For questions or suggestions e-mail us at: ioerger@cs.tamu.edu
NTAEMALPDILDPMYWLGADGIFGSAVLPGILLIVFIETGLLFPLLPGESLLFTGGLLAASAHPPVTIGV LAPAVAVVAVLGDQTAYFIGRRIGPALFKKEDSRFFKKHYVTESHAFFEKYGPWTVILARFVPFARTFVP VIAGVSYMRYPVFLAFDIVGGISWGAGVTLAGYFLGNVPFVHHNLEKIILGILIVSLIPAMTAAWRGYRS RRRSPNNESDSSDPVVLPE*
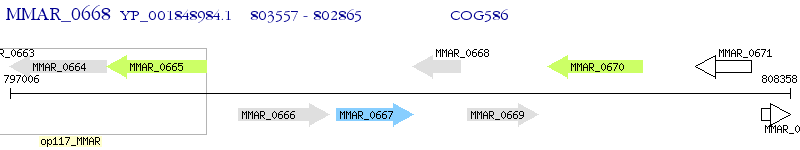
Operon Prediction Model: Genebank
Paralogs
| species | id | gene | e-value | identity (len) | annotation |
| M. marinum M | MMAR_0668 | - | - | 100% (230) | transmembrane protein |
| M. marinum M | MMAR_2063 | dedA | 2e-16 | 33.13% (163) | transmembrane protein DedA |
Closest Orthologs (e-value cutoff: 1e-4)
| species | id | gene | e-value | identity (len) | annotation |
| M. bovis AF2122 / 97 | Mb0371 | - | 1e-107 | 80.87% (230) | transmembrane protein |
| M. gilvum PYR-GCK | Mflv_0237 | - | 1e-45 | 43.35% (203) | hypothetical protein Mflv_0237 |
| M. tuberculosis H37Rv | Rv0364 | - | 1e-107 | 81.30% (230) | transmembrane protein |
| M. leprae Br4923 | MLBr_00287 | - | 3e-92 | 72.60% (219) | hypothetical protein MLBr_00287 |
| M. abscessus ATCC 19977 | MAB_4661 | - | 8e-70 | 57.14% (210) | hypothetical protein MAB_4661 |
| M. avium 104 | MAV_4785 | - | 1e-109 | 84.82% (224) | membrane protein DedA family protein |
| M. smegmatis MC2 155 | MSMEG_0750 | - | 4e-48 | 42.49% (233) | membrane protein |
| M. thermoresistible (build 8) | TH_1640 | - | 3e-81 | 63.27% (226) | POSSIBLE CONSERVED TRANSMEMBRANE PROTEIN |
| M. ulcerans Agy99 | MUL_0104 | - | 1e-131 | 99.57% (230) | transmembrane protein |
| M. vanbaalenii PYR-1 | Mvan_0668 | - | 3e-47 | 43.98% (216) | hypothetical protein Mvan_0668 |
CLUSTAL 2.0.9 multiple sequence alignment
MMAR_0668|M.marinum_M --------MNTAEMALPDILDPMYWLGADGIFGSAVLPGILLIVFIETGL
MUL_0104|M.ulcerans_Agy99 --------MNTAEMALPDILDPMYWLGADGIFGSAVLPGILLIVFIETGL
Mb0371|M.bovis_AF2122/97 --------MSTAVTAMPDILDPMYWLGANGVFGSAVLPGILIIVFIETGL
Rv0364|M.tuberculosis_H37Rv --------MSTAVTAMPDILDPMYWLGANGVFGSAVLPGILIIVFIETGL
MAV_4785|M.avium_104 --------MSTAVTALPDILDPMYWLGADGVFGSAVLPGILVIVFIETGL
