For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
VRTSGRLAARGLIYVIELYRHMVSPLRPASCRFIPTCSQYAVDALSEYGLIRGSWLTVARLAKCGPWHQG GWDPIPERQGCRADEAGCSPAVKQGESEFFV*
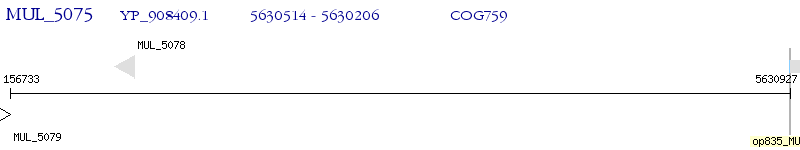
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. ulcerans Agy99 | MUL_5075 | - | - | 100% (102) | hypothetical protein MUL_5075 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3953c | - | 6e-41 | 67.27% (110) | hypothetical protein Mb3953c |
| M. gilvum PYR-GCK | Mflv_0830 | - | 3e-31 | 63.44% (93) | hypothetical protein Mflv_0830 |
| M. tuberculosis H37Rv | Rv3922c | - | 6e-41 | 67.27% (110) | hypothetical protein Rv3922c |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | - | - | - | - | - |
| M. marinum M | MMAR_5578 | - | 1e-58 | 100.00% (102) | hypothetical protein MMAR_5578 |
| M. avium 104 | MAV_5311 | - | 3e-40 | 75.82% (91) | hypothetical protein MAV_5311 |
| M. smegmatis MC2 155 | MSMEG_6944 | - | 2e-32 | 73.42% (79) | hypothetical protein MSMEG_6944 |
| M. thermoresistible (build 8) | TH_0760 | - | 1e-31 | 66.67% (84) | POSSIBLE HEMOLYSIN |
| M. vanbaalenii PYR-1 | Mvan_6076 | - | 6e-31 | 57.58% (99) | hypothetical protein Mvan_6076 |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_0830|M.gilvum_PYR-GCK ---------MSGRS---AVVRAVVYVIQLYRHTISPLRLPTCRFTPTCSQ
Mvan_6076|M.vanbaalenii_PYR-1 ---------MTARS---RAVRAVVYLIQLYRHTISPLRLPTCRFTPTCSQ
MSMEG_6944|M.smegmatis_MC2_155 ---------MRRRIG-VRAVRAVIYVIQLYRHTISPLRLPTCRFMPTCSQ
TH_0760|M.thermoresistible__bu --------VKTLRAAGSASARALIFLIRLYQHTISPLRLPTCRFTPTCSQ
MUL_5075|M.ulcerans_Agy99 ----------MVRTSGRLAARGLIYVIELYRHMVSPLRPASCRFIPTCSQ
MMAR_5578|M.marinum_M ----------MVRTSGRLAARGLIYVIELYRHMVSPLRPASCRFIPTCSQ
Mb3953c|M.bovis_AF2122/97 MSLSRQSCGRVVRVTGRASARGLIFVIQVYRHMLSPLRPASCRFVPTCSQ
Rv3922c|M.tuberculosis_H37Rv MSLSRQSCGRVVRVTGRASARGLIFVIQVYRHMLSPLRPASCRFVPTCSQ
MAV_5311|M.avium_104 MTGPARS-GAAIRGAGRTVARGLIFLIQLYRHMVSPLRPATCRFVPTCSQ
* .*.::::*.:*:* :**** .:*** *****
Mflv_0830|M.gilvum_PYR-GCK YAVDALTEFGFFRGSWLALIRLLKCGPWHRGGWDPIPERTSHSDATADGA
Mvan_6076|M.vanbaalenii_PYR-1 YAVDALTEFGLFRGGWLALVRLLKCGPWHNGGWDPIPDRESHSDATGEGP
MSMEG_6944|M.smegmatis_MC2_155 YAVDALTEYGLIRGTWLATVRLLKCGPWHQGGWDPIPERCAHETETVAGV
TH_0760|M.thermoresistible__bu YAVDALTEYGLLRGIGLTVVRLLKCGPWHRGGWDPIPER--REVRRDAGT
MUL_5075|M.ulcerans_Agy99 YAVDALSEYGLIRGSWLTVARLAKCGPWHQGGWDPIPE----------RQ
MMAR_5578|M.marinum_M YAVDALSEYGLIRGSWLTVARLAKCGPWHQGGWDPIPE----------RQ
Mb3953c|M.bovis_AF2122/97 YAVDALTEYGLLRGSWLTMIRLAKCGPWHRGGWDPIPEG------LTTGR
Rv3922c|M.tuberculosis_H37Rv YAVDALTEYGLLRGSWLTMIRLAKCGPWHRGGWDPIPEG------LTTGR
MAV_5311|M.avium_104 YAVDALDEYGLIRGSWLAAARLTKCGPWHQGGWDPIPE----------RP
****** *:*::** *: ** ******.*******:
Mflv_0830|M.gilvum_PYR-GCK SCAAEVCDTADHRVAPVQQGKSHTSVV
Mvan_6076|M.vanbaalenii_PYR-1 CAAAVTCGTAEHRNRPVQQGKSQTSVV
MSMEG_6944|M.smegmatis_MC2_155 SAETAAAENPVWET-PAKRGESKSRV-
TH_0760|M.thermoresistible__bu TATADRVR-------------SKVRV-
MUL_5075|M.ulcerans_Agy99 GCRADEAGCS----PAVKQGESEFFV-
MMAR_5578|M.marinum_M GCRADEAGCS----PAVKQGESEFFV-
Mb3953c|M.bovis_AF2122/97 SCQTDVDGANDDWNPASKRGERESFV-
Rv3922c|M.tuberculosis_H37Rv SCQTDVDGANDDWNPASKRGERESFV-
MAV_5311|M.avium_104 GCRVNCQDASDAWAVRATRGESGSLV-
. . *