For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
FANIGWGEMLVLVIAGLVILGPERLPGAIRWTSGALRQARDYVSGATSQLRQDLGPEFDDLREPLQELQK LRGMTPRAAITKHLLDGDDSFLTGKFDDERPKQQPQTPAAQDKPEEKPDKPAGPVFDPDAT*
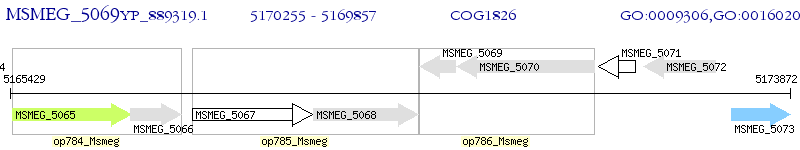
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_5069 | - | - | 100% (132) | sec-independent translocase |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb1256 | tatB | 4e-47 | 69.17% (133) | sec-independent translocase |
| M. gilvum PYR-GCK | Mflv_2203 | - | 5e-52 | 71.83% (142) | sec-independent translocase |
| M. tuberculosis H37Rv | Rv1224 | tatB | 4e-46 | 68.42% (133) | sec-independent translocase |
| M. leprae Br4923 | MLBr_01079 | - | 2e-44 | 65.15% (132) | sec-independent translocase |
| M. abscessus ATCC 19977 | MAB_1365 | - | 9e-43 | 61.76% (136) | sec-independent translocase |
| M. marinum M | MMAR_4213 | tatB | 2e-44 | 71.20% (125) | protein TatB |
| M. avium 104 | MAV_1367 | - | 4e-47 | 68.94% (132) | sec-independent translocase |
| M. thermoresistible (build 8) | TH_0297 | - | 3e-48 | 65.28% (144) | Probable protein TatB |
| M. ulcerans Agy99 | MUL_4516 | tatB | 3e-50 | 72.39% (134) | sec-independent translocase |
| M. vanbaalenii PYR-1 | Mvan_4493 | - | 3e-52 | 70.42% (142) | sec-independent translocase |
CLUSTAL 2.0.9 multiple sequence alignment
Mb1256|M.bovis_AF2122/97 MFANIGWGEMLVLVMVGLVVLGPERLPGAIRWAASALRQARDYLSGVTSQ
Rv1224|M.tuberculosis_H37Rv MFANIGWWEMLVLVMVGLVVLGPERLPGAIRWAASALRQARDYLSGVTSQ
MLBr_01079|M.leprae_Br4923 MFANIGWGEMLVLVVVGLVVLGPERFPGAIRWTLGALRQTRDYLSGVTNQ
MMAR_4213|M.marinum_M ---------MLVLVVVGLVVLGPERLPGAIRWSSGALRQARDYLSGVTSQ
MUL_4516|M.ulcerans_Agy99 MFANIGWGEMLVLVVVGLVVLGPERLPGAIRWSSGALRQARDYLSGVTSQ
MSMEG_5069|M.smegmatis_MC2_155 MFANIGWGEMLVLVIAGLVILGPERLPGAIRWTSGALRQARDYVSGATSQ
TH_0297|M.thermoresistible__bu MFSNVGWGEMVILVIAGLVILGPERLPGAIRWTANALRQVRDYVSGATSQ
Mflv_2203|M.gilvum_PYR-GCK MFANVGWGEMLVLVIAGLVILGPERLPGAIRWTAGAVRQARDYITGATSQ
Mvan_4493|M.vanbaalenii_PYR-1 MFANVGWGEMLVLVIAGLVILGPERLPGAIRWTAGAVRQARDYVTGATSQ
MAV_1367|M.avium_104 MLGSLSWEHMLVLVVVGLVVLGPERLPGAIRWTSNALRQARDYLSGVTTQ
MAB_1365|M.abscessus_ATCC_1997 MFGSVGWGELLVLLIVGLVVLGPERLPGAIRWTTESLRKVRDYASGATAS
:::*::.***:*****:******: ::*:.*** :*.* .
Mb1256|M.bovis_AF2122/97 LREDIGPEFDDLRGHLGELQKLRGMTPRAALTKHLLDGDDSLFTGDFD--
Rv1224|M.tuberculosis_H37Rv LREDIGPEFDDLRGHLGELQKLRGMTPRAALTKHLLDGDDSLFTGDFD--
MLBr_01079|M.leprae_Br4923 LREDIGPEFDDLRGQFGELQKLRGMTPRAALTKHLLDGDDSLFTGNFD--
MMAR_4213|M.marinum_M LRDDMGPEFDDLRGQLGELQKLRGMTPRAALTKHLLDGDDSIFTGNFD--
MUL_4516|M.ulcerans_Agy99 LRDDMGPEFDDLRGQLGELQKLRGMTPRAALTKHLLDGDDSIFTGNFD--
MSMEG_5069|M.smegmatis_MC2_155 LRQDLGPEFDDLREPLQELQKLRGMTPRAAITKHLLDGDDSFLTGKFD--
TH_0297|M.thermoresistible__bu LREEFGPEFDDLREPLSELQKLRGMTPRAALTKHLLDGDDSIFTGRFDTP
Mflv_2203|M.gilvum_PYR-GCK LREDLGPEFDDLREPLSELQKLRGMTPRAALTKHLLDGDDSIFTGRFDST
Mvan_4493|M.vanbaalenii_PYR-1 LREELGTDFDDLREPLSELQRLRGMTPRAALTKHLLDGDDSIFTGKFDQN
MAV_1367|M.avium_104 LREDLGPEFDDLRVPLSELQKLRGMTPRAALTKHLLDGDDSFLTGAFD--
MAB_1365|M.abscessus_ATCC_1997 LREELGPEFDDVRKPLAELQKLRGMTPRAVITKHLLDGDDSVFDSLTR--
**:::*.:***:* : ***:********.:**********.: .
Mb1256|M.bovis_AF2122/97 -----------RPTPKKPDAAGSAGPDATEQIGAGPIPFDSDAT
Rv1224|M.tuberculosis_H37Rv -----------RPTPKKPDAAGSAGPDATEQIGAGPIPFDSDAT
MLBr_01079|M.leprae_Br4923 -----------RPAAAKQQDR------DHHQT-----PFDTDAT
MMAR_4213|M.marinum_M -----------KAASATPAVDAVASAQEAPDEPVRP-PFDSDAT
MUL_4516|M.ulcerans_Agy99 -----------KAASATPAVDAVASAQEAPDEPVRP-PFDSDAT
MSMEG_5069|M.smegmatis_MC2_155 ---------DERPKQQPQ-TPAAQDKPEEKPDKPAGPVFDPDAT
TH_0297|M.thermoresistible__bu GSDGAGSAADSRAPHPPSGTQSNTVQPVDAETPVRTPKIDPDAT
Mflv_2203|M.gilvum_PYR-GCK SSDQPGSGKPPKPQSGPG--PAAAS-GPAATTTPASTPFDPDAT
Mvan_4493|M.vanbaalenii_PYR-1 GKSEKPEQKPEKPQSAPG--PAAAVPDQPAGGRSGSTPYDTDAT
MAV_1367|M.avium_104 -----------RPVNGAAAQPPPAPAPPPEPHRPGQTPFDADAT
MAB_1365|M.abscessus_ATCC_1997 ------PLDDVKKAVTEPAPTPIVNPELAKPAEPGPTRYDADAT
: *.***