MLBr_00287|M.leprae_Br4923 --------MNTAIVALPPTLDPMFWIGPEGFFAAAMLPATLVIIFVETGL
TH_1640|M.thermoresistible__bu --------MSTTVMAMPGILDPMFWIGPDGLFAAAVLPAILIIVFIETGL
MAB_4661|M.abscessus_ATCC_1997 --------MDMG-LAPADVMDPMYWLGEGGLFGGAVLAGVMVIVFIETGL
Mflv_0237|M.gilvum_PYR-GCK -------MVVDNLALMPDFLDPINLLS---YFGTWALIGLLVVVFVESGV
Mvan_0668|M.vanbaalenii_PYR-1 -------MVVDNLALMPDFLDPINLLS---YFGTWALIGLLVVVFVESGV
MSMEG_0750|M.smegmatis_MC2_155 MIDTALPEVTTNLALMPDFMDPLNLIG---YFGTWALVGILLVVFIESGV
: . :**: :. *. * . ::::*:*:*:
MMAR_0668|M.marinum_M LFPLLPGESLLFTGGLLAAS----------AHPPVTIGVLAPAVAVVAVL
MUL_0104|M.ulcerans_Agy99 LLPLLPGESLLFTGGLLAAS----------AHPPVTIGVLAPAVAVVAVL
Mb0371|M.bovis_AF2122/97 LFPLLPGESLLFTGGLLSAS----------PAPPVTIGVLAPCVALVAVL
Rv0364|M.tuberculosis_H37Rv LFPLLPGESLLFTGGLLSAS----------PAPPVTIGVLAPCVALVAVL
MAV_4785|M.avium_104 LFPLLPGESLLFTGGLLAAH----------PNPPASIWVLAPAVAIVAVL
MLBr_00287|M.leprae_Br4923 LFPLLPGESLLFTGGLLATK----------G--TIDIWVLSPSVAVVAVL
TH_1640|M.thermoresistible__bu LFPLLPGDSLLFTGGLLAAG----------G--TMNIWLLCITVTIVAIL
MAB_4661|M.abscessus_ATCC_1997 LFPFLPGDTLLFSAGLIAAQ----------PNSPVAIQVLAPCAALAALL
Mflv_0237|M.gilvum_PYR-GCK LFPILPGDSLLFVAGMLAAGISAQSAEGNVAVVNFQLWQLLVFIPIAAVL
Mvan_0668|M.vanbaalenii_PYR-1 LFPVLPGDSLLFVAGMLAAGTAAASKEGNVANADFQLWQLLVFIPIAAVL
MSMEG_0750|M.smegmatis_MC2_155 LFPVLPGDSLLFVAGMLAAGTAAK---GTAVDANFQLWQLLVFIPIAAIL
*:*.***::*** .*:::: : * .:.*:*
MMAR_0668|M.marinum_M GDQTAYFIGRRIGPALFKKEDSRFFKKHYVTESHAFFEKYGPWTVILARF
MUL_0104|M.ulcerans_Agy99 GDQTAYFIGRRIGPALFKKEDSRFFKKHYVTESHAFFEKYGPWTVILARF
Mb0371|M.bovis_AF2122/97 GDQTAYFIGRRIGPALFKKEDSRFFKKHYVTESHAFFEKYGKWTIILARF
Rv0364|M.tuberculosis_H37Rv GDQTAYFIGRRIGPALFKKEDSRFFKKHYVTESHAFFEKYGKWTIILARF
MAV_4785|M.avium_104 GDQTGYFIGRRIGPALFKKEDSRFFKKHYVTESHAFFEKYGSWAVILARF
MLBr_00287|M.leprae_Br4923 GDQIGYLIGRRIGPALFKKENSRFFKQHYVTESHAFFEKHGRWTIILARF
TH_1640|M.thermoresistible__bu GDQCAYYIGRRIGPALFNKEDSRFFKKSYVEQSHEFFEKYGPYTIILARF
MAB_4661|M.abscessus_ATCC_1997 GSQCGYFIGRRLGPALFKKEDARFFKQRYLTVSREFFDKHGPKTLLIAQF
