For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
GWPSPPPNASKAPAASATPGAGSYPSGGLSHLDERGAAHMVDATEKGATRRTAVAAGFLRTSTEVVALIS SGGLPKGDALATARVAGILAAKRTSDLIPLCHQLALTGVDVDFTMAEADVEITATVRSTDRTGVEMEALT AVIVVALTLYDMIKAVDPAARVDDVRVLSKQGGRSGSWSR*
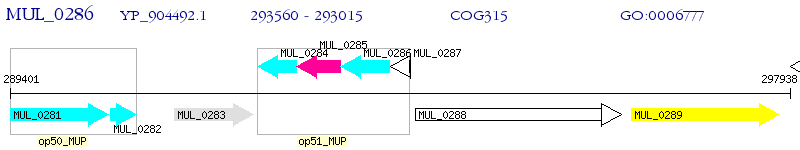
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. ulcerans Agy99 | MUL_0286 | moaC2 | - | 100% (181) | molybdenum cofactor biosynthesis protein C 2 MoaC2 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0888 | moaC | 4e-62 | 82.27% (141) | molybdenum cofactor biosynthesis protein C |
| M. gilvum PYR-GCK | Mflv_1700 | moaC | 1e-50 | 68.97% (145) | molybdenum cofactor biosynthesis protein C |
| M. tuberculosis H37Rv | Rv0864 | moaC | 7e-71 | 81.37% (161) | molybdenum cofactor biosynthesis protein C |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_0864 | - | 3e-54 | 62.96% (162) | molybdenum cofactor biosynthesis protein MoaC2 |
| M. marinum M | MMAR_4668 | moaC2 | 3e-92 | 97.71% (175) | molybdenum cofactor biosynthesis protein C 2 MoaC2 |
| M. avium 104 | MAV_0993 | moaC | 1e-59 | 80.14% (141) | molybdenum cofactor biosynthesis protein C |
| M. smegmatis MC2 155 | MSMEG_5703 | moaC | 2e-61 | 77.63% (152) | molybdenum cofactor biosynthesis protein C |
| M. thermoresistible (build 8) | TH_1354 | moaC | 7e-64 | 75.16% (161) | molybdenum cofactor biosynthesis protein C |
| M. vanbaalenii PYR-1 | Mvan_5052 | moaC | 6e-58 | 71.15% (156) | molybdenum cofactor biosynthesis protein C |
CLUSTAL 2.0.9 multiple sequence alignment
Mflv_1700|M.gilvum_PYR-GCK ----------------------------------------MVDVTAKDAT
Mvan_5052|M.vanbaalenii_PYR-1 -------------------------MPDRLSHLDETGAAHMVDVTAKDAT
MUL_0286|M.ulcerans_Agy99 MGWPSPPPNASKAPAASATPGAGSYPSGGLSHLDERGAAHMVDATEKGAT
MMAR_4668|M.marinum_M ---MAEPPNASKAPAASATPGAGSYPSGGLSHLDERGAAHMVDVTEKGAT
Mb0888|M.bovis_AF2122/97 ----------------------------------------MVDITEKATT
Rv0864|M.tuberculosis_H37Rv ---------------MARASGASDYRSGELSHQDERGAAHMVDITEKATT
MAV_0993|M.avium_104 ----------------------------------------MVDVSGKAPS
TH_1354|M.thermoresistible__bu ---------------VPEPAGAGS-SAPRLSHLDDTGAAHMVDVTAKAPT
MSMEG_5703|M.smegmatis_MC2_155 ---------------------------MELSHLDESGAAHMVDVTNKDVT
MAB_0864|M.abscessus_ATCC_1997 --------------MPEGLRASGSSGPEGLSHIDEHGSAHMVDVSAKQAT
