFGFDVGMTFLEIHGPVPRLRETGVIT
*******.************************************:*****
Mflv_2624|M.gilvum_PYR-GCK RAREFLMRLQPGETYRRTNWTLTIDRRLDVSTERYCEWGPDRTLIQTVPD
Mvan_3957|M.vanbaalenii_PYR-1 RAREFLMRLQPGETYRRTNWSLTIGRRLDVSTELYHEWGPDRTAIQTVSD
MSMEG_4637|M.smegmatis_MC2_155 RARQFLMRLQPGETYRRTNWTLTIGRKLDVSTERYPEWGPDRAMINGVDD
***:****************:***.*:****** * ******: *: * *
Mflv_2624|M.gilvum_PYR-GCK DEFGRRVHLRVEVQHLIRLPESGSICFLIRSYMLPLADLAAFEPWRQRAA
Mvan_3957|M.vanbaalenii_PYR-1 EEFGRLVHLRVEVQHLIRLPESGSICFLIRTYMLPFADLAAFEPWRQRAA
MSMEG_4637|M.smegmatis_MC2_155 ETFGRLVHLRVELQHLIRLPESGAICFLIRTYMLPFADIATVGPWRDRLA
: *** ******:**********:******:****:**:*:. ***:* *
Mflv_2624|M.gilvum_PYR-GCK AVLAELPDDIADYKGIIAYRDRAVAWLRNAG
Mvan_3957|M.vanbaalenii_PYR-1 AVLAELPDDMADYKGIIAYRDRAVAWLRNAG
MSMEG_4637|M.smegmatis_MC2_155 AVLADLPDDMADYKGIIGYRDRAVEWLARH-
****:****:*******.****** ** .