Mflv_0237|M.gilvum_PYR-GCK GAQVGYWIGRNVGTAMFTP-NARFLKQKYLDEAHLFFEQRGPFAIVIARF
Mvan_0668|M.vanbaalenii_PYR-1 GAQVGYWIGRNVGTAMFKP-DARFLKQKYLDEAHLFFEQRGPFAIVVARF
MSMEG_0750|M.smegmatis_MC2_155 GGQVGYIIGRTIGTSMFKP-EARILKQKYLDEAHEFFEQRGPFAIVIARF
* * .* *** :*.::*. ::*::*: *: :: **:: * ::::*:*
MMAR_0668|M.marinum_M VPFARTFVPVIAGVSYMRYPVFLAFDIVGGISWGAGVTLAGYFLGNVPFV
MUL_0104|M.ulcerans_Agy99 VPFARTFVPVIAGVSYMRYPVFLAFDIVGGISWGAGVTLAGYFLGNVPFV
Mb0371|M.bovis_AF2122/97 VPIARTFVPVIAGVSYMRYPVFLGFDIVGGVAWGAGVTLAGYFLGSVPFV
Rv0364|M.tuberculosis_H37Rv VPIARTFVPVIAGVSYMRYPVFLGFDIVGGVAWGAGVTLAGYFLGSVPFV
MAV_4785|M.avium_104 APFVRTFVPVIAGVSYMRYPLFLGFDIVGGIAWGGGATLAGYFLGNVPFV
MLBr_00287|M.leprae_Br4923 LPFMRTFTPVIAGLSYMSYPLYLGFDIVGGILWGGGVTVAGYFLGNVPFV
TH_1640|M.thermoresistible__bu VPIVRTYAPVLAGVSYMRYSVFLTFNAVGGIAWGTGVTLLGYFLGNITFV
MAB_4661|M.abscessus_ATCC_1997 IGVVRTFTPVIAGMSGMRYPTFLLYNAIGSAAWGTGLTVVGYFLGNIAFV
Mflv_0237|M.gilvum_PYR-GCK VPIVRTLAPITAGAAKMNYAVFTLYNAVGAIVWGVGLTLLGFFLGRIEII
Mvan_0668|M.vanbaalenii_PYR-1 VPIVRTLAPITAGAAKMNYAVFTVYNAVGAVVWGAGLTLLGYWLGRFEVI
MSMEG_0750|M.smegmatis_MC2_155 VPIVRTLAPITAGAAKMRYPVFTAFNVLGAIVWGVGLTLLGYWLGQFEII
. ** .*: ** : * *. : :: :*. ** * *: *::** . .:
MMAR_0668|M.marinum_M HHNLEKIILGILIVSLIPAMTAAWRGYRSRRRSPNNESDSSDPVVLPE-
MUL_0104|M.ulcerans_Agy99 HHNLEKIILGILIVSLIPAMTAAWRGYRSRRRSPNNESDSSDPVVLPE-
Mb0371|M.bovis_AF2122/97 HMNFQLIILALVFVSLLPALVSAARVYRARRNAP--QSDP-DPLVLPE-
Rv0364|M.tuberculosis_H37Rv HMNFQLIILAIVFVSLLPALVSAARVYRARRNAP--QSDP-DPLVLPE-
MAV_4785|M.avium_104 HHNLEKIILGILVVSLMPAFVAAWRGYRGRRRPTRTPERESERLPASE-
MLBr_00287|M.leprae_Br4923 RQNLEKIILGILFVSLLPALIAAWHGYRSQSRTAK------SELALPD-
TH_1640|M.thermoresistible__bu KENLEYMILLIVFLSVLPMIFSVTKGMLERRRARSGPALAAEPLPKQSG
MAB_4661|M.abscessus_ATCC_1997 GEHLDIIILVIAILSTLPAAATAAKVYLEKRRAVTENG-----------
Mflv_0237|M.gilvum_PYR-GCK QKLLEPIVIGIVILSVLPIAIEWYKRRKAAKQAGIATPPMEAGEQAG--
Mvan_0668|M.vanbaalenii_PYR-1 QKLLEPIVIGIVLLSVLPMLIEWYRRRKAAKQAGGATEAEPTG------
MSMEG_0750|M.smegmatis_MC2_155 QKLLEPIFILIVVASVAPMFIEWYKRRRAAKRASSTDTATDAASPAS--
:: :.: : . * * : ..