*** : * :
Mflv_1700|M.gilvum_PYR-GCK SRIAVAEGTVHTRPDVVDMITANGLPKGDALATARIAGILAAKRTSDLVP
Mvan_5052|M.vanbaalenii_PYR-1 KRIAVASGTVHTRPDVVELITANGLPKGDALATARVAGIMAAKRTSDLVP
MUL_0286|M.ulcerans_Agy99 RRTAVAAGFLRTSTEVVALISSGGLPKGDALATARVAGILAAKRTSDLIP
MMAR_4668|M.marinum_M RRTAVAAGFLRTSTEVVALISSGGLPKGDALATARVAGILAAKRTSDLIP
Mb0888|M.bovis_AF2122/97 KRTAVAAGILRTSAQVVALISTGGLPKGDALATARVAGIMAAKRTSDLIP
Rv0864|M.tuberculosis_H37Rv KRTAVAAGILRTSAQVVALISTGGLPKGDALATARVAGIMAAKRTSDLIP
MAV_0993|M.avium_104 KRTAVAAGVLRTSADVVALISTGGLPKGDALATARVAGIQAAKRTSDLIP
TH_1354|M.thermoresistible__bu KREAIAAGVLWTRTDVVAMIAAGGLPKGDALATARVAGIMAAKRTSDLIP
MSMEG_5703|M.smegmatis_MC2_155 KRTAVAAGTLLTRPDVVALIASGGLPKGDALATARVAGILAAKRTPDLIP
MAB_0864|M.abscessus_ATCC_1997 AREAVAAGRFVTTPEVLKLLSDGTLPKGDAIATARLAAIMAAKRTSELIP
* *:* * . * .:*: ::: . ******:****:*.* *****.:*:*
Mflv_1700|M.gilvum_PYR-GCK LCHPLAITGVDIDFTVGGDEAPGSVTITATVRTTDRTGVEMEALTAVSVA
Mvan_5052|M.vanbaalenii_PYR-1 LCHPLAITGVDIDFTIGGDDDLGAVAITATVRTTDRTGVEMEALTAVSVA
MUL_0286|M.ulcerans_Agy99 LCHQLALTGVDVDFTMAEAD----VEITATVRSTDRTGVEMEALTAVIVV
MMAR_4668|M.marinum_M LCHQLALTGVDVDFTMAEAD----VEITATVRSTDRTGVEMEALTAVSVA
Mb0888|M.bovis_AF2122/97 LCHQLALTGVDVDFTVGQLD----IEITATVRSTDRTGVEMEALTAVSVA
Rv0864|M.tuberculosis_H37Rv LCHQLALTGVDVDFTVGQLD----IEITATVRSTDRTGVEMEALTAVSVA
MAV_0993|M.avium_104 LCHQLALTGVDVDFAVGESD----IEITATVTSTDRTGVEMEALTAVSVA
TH_1354|M.thermoresistible__bu LCHQLALTGVDVDFDIGEAH----IDITATVRTTDRTGVEMEALTAVSVA
MSMEG_5703|M.smegmatis_MC2_155 LCHQLALTGVDVDFDVRDDA----VGITATVRSTDRTGVEMEALTAVGVA
MAB_0864|M.abscessus_ATCC_1997 LCHQIPLTGIEVDFAIDATAG--SIQITAVARTKDRTGVEMEALTAVTVA
*** :.:**:::** : : ***.. :.************* *.
Mflv_1700|M.gilvum_PYR-GCK ALTVYDMIKAVDRAAGIDGIRVLRKEGGKTGSWTR---
Mvan_5052|M.vanbaalenii_PYR-1 ALTVYDMIKAVDRAARIDDIRVVRKEGGKTGTWVR---
MUL_0286|M.ulcerans_Agy99 ALTLYDMIKAVDPAARVDDVRVLSKQGGRSGSWSR---
MMAR_4668|M.marinum_M ALTLYDMIKAVDPAARIDDVRVLSKQGGRSGSWSR---
Mb0888|M.bovis_AF2122/97 ALTLYDMIKAVDPGALIDDIRVLHKEGGRRGTWTRR--
Rv0864|M.tuberculosis_H37Rv ALTLYDMIKAVDPGALIDDIRVLHKEGGRRGTWTRR--
MAV_0993|M.avium_104 GLTLYDMIKAVDPAARIDDVRVLRKEGGKTGMWTRS--
TH_1354|M.thermoresistible__bu ALTLYDMIKAVDPAARIGDIRVLRKEGGKTGVWTRP--
MSMEG_5703|M.smegmatis_MC2_155 ALTLYDMIKAVDPAARIDNIHVVRKEGGKTGLWEKPAE
MAB_0864|M.abscessus_ATCC_1997 GLTLHDMVKAVDPASRIDDVRVLRKEGGKTGTWTRP--
.**::**:**** .: :..::*: *:**: * * :