For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
KRFVVAGFALVVFVELAALAVPGRSMIGWATGAALAVLLVSVRLALREDPVHDARSTASEDAAELLARWQ SETEILISRADSTRSQWDRHLRPRLAREFAAASGQRLVGDQSAFQATGRMLFGERLWQWVDPNNVASARD DGPGPGRAVLAEILQRLEQL*
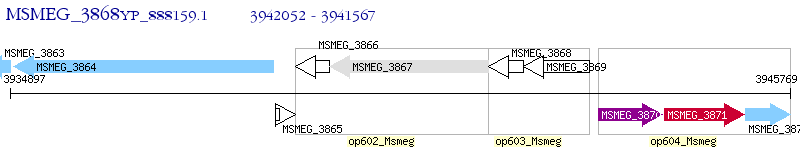
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. smegmatis MC2 155 | MSMEG_3868 | - | - | 100% (161) | hypothetical protein MSMEG_3868 |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb3190c | - | 5e-38 | 50.31% (161) | hypothetical protein Mb3190c |
| M. gilvum PYR-GCK | Mflv_0477 | - | 7e-38 | 49.69% (159) | hypothetical protein Mflv_0477 |
| M. tuberculosis H37Rv | Rv3165c | - | 5e-38 | 50.31% (161) | hypothetical protein Rv3165c |
| M. leprae Br4923 | - | - | - | - | - |
| M. abscessus ATCC 19977 | MAB_1910c | - | 4e-28 | 43.03% (165) | hypothetical protein MAB_1910c |
| M. marinum M | MMAR_1398 | - | 3e-38 | 49.69% (161) | hypothetical protein MMAR_1398 |
| M. avium 104 | MAV_4053 | - | 3e-36 | 47.13% (157) | hypothetical protein MAV_4053 |
| M. thermoresistible (build 8) | TH_0585 | - | 3e-37 | 48.45% (161) | conserved hypothetical protein |
| M. ulcerans Agy99 | - | - | - | - | - |
| M. vanbaalenii PYR-1 | Mvan_0179 | - | 2e-36 | 47.20% (161) | hypothetical protein Mvan_0179 |
CLUSTAL 2.0.9 multiple sequence alignment
Mb3190c|M.bovis_AF2122/97 MKRLIALGIFLIVGIELLALILHDRRLVLAGSGLALALVLLNVRRMLGNR
Rv3165c|M.tuberculosis_H37Rv MKRLIALGIFLIVGIELLALILHDRRLVLAGSGLALALVLLNVRRMLGNR
MAV_4053|M.avium_104 ----MALGLCLIIGIELLALTQHDRRLVLTASGVGLALVLLSLRRALGRG
MMAR_1398|M.marinum_M MKKLIVLGVLLIVAVELLTLAVRDHRLVLAASGVAVALVLLNIRRVVGHA
Mflv_0477|M.gilvum_PYR-GCK MTKLVVGGVVLVIAVQAFALTQLEREVVVAVTGGALACVLIAVRARLDHD
Mvan_0179|M.vanbaalenii_PYR-1 MTKVVVVGAFLIVVVQMLAVTQLDRDIVLTVTGGALACVLIAVRTHLTHQ
MSMEG_3868|M.smegmatis_MC2_155 MKRFVVAGFALVVFVELAALAVPGRSMIGWATGAALAVLLVSVRLALRED
TH_0585|M.thermoresistible__bu MTRLMAAAFAVVILVEAATLAAPERRLVLIVTGIAVAAALLAVRVRLGRA
MAB_1910c|M.abscessus_ATCC_199 MKWLVPAVLLLVVLTEACVAASPLRAFTLPITGVVVGTALGLLWGNLRSA
: ::: : . : . :* :. * : :
Mb3190c|M.bovis_AF2122/97 DELTA-APDSDDLGEGLRRWLSNTETTIRWSESTRADWDRHLRPMLARRF
Rv3165c|M.tuberculosis_H37Rv DELTA-APDSDDLGEGLRRWLSNTETTIRWSESTRADWDRHLRPMLARRF
MAV_4053|M.avium_104 AERES-EHDADDLGDSLRRWLAATETTIRWSESTRADWDRHLRPMLARRY
MMAR_1398|M.marinum_M MAPPA-ETDPDDVGDGLRRWLSGTETTIRRSESTRADWDRHLRPMLARRY
Mflv_0477|M.gilvum_PYR-GCK AVPDD-DARADDAAAAIRRWLSRTETLISWSESSRSDWDRRLRPMLARQF
Mvan_0179|M.vanbaalenii_PYR-1 EGDVD-DPVTDEAAASMRRWLSRTETLISWSESSRSDWDRRLRPMLARQF
MSMEG_3868|M.smegmatis_MC2_155 PVHDARSTASEDAAELLARWQSETEILISRADSTRSQWDRHLRPRLAREF
TH_0585|M.thermoresistible__bu ADPTEPDPAAADRAAALQRWLSRTHTLIGWSEAGRSDWDRHLRPLLARQL
MAB_1910c|M.abscessus_ATCC_199 RADMPSSPASLPADETLRRWHARAEARISWAERTRGDWDRQLRPLIAADF
. : ** : :. * :: *.:***:*** :*
Mb3190c|M.bovis_AF2122/97 EIATGHRQAKDPVAFAATGRMLFGDELWEWVNPNNVTHTGDRQPGPGRAA
Rv3165c|M.tuberculosis_H37Rv EIATGHRQAKDPVAFAATGRMLFGDELWEWVNPNNVTHTGDRQPGPGRAA
MAV_4053|M.avium_104 AIATGQRQAKDPASFQATGQMLFGAELWEWVNPNNVTRTGGRQPGPGRAA
MMAR_1398|M.marinum_M EIATGQRQAKDPAAFHATGRMLFGEDLWEWVNPNNVARTGDRQAGPGRAT
Mflv_0477|M.gilvum_PYR-GCK ELAAGRRKAKDPRAFEATGRMVFGDELWQWVDPDNISRSGSLEPGPGRRT
Mvan_0179|M.vanbaalenii_PYR-1 ELAAGRRKIKDPRAFDATGRMVFGDELWQWVDPENISRTGGQEQGPGRRT
MSMEG_3868|M.smegmatis_MC2_155 AAASGQRLVGDQSAFQATGRMLFGERLWQWVDPNNVASARDDGPGPGRAV
TH_0585|M.thermoresistible__bu EMATGQPRRRDPNGYRAAGRLLLGPRLWPWVDPDNVSSAAAHQPGPGRAV
MAB_1910c|M.abscessus_ATCC_199 ETAAGHRQSRNPAALAATGRMTFGPELWRWVDPNGASDTDRAAPAPGRAV
*:*: : . *:*:: :* ** **:*:. : : .*** .
Mb3190c|M.bovis_AF2122/97 LEEILQKLEQV
Rv3165c|M.tuberculosis_H37Rv LEEILQKLEQV
MAV_4053|M.avium_104 LEEILRKLEQV
MMAR_1398|M.marinum_M LEAILQRLEQV
Mflv_0477|M.gilvum_PYR-GCK LDDILRRLERL
Mvan_0179|M.vanbaalenii_PYR-1 LDEILQRLEKL
MSMEG_3868|M.smegmatis_MC2_155 LAEILQRLEQL
TH_0585|M.thermoresistible__bu LQEILQRLERI
MAB_1910c|M.abscessus_ATCC_199 LQSILERLERM
* **.:**::